Key Points
-
Alternative splicing (AS) is a major contributor to transcriptome and proteome diversity. Evolutionary studies help to address questions that are fundamental to understanding this important process.
-
The main mechanism for exon selection in higher eukaryotes is exon definition: the splicing machinery is placed across exons, constraining their length.
-
Early eukaryotic ancestors are rich in introns, contain degenerate splicing signals and complex spliceosomes, and share homology of splicing factors in different species. These observations suggest an early eukaryotic origin of AS.
-
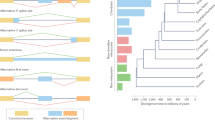
There are three known evolutionary mechanisms that could account for the appearance of an alternatively spliced exon: exon shuffling (a form of gene duplication), exonization of intronic sequences and transition of a constitutive exon to an alternative exon. The formation of an alternative exon permits new functions to be established without eliminating the original function of the protein.
-
Alu elements — primate-specific reteroelements — substantially contribute to the creation of new alternative exons, which can enhance the genomic repertoire.
-
Defining an alternative exon enables understanding of how splicing affects genome evolution. Comparative studies show conservation that indicates functionality, and these studies can help to identify factors that are involved in exon definition.
-
Recently, it was found that exons have increased nucleosome occupancy levels compared with introns; the nucleosome might act as a 'speed bump' on the exons, slowing RNA polymerase II. Exons were also found to be enriched in certain histone modifications.
-
This nucleosome positioning in exons encourages the 'correct' location of molecular interactions across the exon, which contributes to the exon definition mechanism and suggests another level of complexity in eukaryotic splicing regulation.
Abstract
Over the past decade, it has been shown that alternative splicing (AS) is a major mechanism for the enhancement of transcriptome and proteome diversity, particularly in mammals. Splicing can be found in species from bacteria to humans, but its prevalence and characteristics vary considerably. Evolutionary studies are helping to address questions that are fundamental to understanding this important process: how and when did AS evolve? Which AS events are functional? What are the evolutionary forces that shaped, and continue to shape, AS? And what determines whether an exon is spliced in a constitutive or alternative manner? In this Review, we summarize the current knowledge of AS and evolution and provide insights into some of these unresolved questions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Modrek, B. & Lee, C. A genomic view of alternative splicing. Nature Genet. 30, 13–19 (2002).
Tazi, J., Bakkour, N. & Stamm, S. Alternative splicing and disease. Biochim. Biophys. Acta 1792, 14–26 (2009).
Wang, G. S. & Cooper, T. A. Splicing in disease: disruption of the splicing code and the decoding machinery. Nature Rev. Genet. 8, 749–761 (2007).
Cartegni, L., Chew, S. L. & Krainer, A. R. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nature Rev. Genet. 3, 285–298 (2002).
Venables, J. P. Aberrant and alternative splicing in cancer. Cancer Res. 64, 7647–7654 (2004).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Chen, M. & Manley, J. L. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nature Rev. Mol. Cell Biol. 10, 741–754 (2009).
Hartmann, B. & Valcarcel, J. Decrypting the genome's alternative messages. Curr. Opin. Cell Biol. 21, 377–386 (2009).
Hui, J. Regulation of mammalian pre-mRNA splicing. Sci. China C Life Sci. 52, 253–260 (2009).
Licatalosi, D. D. & Darnell, R. B. RNA processing and its regulation: global insights into biological networks. Nature Rev. Genet. 11, 75–87 (2010).
Schwartz, S., Meshorer, E. & Ast, G. Chromatin organization marks exon–intron structure. Nature Struct. Mol. Biol. 16, 990–995 (2009). This paper shows that exons have increased nucleosome occupancy levels compared with introns, and four specific post-translational histone modifications are enriched in exons. This article, together with Tilgner et al . and Andersson et al . (references 98 and 99, respectively), presents evidence that the positioning and modifications of nucleosomes might help to define the exon–intron architecture of genes.
Ast, G. How did alternative splicing evolve? Nature Rev. Genet. 5, 773–782 (2004).
