Abstract
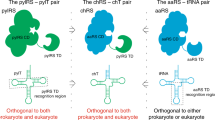
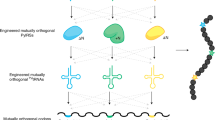
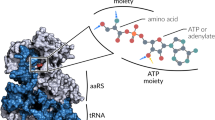
The archaeal/eukaryotic tyrosyl-tRNA synthetase (TyrRS)–tRNATyr pairs do not cross-react with their bacterial counterparts. This 'orthogonal' condition is essential for using the archaeal pair to expand the bacterial genetic code. In this study, the structure of the Methanococcus jannaschii TyrRS–tRNATyr–L-tyrosine complex, solved at a resolution of 1.95 Å, reveals that this archaeal TyrRS strictly recognizes the C1-G72 base pair, whereas the bacterial TyrRS recognizes the G1-C72 in a different manner using different residues. These diverse tRNA recognition modes form the basis for the orthogonality. The common tRNATyr identity determinants (the discriminator, A73 and the anticodon residues) are also recognized in manners different from those of the bacterial TyrRS. Based on this finding, we created a mutant TyrRS that aminoacylates the amber suppressor tRNA with C34 65 times more efficiently than does the wild-type enzyme.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Koide, H. et al. Biosynthesis of a protein containing a nonprotein amino acid by Escherichia coli: L-2-aminohexanoic acid at position 21 in human epidermal growth factor. Proc. Natl. Acad. Sci. USA 85, 6237–6241 (1988).
Nowak, M.W. et al. Nicotinic receptor binding site probed with unnatural amino acid incorporation in intact cells. Science 268, 439–442 (1995).
Short, G.F. III et al. Probing the S1/S1′ substrate binding pocket geometry of HIV-1 protease with modified aspartic acid analogues. Biochemistry 39, 8768–8781 (2000).
Chin, J.W. et al. Addition of p-azido-L-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Chin, J.W., Martin, A.B., King, D.S., Wang, L. & Schultz, P.G. Addition of a photocrosslinking amino acid to the genetic code of Escherichia coli. Proc. Natl. Acad. Sci. USA 99, 11020–11024 (2002).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
Yabuki, T. et al. Dual amino acid-selective and site-directed stable-isotope labeling of the human c-Ha-Ras protein by cell-free synthesis. J. Biomol. NMR 11, 295–306 (1998).
Lu, W., Gong, D., Bar-Sagi, D. & Cole, P.A. Site-specific incorporation of a phosphotyrosine mimetic reveals a role for tyrosine phosphorylation of SHP-2 in cell signaling. Mol. Cell 8, 759–769 (2001).
Wang, L. & Schultz, P.G. Expanding the genetic code. Chem. Commun. (Camb.) 1, 1–11 (2002).
Furter, R. Expansion of the genetic code: site-directed p-fluoro-phenylalanine incorporation in Escherichia coli. Protein Sci. 7, 419–426 (1998).
Wang, L., Brock, A., Herberich, B. & Schultz, P.G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Wang, L., Brock, A. & Schultz, P.G. Adding L-3-(2-naphthyl)alanine to the genetic code of E. coli. J. Am. Chem. Soc. 124, 1836–1837 (2002).
Santoro, S.W., Wang, L., Herberich, B., King, D.S. & Schultz, P.G. An efficient system for the evolution of aminoacyl-tRNA synthetase specificity. Nat. Biotechnol. 20, 1044–1048 (2002).
Wang, L., Zhang, Z., Brock, A. & Schultz, P.G. Addition of the keto functional group to the genetic code of Escherichia coli. Proc. Natl. Acad. Sci. USA 100, 56–61 (2003).
Kiga, D. et al. An engineered Escherichia coli tyrosyl-tRNA synthetase for site-specific incorporation of an unnatural amino acid into proteins in eukaryotic translation and its application in a wheat germ cell-free system. Proc. Natl. Acad. Sci. USA 99, 9715–9720 (2002).
