Abstract
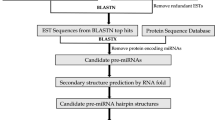
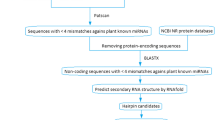
MicroRNAs are small, non-protein-coding RNAs playing regulatory functions in many organisms. Using computational approaches 26 new Brachypodium distachyon miRNAs belonging to 19 miRNA families were identified in expressed sequence tags (EST) and genomic survey sequence databases. EST revealed that predicted miRNAs are expressed in B. distachyon. Detailed nucleotide analyses showed that pre-miRNAs in B. distachyon are in the range of 63–180 nucleotides. Mature miRNAs located in the different positions of precursor RNAs are varied from 19 to 24 nucleotides in length. Quantifying RNAs using realtime PCR (qRT-PCR) analyses validated expression level differences of selected B. distachyon miRNAs. In this study, we detected that the expression level of some of the predicted miRNAs are distinct and some of them are similar in the leaf tissues. In addition, using these miRNAs as queries 27 potential target mRNAs were predicted in B. distachyon NCBI EST database and 246 target mRNA were predicted in NCBI protein-coding nucleotide (mRNA) database of all plant species. The majority of the target mRNAs encode transcription factors regulating plant development, morphology and flowering time. Other newly identified miRNAs target the mRNAs involving metabolic processes, signal transduction and stress response.






Similar content being viewed by others
Abbreviations
- ΔG:
-
Folding-free energies
- AGO1:
-
Argonaute-1
- Ap2:
-
APETALA2
- ARF:
-
Auxin response transcription factor
- DCL1:
-
Dicer-like protein
- GSS:
-
Genomic survey sequence
- miRNA:
-
MicroRNA
- RT:
-
Real time
- EST:
-
Expressed sequence tag
- DCL1:
-
Dicer-like protein
- EREBPs:
-
Ethylene-responsive element-binding proteins
- MFE:
-
Minimal folding-free energy
- MFEI:
-
Minimal folding-free energy index
- miRNA*:
-
Opposite miRNA sequence
- NCBI:
-
National Center of Biotechnology Information
- Nt:
-
Nucleotide(s)
- pre-miRNA:
-
MicroRNA precursor
- pri-miRNAs:
-
MicroRNA primary
- RISC:
-
RNA-induced silencing complex
- qRT-PCR:
-
Quantitative real-time PCR
- SBP:
-
Squamosa promoter-binding proteins
- Ser:
-
Serine
- SPL:
-
SBP-like proteins
- Thr:
-
Threonine
References
Achard P, Herr A, Baulcombe DC, Harberd NP (2004) Modulation of floral development by a gibberellin-regulated microRNA. Development 131:3357–3365
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Bao N, Lye KW, Barton MK (2004) MicroRNA binding sites in Arabidopsis class IIIHD-ZIP mRNAs are required for methylation of the template chromosome. Dev Cell 7:653–662
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bonnet E, Wuyts J, Rouze P, Van de Peer Y (2004) Detection of 91 potential in plant conserved plant microRNAs in Arabidopsis thaliana and Oryza sativa identifies important target genes. Proc Natl Acad Sci USA 101:11511–11516
Cai X, Ballif J, Endo S, Davis E, Liang M, Chen D, DeWald D, Kreps J, Zhu T, Wu Y (2007) A putative CCAAT-binding transcription factor is a regulator of flowering timing in Arabidopsis. Plant Physiol 145(1):98–105
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301:336–338
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Chen J, Li WX, Xie D, Peng JR, Ding SW (2004) Viral virulence protein suppresses RNA silencing-mediated defense but upregulates the role of microRNA in host gene expression. Plant Cell 16:1302–1313
Ergen ZN, Dinler G, Shearman R, Budak H (2007) Identifying, cloning and structural analysis of differentially expressed genes upon Puccinia infection of Festuca rubra var. rubra. Gene 393:145–152
Goetz M, Hooper LC, Johnson SD, Rodrigues JC, Vivian-Smith A, Koltunow AM (2007) Expression of aberrant forms of AUXIN RESPONSE FACTOR8 stimulates parthenocarpy in Arabidopsis and tomato. Plant Physiol 145(2):351–366
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Guo Q, Xiang AL, Yang Q, Yang ZM (2007) Bioinformatic identification of microRNAs and their target genes from Solanum tuberosum expressed sequence tags. Chin Sci Bull 52:2380–2389
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Jung JH, Park CM (2007) MIR166/165 genes exhibit dynamic expression patterns in regulating shoot apical meristem and floral development in Arabidopsis. Planta 225(6):1327–1338
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA unction. Dev Cell 4:205–217
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim VN, Chua NH, Park CM (2005) microRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42:84–94