Ram, O. & Ast, G. SR proteins: a foot on the exon before the transition from intron to exon definition. Trends Genet. 23, 5–7 (2007).
Edgell, D. R., Belfort, M. & Shub, D. A. Barriers to intron promiscuity in bacteria. J. Bacteriol. 182, 5281–5289 (2000).
Watanabe, Y. et al. Introns in protein-coding genes in archaea. FEBS Lett. 510, 27–30 (2002).
Yokobori, S. et al. Gain and loss of an intron in a protein-coding gene in archaea: the case of an archaeal RNA pseudouridine synthase gene. BMC Evol. Biol. 9, 198 (2009).
Alekseyenko, A. V., Kim, N. & Lee, C. J. Global analysis of exon creation versus loss and the role of alternative splicing in 17 vertebrate genomes. RNA 13, 661–670 (2007).
Artamonova, I. I. & Gelfand, M. S. Comparative genomics and evolution of alternative splicing: the pessimists' science. Chem. Rev. 107, 3407–3430 (2007).
Kim, E., Goren, A. & Ast, G. Alternative splicing: current perspectives. Bioessays 30, 38–47 (2008).
Koren, E., Lev-Maor, G. & Ast, G. The emergence of alternative 3′ and 5′ splice site exons from constitutive exons. PLoS Comput. Biol. 3, e95 (2007).
Fox-Walsh, K. L. et al. The architecture of pre-mRNAs affects mechanisms of splice-site pairing. Proc. Natl Acad. Sci. USA 102, 16176–16181 (2005).
Sterner, D. A., Carlo, T. & Berget, S. M. Architectural limits on split genes. Proc. Natl Acad. Sci. USA 93, 15081–15085 (1996).
Carmel, L., Rogozin, I. B., Wolf, Y. I. & Koonin, E. V. Patterns of intron gain and conservation in eukaryotic genes. BMC Evol. Biol. 7, 192 (2007).
Li, W., Tucker, A. E., Sung, W., Thomas, W. K. & Lynch, M. Extensive, recent intron gains in Daphnia populations. Science 326, 1260–1262 (2009).
Farlow, A., Meduri, E., Dolezal, M., Hua, L. & Schlötterer, C. Nonsense-mediated decay enables intron gain in Drosophila. PLoS Genet. 6, e1000819 (2010).
Sela, N. et al. Comparative analysis of transposed element insertion within human and mouse genomes reveals Alu's unique role in shaping the human transcriptome. Genome Biol. 8, R127 (2007).
Schwartz, S. H. et al. Large-scale comparative analysis of splicing signals and their corresponding splicing factors in eukaryotes. Genome Res. 18, 88–103 (2008).
Barbosa-Morais, N. L., Carmo-Fonseca, M. & Aparicio, S. Systematic genome-wide annotation of spliceosomal proteins reveals differential gene family expansion. Genome Res. 16, 66–77 (2006).
Fedorov, A., Merican, A. F. & Gilbert, W. Large-scale comparison of intron positions among animal, plant, and fungal genes. Proc. Natl Acad. Sci. USA 99, 16128–16133 (2002).
Csuros, M., Rogozin, I. B. & Koonin, E. V. Extremely intron-rich genes in the alveolate ancestors inferred with a flexible maximum-likelihood approach. Mol. Biol. Evol. 25, 903–911 (2008).
Nguyen, H. D., Yoshihama, M. & Kenmochi, N. New maximum likelihood estimators for eukaryotic intron evolution. PLoS Comput. Biol. 1, e79 (2005).
Roy, S. W. & Irimia, M. Splicing in the eukaryotic ancestor: form, function and dysfunction. Trends Ecol. Evol. 24, 447–455 (2009). This review covers important aspects of eukaryotic evolution.
Rogozin, I. B., Wolf, Y. I., Sorokin, A. V., Mirkin, B. G. & Koonin, E. V. Remarkable interkingdom conservation of intron positions and massive, lineage-specific intron loss and gain in eukaryotic evolution. Curr. Biol. 13, 1512–1517 (2003).
