Abstract
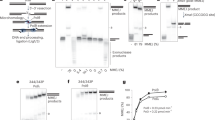
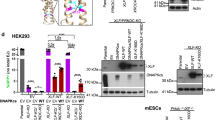
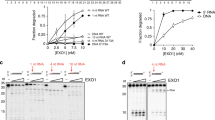
DNA double-strand breaks (DSBs) can be repaired either via homologous recombination (HR) or nonhomologous end-joining (NHEJ). Both pathways are operative in eukaryotes, but bacteria had been thought to rely on HR alone. Here we provide direct evidence that mycobacteria have a robust NHEJ pathway that requires Ku and a specialized polyfunctional ATP-dependent DNA ligase (LigD). NHEJ of blunt-end and complementary 5′-overhang DSBs is highly mutagenic (∼50% error rate). Analysis of the recombination junctions ensuing from individual NHEJ events highlighted the participation of several DNA end-remodeling activities, including template-dependent fill-in of 5′ overhangs, nontemplated addition of single nucleotides at blunt ends, and nucleolytic resection. LigD itself has the template-dependent and template-independent polymerase functions in vitro that compose the molecular signatures of NHEJ in vivo. Another ATP-dependent DNA ligase (LigC) provides a backup mechanism for LigD-independent error-prone repair of blunt-end DSBs. We speculate that NHEJ allows mycobacteria to evade genotoxic host defense.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Buchmeier, N.A. et al. DNA repair is more important than catalase for Salmonella virulence in mice. J. Clin. Invest. 95, 1047–1053 (1995).
Buchmeier, N.A., Lipps, C.J., So, M.Y. & Heffron, F. Recombination-deficient mutants of Salmonella typhimurium are avirulent and sensitive to the oxidative burst of macrophages. Mol. Microbiol. 7, 933–936 (1993).
O'Rourke, E.J. et al. Pathogen DNA as target for host-generated oxidative stress: role for repair of bacterial DNA damage in Helicobacter pylori colonization. Proc. Natl. Acad. Sci. USA 100, 2789–2794 (2003).
Suvarnapunya, A.E., Lagasse, H.A. & Stein, M.A. The role of DNA base excision repair in the pathogenesis of Salmonella enterica serovar Typhimurium. Mol. Microbiol. 48, 549–559 (2003).
Sander, P. et al. Mycobacterium bovis BCG recA deletion mutant shows increased susceptibility to DNA-damaging agents but wild-type survival in a mouse infection model. Infect. Immun. 69, 3562–3568 (2001).
Darwin, K.H., Ehrt, S., Gutierrez-Ramos, J.C., Weich, N. & Nathan, C.F. The proteasome of Mycobacterium tuberculosis is required for resistance to nitric oxide. Science 302, 1963–1966 (2003).
Boshoff, H.I., Reed, M.B., Barry, C.E. & Mizrahi, V. DnaE2 polymerase contributes to in vivo survival and the emergence of drug resistance in Mycobacterium tuberculosis. Cell 113, 183–193 (2003).
Cromie, G.A., Connelly, J.C. & Leach, D.R. Recombination at double-strand breaks and DNA ends: conserved mechanisms from phage to humans. Mol. Cell 8, 1163–1174 (2001).
Lieber, M.R., Ma, Y., Pannicke, U. & Schwarz, K. Mechanism and regulation of human non-homologous DNA end-joining. Nat. Rev. Mol. Cell Biol. 4, 712–720 (2003).
Downs, J.A. & Jackson, S.P. A means to a DNA end: the many roles of Ku. Nat. Rev. Mol. Cell Biol. 5, 367–378 (2004).
Riballo, E. et al. Identification of a defect in DNA ligase IV in a radiosensitive leukaemia patient. Curr. Biol. 9, 699–702 (1999).
Barnes, D.E., Tomkinson, A.E., Lehmann, A.R., Webster, A.D. & Lindahl, T. Mutations in the DNA ligase I gene of an individual with immunodeficiencies and cellular hypersensitivity to DNA-damaging agents. Cell 69, 495–503 (1992).
Schar, P., Herrmann, G., Daly, G. & Lindahl, T. A newly identified DNA ligase of Saccharomyces cerevisiae involved in RAD52-independent repair of DNA double-strand breaks. Genes Dev. 11, 1912–1924 (1997).
Teo, S.H. & Jackson, S.P. Identification of Saccharomyces cerevisiae DNA ligase IV: involvement in DNA double-strand break repair. EMBO J. 16, 4788–4795 (1997).
Wilson, T.E., Grawunder, U. & Lieber, M.R. Yeast DNA ligase IV mediates non-homologous DNA end joining. Nature 388, 495–498 (1997).
Frank, K.M. et al. Late embryonic lethality and impaired V(D)J recombination in mice lacking DNA ligase IV. Nature 396, 173–177 (1998).
Grawunder, U., Zimmer, D., Fugmann, S., Schwarz, K. & Lieber, M.R. DNA ligase IV is essential for V(D)J recombination and DNA double-strand break repair in human precursor lymphocytes. Mol. Cell 2, 477–484 (1998).
