Abstract—
The control region of mitochondrial DNA (mtDNA) in sturgeon contains one to seven tandem nucleotide repeats 78‒83 bp in size. Some sturgeon species are homoplasmic by the D-loop size (Acipenser nudiventris, A. oxyrinchus, A. sturio), some are mildly heteroplasmic (A. fulvescens, Huso huso) and some are markedly heteroplasmic (A. brevirostrum, A. medirostris, A. mikadoi, A. naccarii, and A. transmontanus). This work presents a comparison of the D-loop sequences associated with the termination of mtDNA replication in fish and the conservative sequences determining the termination of replication (TAS) in these organisms. It is proposed that the D-loop heteroplasmy in sturgeon may be associated with variation in the number of tandem repeat sequences, which can form stable spatial structures during mtDNA replication. In most sturgeon species with pronounced heteroplasmy, the energy levels required for the folding of tandem repeats containing variable number of repeated units differ minimally.





Similar content being viewed by others
REFERENCES
Smith D.R. 2016. The past, present and future of mitochondrial genomics: Have we sequenced enough mtDNAs? Brief Funct. Genomics. 15, 47‒54.
Anderson S., Bankier A.T., Barrell B.G., de Bruijn M.H., Coulson A.R., Drouin J., Eperon I.C., Nierlich D.P., Roe B.A., Sanger F., Schreier P.H., Smith A.J., Staden R., Young I.G. 1981. Sequence and organization of the human mitochondrial genome. Nature. 290, 457–465.
Kasamatsu H., Robberson D.L., Vinograd J. 1971. A novel closed-circular mitochondrial DNA with properties of a replicating intermediate. Proc. Natl. Acad. Sci. U. S. A. 68, 2252–2257.
Kaguni L.S. 2004. DNA polymerase γ, the mitochondrial replicase. Annu. Rev. Biochem. 73, 293–320.
Jemt E., Persson Ö., Shi Y., Mehmedovic M., Uhler J.P., Dávila López M., Freyer C., Gustafsson C.M., Samuelsson T., Falkenberg M. 2015. Regulation of DNA replication at the end of the mitochondrial D-loop involves the helicase TWINKLE and a conserved sequence element. Nucleic Acids Res. 43, 9262–9275.
Shadel G.S., Clayton D.A. 1997. Mitochondrial DNA maintenance in vertebrates. Annu. Rev. Biochem. 66, 409–435.
Nicholls T.J, Minczuk M. 2014. In D-loop: 40 years of mitochondrial 7S DNA. Exp. Gerontol. 56, 175–181.
Buroker N.E., Brown J.R., Gilbert T.A., O’Hara P.J., Beckenbach A.T., Thomas W.K., Smith M.J. 1990. Length heteroplasmy of sturgeon mitochondrial DNA: An illegitimate elongation model. Genetics. 124, 157–163.
Brown J.R., Beckenbach K., Beckenbach A.T., Smith M.J. 1996. Length variation, heteroplasmy and sequence divergence in the mitochondrial DNA of four species of sturgeon (Acipenser). Genetics. 142, 525–535.
Ludwig A., May B., Debus L., Jenneckens I. 2000. Heteroplasmy in the mtDNA control region of sturgeon (Acipenser, Huso and Scaphirhynchus). Genetics. 156, 1933–1947.
Larkin M.A., Blackshields G., Brown N.P., Chenna R., McGettigan P.A., McWilliam H., Valentin F., Wallace I.M., Wilm A., Lopez R., Thompson J.D., Gibson T.J., Higgins D.G. 2007. Clustal W and Clustal X version 2.0. Bioinformatics. 23, 2947–2948.
Okonechnikov K., Golosova O., Fursov M., the UGENE team. 2012. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics. 28, 1166–1167.
Benson G. 1999. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580.
Zuker M. 1989. On finding all suboptimal foldings of an RNA molecule. Science. 244, 48–52.
Zuker M. 2003. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415.
Dettlaff T.A., Ginsburg A.S., Schmalhausen O.I. 1993. Sturgeon Fishes: Developmental Biology and Aquaculture. Berlin: Springer-Verlag, 200–203.
Doroshov S.I., Moberg G.P., Van Eenennaam J.P. 1997. Observations on the reproductive cycle of cultured white sturgeon, Acipenser transmontanus. Environ. Biol. Fishes. 48, 265–278.
Benjamini Y., Hochberg Y. 1995. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. Roy. Statist. Soc. Ser. B (Methodological). 57, 289–300.
Tzur S., Rosset S. 2017. Strictly conserved tri-nucleotide motif “CAT” is associated with TAS DNA protein-binding sites in human mitochondrial DNA control region. Mitochondr. DNA. 28 (2), 250–253.
Vodolazhskii D.I., Kornienko I.V., Voinova N.V. 2008. Hypervariability of the D-loop region in mitochondrial DNA of Russian sturgeon Acipenser gueldenstaedtii (Acipenseriformes, Acipenseridae). J. Ichthyol. 48 (2), 188–197.
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated by D. Timchenko
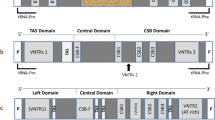
Abbreviations: OL, origin of L-strand replication; OH, origin of H-strand replication; TAS, termination-associated sequence of mtDNA, VNTR, variable number tandem repeat.
Rights and permissions
About this article
Cite this article
Kornienko, I.V., Chebotarev, D.A., Makhotkin, M.A. et al. Termination of Replication and Mechanisms of Heteroplasmy in Sturgeon Mitochondrial DNA. Mol Biol 53, 107–117 (2019). https://doi.org/10.1134/S0026893319010060
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893319010060