Roy, S. W. & Gilbert, W. Rates of intron loss and gain: implications for early eukaryotic evolution. Proc. Natl Acad. Sci. USA 102, 5773–5778 (2005).
Carmel, L., Wolf, Y. I., Rogozin, I. B. & Koonin, E. V. Three distinct modes of intron dynamics in the evolution of eukaryotes. Genome Res. 17, 1034–1044 (2007).
Jaillon, O. et al. Translational control of intron splicing in eukaryotes. Nature 451, 359–362 (2008).
Kerenyi, Z. et al. Inter-kingdom conservation of mechanism of nonsense-mediated mRNA decay. EMBO J. 27, 1585–1595 (2008).
Plass, M., Agirre, E., Reyes, D., Camara, F. & Eyras, E. Co-evolution of the branch site and SR proteins in eukaryotes. Trends Genet. 24, 590–594 (2008).
Gal-Mark, N., Schwartz, S., Ram, O., Eyras, E. & Ast, G. The pivotal roles of TIA proteins in 5′ splice-site selection of Alu exons and across evolution. PLoS Genet. 5, e1000717 (2009).
Irimia, M., Rukov, J. L., Penny, D. & Roy, S. W. Functional and evolutionary analysis of alternatively spliced genes is consistent with an early eukaryotic origin of alternative splicing. BMC Evol. Biol. 7, 188 (2007).
Lev-Maor, G., Sorek, R., Shomron, N. & Ast, G. The birth of an alternatively spliced exon: 3′ splice-site selection in Alu exons. Science 300, 1288–1291 (2003).
Nurtdinov, R. N., Artamonova, I. I., Mironov, A. A. & Gelfand, M. S. Low conservation of alternative splicing patterns in the human and mouse genomes. Hum. Mol. Genet. 12, 1313–1320 (2003).
Thanaraj, T. A., Clark, F. & Muilu, J. Conservation of human alternative splice events in mouse. Nucleic Acids Res. 31, 2544–2552 (2003).
Kondrashov, F. A. & Koonin, E. V. Evolution of alternative splicing: deletions, insertions and origin of functional parts of proteins from intron sequences. Trends Genet. 19, 115–119 (2003).
Gilbert, W. Why genes in pieces? Nature 271, 501 (1978).
Kondrashov, F. A. & Koonin, E. V. Origin of alternative splicing by tandem exon duplication. Hum. Mol. Genet. 10, 2661–2669 (2001).
Doolittle, R. F. The multiplicity of domains in proteins. Annu. Rev. Biochem. 64, 287–314 (1995).
Kolkman, J. A. & Stemmer, W. P. Directed evolution of proteins by exon shuffling. Nature Biotech. 19, 423–428 (2001).
Liu, M. & Grigoriev, A. Protein domains correlate strongly with exons in multiple eukaryotic genomes — evidence of exon shuffling? Trends Genet. 20, 399–403 (2004). The authors found a strong correlation between borders of exons and protein domains in multiple eukaryotic genomes. This is consistent with the principles of exon shuffling.
Peng, T. & Li, Y. Tandem exon duplication tends to propagate rather than to create de novo alternative splicing. Biochem. Biophys. Res. Commun. 383, 163–166 (2009).
Letunic, I., Copley, R. R. & Bork, P. Common exon duplication in animals and its role in alternative splicing. Hum. Mol. Genet. 11, 1561–1567 (2002).
Gal-Mark, N., Schwartz, S. & Ast, G. Alternative splicing of Alu exons — two arms are better than one. Nucleic Acids Res. 36, 2012–2023 (2008).
Long, M., Rosenberg, C. & Gilbert, W. Intron phase correlations and the evolution of the intron/exon structure of genes. Proc. Natl Acad. Sci. USA 92, 12495–12499 (1995).
