Abstract
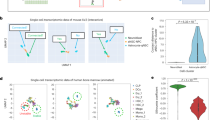
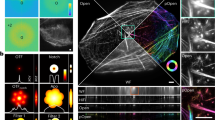
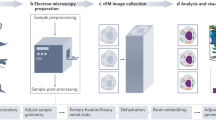
Open-Source 3D Visualization-Assisted Analysis (Vaa3D) is a software platform for the visualization and analysis of large-scale multidimensional images. In this protocol we describe how to use several popular features of Vaa3D, including (i) multidimensional image visualization, (ii) 3D image object generation and quantitative measurement, (iii) 3D image comparison, fusion and management, (iv) visualization of heterogeneous images and respective surface objects and (v) extension of Vaa3D functions using its plug-in interface. We also briefly demonstrate how to integrate these functions for complicated applications of microscopic image visualization and quantitative analysis using three exemplar pipelines, including an automated pipeline for image filtering, segmentation and surface generation; an automated pipeline for 3D image stitching; and an automated pipeline for neuron morphology reconstruction, quantification and comparison. Once a user is familiar with Vaa3D, visualization usually runs in real time and analysis takes less than a few minutes for a simple data set.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Long, F., Zhou, J. & Peng, H. Visualization and analysis of 3D microscopic images. PLoS Comput. Biol. 8, e1002519 (2012).
Clendenon, J.L., Phillips, C.L., Sandoval, R.M., Fang, S. & Dunn, K.W. Voxx: a PC-based, near real-time volume rendering system for biological microscopy. Am. J. Physiol. Cell Physiol. 282, C213–C218 (2002).
Swedlow, J.R., Goldberg, I., Brauner, E. & Sorger, P.K. Informatics and quantitative analysis in biological imaging. Science 300, 100–102 (2003).
Abràmoff, M.D., Magalhães, P.J. & Ram, S.J. Image processing with ImageJ. Biophotonics Internatl. 11, 36–42 (2004).
de Chaumont, F. et al. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Sommer, C., Straehle, C., Kothe, U. & Hamprecht, F.A. Ilastik: Interactive learning and segmentation toolkit. Biomedical Imaging: From Nano to Macro, 2011 IEEE International Symposium on 230–233 (2011).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Murphy, R.F. Cell organizer: image-derived models of subcellular organization and protein distribution. Comput. Methods Cell Biol. 110, 179 (2012).
Long, F., Peng, H., Liu, X., Kim, S.K. & Myers, E. A 3D digital atlas of C. elegans and its application to single-cell analyses. Nat. Methods 6, 667–672 (2009).
Luisi, J., Narayanaswamy, A., Galbreath, Z. & Roysam, B. The FARSIGHT trace editor: an open source tool for 3-D inspection and efficient pattern analysis aided editing of automated neuronal reconstructions. Neuroinformatics 9, 305–315 (2011).
Kvilekval, K., Fedorov, D., Obara, B., Singh, A. & Manjunath, B. Bisque: a platform for bioimage analysis and management. Bioinformatics 26, 544–552 (2010).
Lau, C. et al. Exploration and visualization of gene expression with neuroanatomy in the adult mouse brain. BMC Bioinformatics 9, 153 (2008).
Peng, H. et al. BrainAligner: 3D registration atlases of Drosophila brains. Nat. Methods 8, 493–498 (2011).
Peng, H. Bioimage informatics: a new area of engineering biology. Bioinformatics 24, 1827–1836 (2008).
Swedlow, J.R., Goldberg, I.G. & Eliceiri, K.W. Bioimage informatics for experimental biology. Ann. Rev. Biophys. 38, 327 (2009).
Shamir, L., Delaney, J.D., Orlov, N., Eckley, D.M. & Goldberg, I.G. Pattern recognition software and techniques for biological image analysis. PLoS Comput. Biol. 6, e1000974 (2010).
Danuser, G. Computer vision in cell biology. Cell 147, 973–978 (2011).
Eliceiri, K.W. et al. Biological imaging software tools. Nat. Methods 9, 697–710 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Peng, H., Ruan, Z., Long, F., Simpson, J.H. & Myers, E.W. V3D enables real-time 3D visualization and quantitative analysis of large-scale biological image data sets. Nat. Biotechnol. 28, 348–353 (2010).
Gonzalez-Bellido, P.T., Peng, H., Yang, J., Georgopoulos, A.P. & Olberg, R.M. Eight pairs of descending visual neurons in the dragonfly give wing motor centers accurate population vector of prey direction. Proc. Natl. Acad. Sci. USA 110, 696–701 (2013).
Yu, Y. & Peng, H. Automated high speed stitching of large 3D microscopic images. Biomedical Imaging: From Nano to Macro, 2011 IEEE International Symposium on Biomedical Imaging: From Nano to Macro (ISBI'2011). 238–241 (2011).
Bria, A. & Iannello, G. TeraStitcher-a tool for fast automatic 3D-stitching of teravoxel-sized microscopy images. BMC Bioinformatics 13, 316 (2012).
Silvestri, L., Bria, A., Sacconi, L., Iannello, G. & Pavone, F. Confocal light sheet microscopy: micron-scale neuroanatomy of the entire mouse brain. Optics Express 20, 20582–20598 (2012).
Peng, H., Long, F. & Myers, G. Automatic 3D neuron tracing using all-path pruning. Bioinformatics 27, i239–i247 (2011).
Scorcioni, R., Polavaram, S. & Ascoli, G.A. L-Measure: a web-accessible tool for the analysis, comparison and search of digital reconstructions of neuronal morphologies. Nat. Protoc. 3, 866–876 (2008).
Acknowledgements
We thank Z. Ruan, H. Xiao and Y. Wan for developing several modules briefly discussed in this article; L. Qu, Y. Yu, J. Zhou, L. Ibanez, P. Yu, C. Bruns and many other contributors to the Vaa3D project; C. Doe, E. Heckscher, R. Kerr, J. Simpson, P. Chung, G. Rubin, L. Silvestri, L. Sacconi, F.S. Pavone, http://NeuroMorpho.org/, the Janelia FlyLight project and many other collaborators for providing test data for the demonstrations. We thank the Janelia Farm Research Campus of the Howard Hughes Medical Institute and the Allen Institute for Brain Science for support of this work.
Author information
Authors and Affiliations
Contributions
H.P. conceived and developed this study; A.B. and G.I. developed the TeraStitcher plug-in; F.L. developed the cell segmentation plug-in; H.P. supervised or assisted all plug-in developments; H.P. wrote the manuscript with the contributions from co-authors. Z.Z. helped in testing the options and in editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Peng, H., Bria, A., Zhou, Z. et al. Extensible visualization and analysis for multidimensional images using Vaa3D. Nat Protoc 9, 193–208 (2014). https://doi.org/10.1038/nprot.2014.011
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.011
This article is cited by
-
Mapping of individual sensory nerve axons from digits to spinal cord with the transparent embedding solvent system
Cell Research (2024)
-
Vesicular HMGB1 release from neurons stressed with spreading depolarization enables confined inflammatory signaling to astrocytes
Journal of Neuroinflammation (2023)
-
In-silico clearing approach for deep refractive index tomography by partial reconstruction and wave-backpropagation
Light: Science & Applications (2023)
-
CellRemorph: A Toolkit for Transforming, Selecting, and Slicing 3D Cell Structures on the Road to Morphologically Detailed Astrocyte Simulations
Neuroinformatics (2023)
-
nGauge: Integrated and Extensible Neuron Morphology Analysis in Python
Neuroinformatics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.