Laufs P, Peaucelle A, Morin H, Traas J (2004) MicroRNA regulation of theCUC genes is required for boundary size control in Arabidopsis meristems. Development 131:4311–4322
Lauter N, Kampani A, Carlson S, Goebel M, Moose SP (2005) microRNA172 down-regulates glossy15 to promote vegetative phase change in maize. Proc Natl Acad Sci USA 102:9412–9417
Li Y, Li W, Jin YX (2005) Computational identification of novel family members of microRNA genes in Arabidopsis thaliana and Oryza sativa. Acta Biochim Biophys Sin (Shanghai) 37:75–87
Llave C, Xie ZX, Kasschau KD, Carrington JC (2002) Cleavage of scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 297:2053–2056
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14:1035–1046
Millar AA, Gubler F (2005) The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 17:705–721
Mlotshwa S, Yang Z, Kim Y, Chen X (2006) Floral patterning defects induced by Arabidopsis APETALA2 and microRNA172 expression in Nicotiana benthamiana. Plant Mol Biol 61:781–793
Ochando I, Jover-Gil S, Ripoll JJ, Candela H, Vera A, Ponce MR, Martínez-Laborda A, Micol JL (2006) Mutations in the microRNA complementarity site of the INCURVATA4 gene perturb meristem function and adaxialize lateral organs in Arabidopsis. Plant Physiol 14:607–619
Ozdemir BS, Hernandez P, Filiz E, Budak H (2008) Brachypodium genomics. Int J Plant Genomics 2008:536104
Palatnik JF, Allen E, Wu X et al (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263
Park W, Li JJ, Song RT, Messing J, Chen XM (2002) CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 12:1484–1495
Prigge MJ, Otsuga D, Alonso JM, Ecker JR, Drews GN, Clark SE (2005) Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17:61–76
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP (2002) MicroRNAs in plants. Genes Dev 16:1616–1626
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110:513–520
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Dev Cell 8:517–527
Schwarz S, Grande AV, Bujdoso N, Saedler H, Huijser P (2008) The microRNA regulated SBP-box genes SPL9 and SPL15 control shoot maturation in Arabidopsis. Plant Mol Biol 67(1–2):183–195
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP (2007) Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 3:1–12
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17:2204–2216
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHDZIP target genes. Development 132:3657–3668
Wu G, Poethig RS (2006) Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 133:3539–3547
Xie FL, Huang SQ, Guo K, Xiang AL, Zhu YY, Nie L, Yang ZM (2007) Computational identification of novel microRNAs and targets in Brassica napus. FEBS Lett 581:1464–1474
Yin Z, Li C, Han X, Shen F (2008) Identification of conserved microRNAs and their target genes in tomato (Lycopersicon esculentum). Gene 414:60–66
Zhang Y (2005) miRU: an automated plant miRNA target prediction server. Nucleic Acids Res 33:W701–W704
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2005) Identification and characterization of new plant microRNAs using EST analysis. Cell Res 15:336–360
Zhang BH, Pan XP, Anderson TA (2006a) Identification of 188 conserved maize microRNAs and their targets. FEBS Lett 580:3753–3762
Zhang BH, Pan XP, Cobb GP, Anderson TA (2006b) Plant microRNA: a small regulatory molecule with big impact. Dev Biol 289:3–16
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2006c) Computational identification of microRNAs and their targets. Comput Biol Chem 30:395–407
Zhang BH, Pan XP, Cannon CH, Cobb GP, Anderson TA (2006d) Conservation and divergence of plant microRNA genes. Plant J 46:243–259
Zhang BH, Wang QL, Pan XP (2007a) MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol 210:279–289
Zhang BH, Wang QL, Wang K, Pan X, Liu F, Gou T, Cobb GP, Anderson TA (2007b) Identification of cotton microRNAs and their targets. Gene 397:26–37
Zhang B, Pan X, Stellwag EJ (2008) Identification of soybean microRNAs and their targets. Planta 229:161–182
Zhang W, Luo Y, Gong X, Zeng W, Li S (2009) Computational identification of 48 potato microRNAs and their targets. Comput Biol Chem 33:84–93
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Acknowledgment
We would like to thank Mine Bakar for technical assistance in performing qRT-PCR experiments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Unver, T., Budak, H. Conserved microRNAs and their targets in model grass species Brachypodium distachyon . Planta 230, 659–669 (2009). https://doi.org/10.1007/s00425-009-0974-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-0974-7