Patthy, L. Genome evolution and the evolution of exon-shuffling — a review. Gene 238, 103–114 (1999).
Patthy, L. Intron-dependent evolution: preferred types of exons and introns. FEBS Lett. 214, 1–7 (1987).
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000).
De Grassi, A. & Ciccarelli, F. D. Tandem repeats modify the structure of human genes hosted in segmental duplications. Genome Biol. 10, R137 (2009).
Parma, J., Christophe, D., Pohl, V. & Vassart, G. Structural organization of the 5′ region of the thyroglobulin gene. Evidence for intron loss and 'exonization' during evolution. J. Mol. Biol. 196, 769–779 (1987).
Makalowski, W., Mitchell, G. A. & Labuda, D. Alu sequences in the coding regions of mRNA: a source of protein variability. Trends Genet. 10, 188–193 (1994).
Sorek, R., Ast, G. & Graur, D. Alu-containing exons are alternatively spliced. Genome Res. 12, 1060–1067 (2002).
Nekrutenko, A. & Li, W. H. Transposable elements are found in a large number of human protein-coding genes. Trends Genet. 17, 619–621 (2001).
Wang, W. & Kirkness, E. F. Short interspersed elements (SINEs) are a major source of canine genomic diversity. Genome Res. 15, 1798–1808 (2005).
Wang, W. et al. Origin and evolution of new exons in rodents. Genome Res. 15, 1258–1264 (2005).
Kandul, N. P. & Noor, M. A. Large introns in relation to alternative splicing and gene evolution: a case study of Drosophila bruno-3. BMC Genet. 10, 67 (2009).
Kent, L. B. & Robertson, H. M. Evolution of the sugar receptors in insects. BMC Evol. Biol. 9, 41 (2009).
Fu, Y. et al. Alternative splicing of anciently exonized 5S rRNA regulates plant transcription factor TFIIIA. Genome Res. 19, 913–921 (2009).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Amit, M. et al. Biased exonization of transposed elements in duplicated genes: a lesson from the TIF-IA gene. BMC Mol. Biol. 8, 109 (2007).
Mersch, B., Sela, N., Ast, G., Suhai, S. & Hotz-Wagenblatt, A. SERpredict: detection of tissue- or tumor-specific isoforms generated through exonization of transposable elements. BMC Genet. 8, 78 (2007).
Lin, L. et al. Diverse splicing patterns of exonized Alu elements in human tissues. PLoS Genet. 4, e1000225 (2008).
Toll-Riera, M. et al. Origin of primate orphan genes: a comparative genomics approach. Mol. Biol. Evol. 26, 603–612 (2009).
Sorek, R. The birth of new exons: mechanisms and evolutionary consequences. RNA 13, 1603–1608 (2007).
Singer, S. S., Mannel, D. N., Hehlgans, T., Brosius, J. & Schmitz, J. From 'junk' to gene: curriculum vitae of a primate receptor isoform gene. J. Mol. Biol. 341, 883–886 (2004).
Krull, M., Brosius, J. & Schmitz, J. Alu-SINE exonization:en route to protein-coding function. Mol. Biol. Evol. 22, 1702–1711 (2005).
Kreahling, J. & Graveley, B. R. The origins and implications of Aluternative splicing. Trends Genet. 20, 1–4 (2004).
Schwartz, S. et al. Alu exonization events reveal features required for precise recognition of exons by the splicing machinery. PLoS Comput. Biol. 5, e1000300 (2009). This article describes the features that are required for precise recognition of exons by the splicing machinery by analysing Alu exonization events.
Sorek, R. et al. Minimal conditions for exonization of intronic sequences: 5′ splice site formation in Alu exons. Mol. Cell 14, 221–231 (2004).
Ram, O., Schwartz, S. & Ast, G. Multifactorial interplay controls the splicing profile of Alu-derived exons. Mol. Cell. Biol. 28, 3513–3525 (2008).
Corvelo, A. & Eyras, E. Exon creation and establishment in human genes. Genome Biol. 9, R141 (2008). The authors show that specific sequence environments are required for exonization and that these can change with time.