Sakamoto, K. et al. Site-specific incorporation of an unnatural amino acid into proteins in mammalian cells. Nucleic Acids Res. 30, 4692–4699 (2002).
Chow, C.M. & RajBhandary, U.L. Saccharomyces cerevisiae cytoplasmic tyrosyl-tRNA synthetase gene. Isolation by complementation of a mutant Escherichia coli suppressor tRNA defective in aminoacylation and sequence analysis. J. Biol. Chem. 268, 12855–12863 (1993).
Quinn, C.L., Tao, N. & Schimmel, P. Species-specific microhelix aminoacylation by a eukaryotic pathogen tRNA synthetase dependent on a single base pair. Biochemistry 34, 12489–12495 (1995).
Fechter, P., Rudinger-Thirion, J., Tukalo, M. & Giegé, R. Major tyrosine identity determinants in Methanococcus jannaschii and Saccharomyces cerevisiae tRNATyr are conserved but expressed differently. Eur. J. Biochem. 268, 761–767 (2001).
Wang, L., Magliery, T.J, Liu, D.R. & Schultz, P.G. A new functional suppressor tRNA/aminoacyl-tRNA synthetase pair for the in vivo incorporation of unnatural amino acids into proteins. J. Am. Chem. Soc. 122, 5010–5011 (2000).
Giegé, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Marck, C. & Grosjean, H. tRNomics: analysis of tRNA genes from 50 genomes of eukarya, archaea, and bacteria reveals anticodon-sparing strategies and domain-specific features. RNA 8, 1189–1232 (2002).
Himeno, H., Hasegawa, T., Ueda, T., Watanabe, K. & Shimizu, M. Conversion of aminoacylation specificity from tRNATyr to tRNASerin vitro. Nucleic Acids Res. 18, 6815–6819 (1990).
Lee, C.P. & RajBhandary, U.L. Mutants of Escherichia coli initiator tRNA that are aminoacylated with tyrosine by yeast extracts. Proc. Natl. Acad. Sci. USA 88, 11378–11382 (1991).
Fechter, P., Rudinger-Thirion, J., Théobald-Dietrich, A. & Giegé, R. Identity of tRNA for yeast tyrosyl-tRNA synthetase: tyrosylation is more sensitive to identity nucleotides than to structural features. Biochemistry 39, 1725–1733 (2000).
Brick, P., Bhat, T.N. & Blow, D.M. Structure of tyrosyl-tRNA synthetase refined at 2.3 Å resolution. Interaction of the enzyme with the tyrosyl adenylate intermediate. J. Mol. Biol. 208, 83–98 (1989).
Qiu, X. et al. Crystal structure of Staphylococcus aureus tyrosyl-tRNA synthetase in complex with a class of potent and specific inhibitors. Protein Sci. 10, 2008–2016 (2001).
Yaremchuk, A., Kriklivyi, I., Tukalo, M. & Cusack, S. Class I tyrosyl-tRNA synthetase has a class II mode of cognate tRNA recognition. EMBO J. 21, 3829–3840 (2002).
Yang, X.L., Skene, R.J., McRee, D.E. & Schimmel, P. Crystal structure of a human aminoacyl-tRNA synthetase cytokine. Proc. Natl. Acad. Sci. USA 99, 15369–15374 (2002).
Steer, B.A. & Schimmel, P. Major anticodon-binding region missing from an archaebacterial tRNA synthetase. J. Biol. Chem. 274, 35601–35606 (1999).
Eriani, G., Delarue, M., Poch, O., Gangloff, J. & Moras, D. Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347, 203–206 (1990).
Starzyk, R.M., Webster, T.A. & Schimmel, P. Evidence for dispensable sequences inserted into a nucleotide fold. Science 237, 1614–1618 (1987).
