Abstract
Background and Purpose
Nuclear grades of clear cell renal cell carcinoma (ccRCC) are usually confirmed by invasive methods. Radiomics is a quantitative tool that uses non-invasive medical imaging for tumor diagnosis and prognosis. In this study, a radiomics approach was proposed to analyze the association between preoperative computed tomography (CT) images and nuclear grades of ccRCC.
Methods
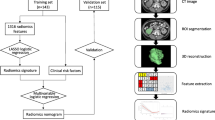
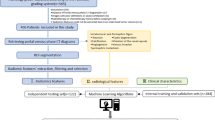
Our dataset included 320 ccRCC patients from two centers and was divided into a training set (n = 124), an internal test set (n = 123), and an external test set (n = 73). A radiomic feature set was extracted from unenhanced, corticomedullary phase, and nephrographic phase CT images. The maximizing independent classification information criteria function and recursive feature elimination with cross-validation were used to select effective features. Random forests were used to build a final model for predicting nuclear grades, and area under the receiver operating characteristic curve (AUC) was used to evaluate the performance of radiomic features and models.
Results
The radiomic features from the three CT phases could effectively distinguished the four nuclear grades. A combined model, merging radiomic features and clinical characteristics, obtained good predictive performances in the internal test set (AUC 0.77, 0.75, 0.79, and 0.85 for the four grades, respectively), and performance was further confirmed in the external test set, with AUCs of 0.75, 0.68, and 0.73 (no fourth-level data).
Conclusion
The combination of CT radiomic features and clinical characteristics could discriminate the nuclear grades in ccRCC, which may help in assisting treatment decision making.





Similar content being viewed by others
References
Bhatt JR, Finelli A. Landmarks in the diagnosis and treatment of renal cell carcinoma. Nat Rev Urol. 2014;11:517–25.
Störkel S, Eble JN, Adlakha MD, et al. Classification of renal cell carcinoma. Cancer. 1997;80:987.
Rabjerg M. Identification and validation of novel prognostic markers in Renal Cell Carcinoma. Dan Med J. 2017;64:B5339.
Moch H, Cubilla AL, Humphrey PA, et al. The 2016 WHO classification of tumours of the urinary system and male genital organs—part A: renal, penile, and testicular tumours. Histopathology. 2016;46:93–105.
Patard JJ, Leray E, Rioux-Leclercq N, et al. Prognostic value of histologic subtypes in renal cell carcinoma: a multicenter experience. J Urol. 2006;175:2763–71.
Ljungberg B, Bensalah K, Canfield S, et al. EAU guidelines on renal cell carcinoma: 2014 update. Eur Urol. 2015;67:913–24.
Zhu YH, Wang X, Zhang J, et al. Low enhancement on multiphase contrast-enhanced CT images: an independent predictor of the presence of high tumor grade of clear cell renal cell carcinoma. AJR Am J Roentgenol. 2014;203:295–300.
Coy H, Douek M, Young J, et al. Differentiation of low grade from high grade clear cell renal cell carcinoma neoplasms using a CAD algorithm on four-phase CT. J Clin Oncol. 2016;34(15 Suppl):4564.
Leibovich BC, Blute ML, Cheville JC, et al. Prediction of progression after radical nephrectomy for patients with clear cell renal cell carcinoma: a stratification tool for prospective clinical trials. Cancer. 2003;97:1663–71.
Erdoğan F, et al. Prognostic significance of morphologic parameters in renal cell carcinoma. Am J Surg Pathol. 1982;58:655–63.
Bretheau D, Lechevallier E, De FM, et al. Prognostic value of nuclear grade of renal cell carcinoma. Cancer. 1995;76:2543.
Sekar RR, Patil D, Pearl J, et al. The relationship between preoperative c-reactive protein and Fuhrman nuclear grade in stage T1 renal cell carcinoma. J Urol. 2016;195:e1033.
Lambin P, Rios-Velazquez E, Leijenaar R, et al. Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer. 2012;48:441–6.
Zhou H, Dong D, Chen B, et al. Diagnosis of distant metastasis of lung cancer: based on clinical and radiomic features. Transl Oncol. 2018;11:31–6.
Song J, Shi J, Dong D, et al. A new approach to predict progression-free survival in stage IV EGFR-mutant NSCLC patients with EGFR-TKI therapy. Clin Cancer Res. 2018;24(15):3583–92.