Gombart, A. F., Saito, T. & Koeffler, H. P. Exaptation of an ancient Alu short interspersed element provides a highly conserved vitamin D-mediated innate immune response in humans and primates. BMC Genomics 10, 321 (2009).
Lee, J. R. et al. Lineage specific evolutionary events on SFTPB gene: Alu recombination-mediated deletion (ARMD), exonization, and alternative splicing events. Gene 435, 29–35 (2009).
Pan, Q. et al. Alternative splicing of conserved exons is frequently species-specific in human and mouse. Trends Genet. 21, 73–77 (2005).
Lev-Maor, G. et al. The 'alternative' choice of constitutive exons throughout evolution. PLoS Genet. 3, e203 (2007). This paper shows that exons shift from constitutive to alternative splicing during evolution, and relaxation of the 5′ splice site sequence is one of the molecular mechanisms that leads to this shift.
Ke, S., Zhang, X. H. & Chasin, L. A. Positive selection acting on splicing motifs reflects compensatory evolution. Genome Res. 18, 533–543 (2008).
Lev-Maor, G. et al. Intronic Alus influence alternative splicing. PLoS Genet. 4, e1000204 (2008). This article shows that Alu insertions into introns change the mode of splicing of the flanking exons.
Tappino, B., Regis, S., Corsolini, F. & Filocamo, M. An Alu insertion in compound heterozygosity with a microduplication in GNPTAB gene underlies Mucolipidosis II. Mol. Genet. Metab. 93, 129–133 (2008).
Mola, G., Vela, E., Fernandez-Figueras, M. T., Isamat, M. & Munoz-Marmol, A. M. Exonization of Alu-generated splice variants in the survivin gene of human and non-human primates. J. Mol. Biol. 366, 1055–1063 (2007).
Sorek, R., Shamir, R. & Ast, G. How prevalent is functional alternative splicing in the human genome? Trends Genet. 20, 68–71 (2004).
Yeo, G. W., Van Nostrand, E., Holste, D., Poggio, T. & Burge, C. B. Identification and analysis of alternative splicing events conserved in human and mouse. Proc. Natl Acad. Sci. USA 102, 2850–2855 (2005).
Modrek, B. & Lee, C. J. Alternative splicing in the human, mouse and rat genomes is associated with an increased frequency of exon creation and/or loss. Nature Genet. 34, 177–180 (2003).
Magen, A. & Ast, G. The importance of being divisible by three in alternative splicing. Nucleic Acids Res. 33, 5574–5582 (2005).
Irimia, M. et al. Widespread evolutionary conservation of alternatively spliced exons in Caenorhabditis. Mol. Biol. Evol. 25, 375–382 (2008).
Melamud, E. & Moult, J. Stochastic noise in splicing machinery. Nucleic Acids Res. 37, 4873–4886 (2009).
Xing, Y. & Lee, C. Alternative splicing and RNA selection pressure — evolutionary consequences for eukaryotic genomes. Nature Rev. Genet. 7, 499–509 (2006).
Ermakova, E. O., Nurtdinov, R. N. & Gelfand, M. S. Fast rate of evolution in alternatively spliced coding regions of mammalian genes. BMC Genomics 7, 84 (2006).
Wang, Z. & Burge, C. B. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA 14, 802–813 (2008).
Plass, M. & Eyras, E. Differentiated evolutionary rates in alternative exons and the implications for splicing regulation. BMC Evol. Biol. 6, 50 (2006).
Sakabe, N. J. & de Souza, S. J. Sequence features responsible for intron retention in human. BMC Genomics 8, 59 (2007).
Tilgner, H. et al. Nucleosome positioning as a determinant of exon recognition. Nature Struct. Mol. Biol. 16, 996–1001 (2009). The authors found stronger nucleosome occupancy in exons than in exons with weak splice sites and in pseudoexons.
Andersson, R., Enroth, S., Rada-Iglesias, A., Wadelius, C. & Komorowski, J. Nucleosomes are well positioned in exons and carry characteristic histone modifications. Genome Res. 19, 1732–1741 (2009). The authors found higher nucleosome occupancy in exons. The exons were enriched with specific histone modifications.
Hodges, C., Bintu, L., Lubkowska, L., Kashlev, M. & Bustamante, C. Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II. Science 325, 626–628 (2009).
Kolasinska-Zwierz, P. et al. Differential chromatin marking of introns and expressed exons by H3K36me3. Nature Genet. 41, 376–381 (2009).
Luco, R. F. et al. Regulation of alternative splicing by histone modifications. Science 327, 996–1000 (2010). The authors show the first direct link between histone modification and AS: the modulation of AS resulted in splice-site switching.
Nahkuri, S., Taft, R. J. & Mattick, J. S. Nucleosomes are preferentially positioned at exons in somatic and sperm cells. Cell Cycle 8, 3420–3424 (2009).
Lavelle, C. & Prunell, A. Chromatin polymorphism and the nucleosome superfamily: a genealogy. Cell Cycle 6, 2113–2119 (2007).
Sivolob, A. & Prunell, A. Nucleosome conformational flexibility and implications for chromatin dynamics. Philos. Transact. A Math. Phys. Eng. Sci. 362, 1519–1547 (2004).
Alilat, M., Sivolob, A., Revet, B. & Prunell, A. Nucleosome dynamics. Protein and DNA contributions in the chiral transition of the tetrasome, the histone (H3–H4)2 tetramer–DNA particle. J. Mol. Biol. 291, 815–841 (1999).
Kaplan, C. D. Revealing the hidden relationship between nucleosomes and splicing. Cell Cycle 8, 3633–3634 (2009).
Garcia-Blanco, M. A., Baraniak, A. P. & Lasda, E. L. Alternative splicing in disease and therapy. Nature Biotech. 22, 535–546 (2004).
Wood, M., Yin, H. & McClorey, G. Modulating the expression of disease genes with RNA-based therapy. PLoS Genet. 3, e109 (2007).
Sugnet, C. W., Kent, W. J., Ares, M. Jr & Haussler, D. Transcriptome and genome conservation of alternative splicing events in humans and mice. Pac. Symp. Biocomput. 9, 66–77 (2004).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Labrador, M. & Corces, V. G. Extensive exon reshuffling over evolutionary time coupled to trans-splicing in Drosophila. Genome Res. 13, 2220–2228 (2003).
van Rijk, A. & Bloemendal, H. Molecular mechanisms of exon shuffling: illegitimate recombination. Genetica 118, 245–249 (2003).
Babushok, D. V., Ostertag, E. M. & Kazazian, H. H. Jr. Current topics in genome evolution: molecular mechanisms of new gene formation. Cell. Mol. Life Sci. 64, 542–554 (2007).
Patthy, L. Exon shuffling and other ways of module exchange. Matrix Biol. 15, 301–310; discussion 311–312 (1996).
van Rijk, A. A., de Jong, W. W. & Bloemendal, H. Exon shuffling mimicked in cell culture. Proc. Natl Acad. Sci. USA 96, 8074–8079 (1999).
Bass, B. L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2, e391 (2004).
Lev-Maor, G. et al. RNA-editing-mediated exon evolution. Genome Biol. 8, R29 (2007).
Moller-Krull, M., Zemann, A., Roos, C., Brosius, J. & Schmitz, J. Beyond DNA: RNA editing and steps toward Alu exonization in primates. J. Mol. Biol. 382, 601–609 (2008).
Gommans, W. M., Mullen, S. P. & Maas, S. RNA editing: a driving force for adaptive evolution? Bioessays 31, 1137–1145 (2009).
Acknowledgements
We thank D. Hollander for preparing the figures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Spliceosome
-
A ribonucleoprotein complex that is involved in splicing of nuclear precursor mRNA (pre-mRNA). It is composed of five small nuclear ribonucleoproteins (snRNPs) and more than 50 non-snRNPs, which recognize and assemble on exon–intron boundaries to catalyse intron processing of the pre-mRNA.
- Nucleosome
-
The basic unit of chromatin, containing ∼147 bp of DNA wrapped around a histone octamer (which is composed of two copies each of histone 3 (H3), H4, H2A and H2B).
- Basal splicing
-
A conserved mRNA splicing mechanism. It is composed of the splicing signals and the core of the machinery is formed by five spliceosomal small nuclear ribonucleoproteins and an unknown number of proteins.
- Alu
-
An interspersed DNA sequence of 300 bp that belongs to the short interspersed element (SINE) family and is found in the genome of primates. Alu elements are composed of a head-to-tail dimer in which the first monomer is 140 bp long and the second is 170 bp long. In humans, there are ∼1.1 million copies of Alu elements, of which ∼500,000 copies are located in introns.
- Retroelement
-
A mobile genetic element. Its DNA is transcribed into RNA, which is reverse-transcribed into DNA and then inserted into a new location in the genome.
- Serine/arginine proteins
-
A group of highly conserved serine- and arginine-rich splicing regulatory proteins in metazoans.
- Heterogenous nuclear ribonucleoproteins
-
A large set of proteins that bind the precursor mRNA and regulate splicing.
- Nonsense-mediated decay
-
The process by which the cell destroys mRNAs that are untranslatable due to the presence of a premature stop codon in the coding region.
- Excavates
-
A major kingdom of unicellular eukaryotes, often known as Excavata. The phylogenetic category Excavata contains a variety of free-living and symbiotic forms, and also includes some important parasites of humans.
- Chromalveolates
-
A hypothetical 'supergroup' of protists, including apicomplexa, dinoflagellates, ciliates, heterokonts, haptophytes and cryptomonads, all of which are suggested to have diverged from an ancient common ancestor that acquired a plastid by secondary endosymbiosis with a red alga.
- Substitution alternative splicing
-
An alternative splicing pattern in which one of two amino acid sequences is included in the protein.
- Mutually exclusive alternative splicing
-
Only one of a set of two or more exons in a gene is included in the final transcript.
- Modular proteins
-
Proteins created by intronic recombination. According to the exon shuffling theory, each exon encodes a single protein domain (a 'module'), and the process of shuffling creates a new chimeric protein from the combination of domains (or 'modules').
- Transposable elements
-
Segments of genetic material that are capable of changing their location in the genome of an organism.
- Orphan genes
-
Genes that do not share any homology with genes from other species.
- Splicing regulatory elements
-
Specific cis-acting RNA sequence elements that are present in introns or in exons. They are bound by trans-acting splicing regulatory proteins (repressors and activators), which regulate alternative splicing.
- Purifying selection
-
Selection against deleterious alleles that arise in a population, preventing their increase in frequency and assuring their eventual disappearance from the gene pool.
- Pseudoexons
-
Precursor mRNA sequences that resemble exons — both in their size and in the presence of flanking splice-site sequences — but that are not normally recognized by the splicing machinery.
Rights and permissions
About this article
Cite this article
Keren, H., Lev-Maor, G. & Ast, G. Alternative splicing and evolution: diversification, exon definition and function. Nat Rev Genet 11, 345–355 (2010). https://doi.org/10.1038/nrg2776
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2776
This article is cited by
-
The physiology of alternative splicing
Nature Reviews Molecular Cell Biology (2023)
-
A presumed missense variant in the U2AF2 gene causes exon skipping in neurodevelopmental diseases
Journal of Human Genetics (2023)
-
The role of alternative splicing in lung cancer
Cancer Chemotherapy and Pharmacology (2023)
-
A comparison of mRNA sequencing (RNA-Seq) library preparation methods for transcriptome analysis
BMC Genomics (2022)
-
ESRP1-regulated isoform switching of LRRFIP2 determines metastasis of gastric cancer
Nature Communications (2022)