Ruff, M. et al. Class II aminoacyl transfer RNA synthetases: crystal structure of yeast aspartyl-tRNA synthetase complexed with tRNAAsp. Science 252, 1682–1689 (1991).
Rould, M.A., Perona, J.J., Söll, D. & Steitz, T.A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNAGln and ATP at 2.8 Å resolution. Science 246, 1135–1142 (1989).
Wakasugi, K., Quinn, C.L., Tao, N. & Schimmel, P. Genetic code in evolution: switching species-specific aminoacylation with a peptide transplant. EMBO J. 17, 297–305 (1998).
Steer, B.A. & Schimmel, P. Domain-domain communication in a miniature archaebacterial tRNA synthetase. Proc. Natl. Acad. Sci. USA 96, 13644–13649 (1999).
van Tol, H., Stange, N., Gross, H.J. & Beier, H. A human and a plant intron-containing tRNATyr gene are both transcribed in a HeLa cell extract but spliced along different pathways. EMBO J. 6, 35–41 (1987).
Brick, P. & Blow, D.M. Crystal structure of a deletion mutant of a tyrosyl-tRNA synthetase complexed with tyrosine. J. Mol. Biol. 194, 287–297 (1987).
Zhang, D., Vaidehi, N., Goddard, W.A. III, Danzer, J.F. & Debe, D. Structure-based design of mutant Methanococcus jannaschii tyrosyl-tRNA synthetase for incorporation of O-methyl-L-tyrosine. Proc. Natl. Acad. Sci. USA 99, 6579–6584 (2002).
Fechter, P., Rudinger, J., Giegé, R. & Théobald-Dietrich, A. Ribozyme processed tRNA transcripts with unfriendly internal promoter for T7 RNA polymerase: production and activity. FEBS Lett. 436, 99–103 (1998).
LeMaster, D.M. & Richards, F.M. 1H-15N heteronuclear NMR studies of Escherichia coli thioredoxin in samples isotopically labeled by residue type. Biochemistry 24, 7263–7268 (1985).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collection in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Weeks, C.M. & Miller, R. Optimizing Shake-and-Bake for proteins. Acta Crystallogr. D 55, 492–500 (1999).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F. & Higgins, D.G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Gouet, P., Courcelle, E., Stuart, D.I. & Métoz, F. ESPript: multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Acknowledgements
We acknowledge the contributions of M. Kawamoto and H. Sakai for the synchrotron data collection at BL41XU in SPring-8 (Harima, Japan). We thank D. Kiga for helpful advice on the steady-state kinetics of the aminoacylation. We also thank K. Ohtake and M. Takahashi for help in the construction of the TyrRS expression system and R. Ishii for help in the data collection at BL41XU in SPring-8. This work was supported in part by Grants-in-Aid for Scientific Research on Priority Areas from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT) of Japan (S.Y. and O.N.) and by the National Project on Protein Structural and Functional Analyses of MEXT (S.Y.) and the Human Frontier Science Program (O.N. and S.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kobayashi, T., Nureki, O., Ishitani, R. et al. Structural basis for orthogonal tRNA specificities of tyrosyl-tRNA synthetases for genetic code expansion. Nat Struct Mol Biol 10, 425–432 (2003). https://doi.org/10.1038/nsb934
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb934
This article is cited by
-
A swapped genetic code prevents viral infections and gene transfer
Nature (2023)
-
Expanding the substrate scope of pyrrolysyl-transfer RNA synthetase enzymes to include non-α-amino acids in vitro and in vivo
Nature Chemistry (2023)
-
Rapid discovery and evolution of orthogonal aminoacyl-tRNA synthetase–tRNA pairs
Nature Biotechnology (2020)
-
Site-Specific Labelling of Multidomain Proteins by Amber Codon Suppression
Scientific Reports (2018)
-
New in silico approach to assessing RNA secondary structures with non-canonical base pairs
BMC Bioinformatics (2015)