Dong D, Tang L, Li Z-Y, et al. Development and validation of an individualized nomogram to identify occult peritoneal metastasis in patients with advanced gastric cancer. Ann Oncol. 2019;30:431–8.
Dong D, Zhang F, Zhong L-Z, et al. Development and validation of a novel MR imaging predictor of response to induction chemotherapy in locoregionally advanced nasopharyngeal cancer: a randomized controlled trial substudy (NCT01245959). BMC Med. 2019;17(1):190.
Peng H, Dong D, Fang M, et al. Prognostic value of deep learning PET/CT-based radiomics: potential role for future individual induction chemotherapy in advanced nasopharyngeal carcinoma. Clin Cancer Res. 2019;25(14):4271–9.
Zhu X, Dong D, Chen Z, et al. Radiomic signature as a diagnostic factor for histologic subtype classification of non-small cell lung cancer. Eur Radiol. 2018;28(7):2772–8.
Yang L, Dong D, Fang M, et al. Can CT-based radiomics signature predict KRAS/NRAS/BRAF mutations in colorectal cancer? Eur Radiol. 2018;28(5):2058–67.
Wang S, Zhou M, Liu Z, et al. Central focused convolutional neural networks: developing a data-driven model for lung nodule segmentation. Med Image Anal. 2017;40:172–83.
Edge SB, Byrd DR, Compton CC, et al. American Joint Committee on Cancer (AJCC) cancer staging manual; 2010.
Rios VE, Parmar C, Liu Y, et al. Somatic mutations drive distinct imaging phenotypes in lung cancer. Cancer Res. 2017;77:3922.
Aerts HJWL, Velazquez ER, Leijenaar RTH, et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun. 2014;5:4006.
Lambin P, Rth L, Deist TM, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017;14:749.
Wang J, Wei JM, Yang Z, Wang SQ. Feature selection by maximizing independent classification information. IEEE Trans Knowl Data Eng. 2017;29:828–41.
Granitto PM, Furlanello C, Biasioli F, Gasperi F. Recursive feature elimination with random forest for PTR-MS analysis of agroindustrial products. Chemom Intell Lab Syst. 2006;83:83–90.
Buitinck L, Louppe G, Blondel M, et al. API design for machine learning software: experiences from the scikit-learn project. Eprint Arxiv; 2013.
Pedregosa F, Gramfort A, Michel V, et al. Scikit-learn: machine learning in python. J Mach Learn Res. 2016;12:2825–30.
Ding J, Xing Z, Jiang Z, et al. CT-based radiomic model predicts high grade of clear cell renal cell carcinoma. Eur J Radiol. 2018;103:51–6.
Acknowledgement
The authors would like to acknowledge the instrumental and technical support of the Multi-modal biomedical imaging experimental platform, Institute of Automation, Chinese Academy of Sciences.
Funding
This work was supported by the National Key R&D Program of China (2017YFC1308700, 2017YFA0205200, 2017YFC1309100, 2016YFC0103803, 2016YFC0103001), National Natural Science Foundation of China (91959130, 81971776, 81771924, 812716298, 81271629, 81227901), Beijing Natural Science Foundation (L182061), Wuxi Medical Innovation Team Program (CXTD002), Bureau of International Cooperation of Chinese Academy of Sciences (173211KYSB20160053), and the Youth Innovation Promotion Association CAS (2017175).
Author information
Authors and Affiliations
Contributions
Research idea and study design: XF, HZ, JT. Data acquisition: HM, XL, MX, SY, JZ, RY. Data analysis/interpretation: HZ, DD, MF, DG, HZ, XF, JT. Supervision or mentorship: DD, HZ, XF, JT, CP, and XF.
Corresponding authors
Ethics declarations
Disclosure
Hongyu Zhou, Haixia Mao, Di Dong, Mengjie Fang, Dongsheng Gu, Xueling Liu, Min Xu, Shudong Yang, Jian Zou, Ruohan Yin, Hairong Zheng, Jie Tian, Changjie Pan, and Xiangming Fang have declared no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, H., Mao, H., Dong, D. et al. Development and External Validation of Radiomics Approach for Nuclear Grading in Clear Cell Renal Cell Carcinoma. Ann Surg Oncol 27, 4057–4065 (2020). https://doi.org/10.1245/s10434-020-08255-6
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-020-08255-6