Konrad, E.B., Modrich, P. & Lehman, I.R. Genetic and enzymatic characterization of a conditional lethal mutant of Escherichia coli K12 with a temperature-sensitive DNA ligase. J. Mol. Biol. 77, 519–529 (1973).
Magnet, S. & Blanchard, J.S. Mechanistic and kinetic study of the ATP-dependent DNA ligase of Neisseria meningitidis. Biochemistry 43, 710–717 (2004).
Wilkinson, A., Day, J. & Bowater, R. Bacterial DNA ligases. Mol. Microbiol. 40, 1241–1248 (2001).
Cheng, C. & Shuman, S. Characterization of an ATP-dependent DNA ligase encoded by Haemophilus influenzae. Nucleic Acids Res. 25, 1369–1374 (1997).
Gong, C., Martins, A., Bongiorno, P., Glickman, M. & Shuman, S. Biochemical and genetic analysis of the four DNA ligases of mycobacteria. J. Biol. Chem. 279, 20594–20606 (2004).
Aravind, L. & Koonin, E.V. Prokaryotic homologs of the eukaryotic DNA-end-binding protein Ku, novel domains in the Ku protein and prediction of a prokaryotic double-strand break repair system. Genome Res. 11, 1365–1374 (2001).
Doherty, A.J., Jackson, S.P. & Weller, G.R. Identification of bacterial homologues of the Ku DNA repair proteins. FEBS Lett. 500, 186–188 (2001).
Della, M. et al. Mycobacterial Ku and ligase proteins constitute a two-component NHEJ repair machine. Science 306, 683–685 (2004).
Zhu, H. & Shuman, S. A primer-dependent polymerase function of Pseudomonas aeruginosa ATP-dependent DNA ligase (LigD). J. Biol. Chem. 280, 418–427 (2005).
Weller, G.R. et al. Identification of a DNA nonhomologous end-joining complex in bacteria. Science 297, 1686–1689 (2002).
Braunstein, M., Brown, A.M., Kurtz, S. & Jacobs, W.R. Jr. Two nonredundant SecA homologues function in mycobacteria. J. Bacteriol. 183, 6979–6990 (2001).
Lipps, G., Weinzierl, A.O., von Scheven, G., Buchen, C. & Cramer, P. Structure of a bifunctional DNA primase-polymerase. Nat. Struct. Mol. Biol. 11, 157–162 (2004).
Ito, N., Nureki, O., Shirouzu, M., Yokoyama, S. & Hanaoka, F. Crystal structure of the Pyrococcus horikoshii DNA primase-UTP complex: implications for the mechanism of primer synthesis. Genes Cells 8, 913–923 (2003).
Augustin, M.A., Huber, R. & Kaiser, J.T. Crystal structure of a DNA-dependent RNA polymerase (DNA primase). Nat. Struct. Biol. 8, 57–61 (2001).
Glickman, M.S. The mmaA2 gene of Mycobacterium tuberculosis encodes the distal cyclopropane synthase of the α-mycolic acid. J. Biol. Chem. 278, 7844–7849 (2003).
Manolis, K.G. et al. Novel functional requirements for non-homologous DNA end joining in Schizosaccharomyces pombe. EMBO J. 20, 210–221 (2001).
Walker, J.R., Corpina, R.A. & Goldberg, J. Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair. Nature 412, 607–614 (2001).
Aylon, Y., Liefshitz, B. & Kupiec, M. The CDK regulates repair of double-strand breaks by homologous recombination during the cell cycle. EMBO J. 23, 4868–4875 (2004).
Heidenreich, E., Novotny, R., Kneidinger, B., Holzmann, V. & Wintersberger, U. Non-homologous end joining as an important mutagenic process in cell cycle– arrested cells. EMBO J. 22, 2274–2283 (2003).
Ferreira, M.G. & Cooper, J.P. Two modes of DNA double-strand break repair are reciprocally regulated through the fission yeast cell cycle. Genes Dev. 18, 2249–2254 (2004).
Timm, J., Lim, E.M. & Gicquel, B. Escherichia coli– mycobacteria shuttle vectors for operon and gene fusions to lacZ: the pJEM series. J. Bacteriol. 176, 6749–6753 (1994).
Golemis, E. et al. The interaction trap. In Current Protocols in Molecular Biology (eds. Ausubel, F.M. et al.) 20.1.1–20.1.2 (Wiley, New York, 1999).
Acknowledgements
This research was supported by US National Institutes of Health grants AI53417 (to M.S.G.) and GM63611 (to S.S.). M.S.G. is the recipient of research awards from the Ellison Medical Foundation and the New York Academy of Medicine Speakers Fund for Biomedical Research. S.S. is an American Cancer Society Research Professor.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Mycobacterium smegmatis deletion strains. (PDF 19 kb)
Rights and permissions
About this article
Cite this article
Gong, C., Bongiorno, P., Martins, A. et al. Mechanism of nonhomologous end-joining in mycobacteria: a low-fidelity repair system driven by Ku, ligase D and ligase C. Nat Struct Mol Biol 12, 304–312 (2005). https://doi.org/10.1038/nsmb915
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb915