Abstract
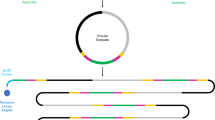
A novel method called “multiplex-terminal restriction fragment length polymorphism (M-TRFLP)” has been recently developed which can be used for simultaneous analysis of the community composition of two or more microbial taxa (up to four). This method can also be used for microbial diagnostic purposes. For M-TRFLP analysis, primers specific to different target genes are used for multiplex-PCR, with one primer for each target being labeled with a unique fluorescent dye at its 5′ end. Restriction digestion of the amplified products followed by fragment size analysis on a DNA sequencer produces profiles for targeted genes, which can be distinguished from each other by the color of the terminal fragments imparted by the unique fluorescent dye used for primer labeling. In contrast to current protocols, M-TRFLP allows multiple communities or multiple targets (genes) data to be obtained in just one reaction and therefore saves time, cost and labor. This protocol can be completed in 5–8 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Prosser, J.I. Molecular and functional diversity in soil micro-organisms. Plant Soil 244, 9–17 (2002).
Singh, B.K. et al. Investigating microbial community structure in soil by physiological, biochemical and molecular fingerprinting methods. Eur. J. Soil Sci. 57, 72–82 (2006).
Singh, B.K., Millard, P., Whiteley, A.S. & Murrell, J.C. Unravelling rhizosphere–microbial interactions: opportunities and limitations. Trends Microbiol. 12, 386–393 (2004).
Jacobs, T. Any diamond in the diagnostic coal. Nat. Biotechnol. 24, 930 (2006).
Moore, M.M. & Feist, M.D. Real-time PCR method for Salmonella spp. targeting the stn gene. J. Appl. Microbiol. (in the press) (2006) doi:10.1111/j.1365–2672.2006.03073.x.
Hein, I., Flekna, G., Krassnig, M. & Wagner, M. Real-time PCR for the detection of Salmonella spp. In food: an alternative approach to a conventional PCR system suggested by the FOOD-PCR project. J. Microbiol. Method (in the press) (2006).
Chiang, Y.-C., Yang, C.-Y., Li, C., Ho, Y.-C. & Tsen, H.-Y. Identification of Bacillus spp. Escherichia coli, Salmonella spp., Staphylococcus spp. and Vibrio spp. with 16s ribosomal DNA based oligonucleotide array hybridisation. Int. J. Food Microbiol. 107, 131–137 (2006).
Christensen, J.E., Stencil, J.A. & Reed, K.D. Rapid identification of bacteria from positive blood cultures by terminal restriction fragment length polymorphism profile analysis of the 16S rRNA gene. J. Clin. Microbiol. 41, 3790–3800 (2003).
Avaniss-Aghajani, E. et al. A molecular technique for rapid identification of mycobacteria. J. Clin. Microbiol. 34, 98–102 (1996).
Song, Y., Kato, N., Liu, C., Matsumiya, Y., Kato, H. & Watanabe, K. Rapid identification of 11 human intestinal Lactobacillus species by multiplex PCR assays using group- and species-specific primers derived from the 16S–23S rRNA intergenic spacer region and its flanking 23S rRNA. FEMS Microbiol. Lett. 187, 167–173 (2000).
Veiga, M.I., Ferreira, P.E., Bjorkman, A. & Gil, J.P. Multiplex PCR-RFLP methods for pfcrt, pfmdr1 and pfdhfr mutations in Plasmodium falciparum. Mol. Cell Probes 20, 100–104 (2006).
Singh, B.K., Nazaries, L., Munro, S., Anderson, I.C. & Campbell, C.D. Multiplex-terminal restriction fragment length polymorphism for rapid and simultaneous analysis of different components of the soil microbial community. Appl. Environ. Microbiol. 72, 7278–7285 (2006).
Yu, C.P., Ahuja, R., Sayler, G. & Chu, K.H. Quantitative molecular assay for fingerprinting microbial communities of wastewater and estrogen-degrading consortia. Appl. Environ. Microbiol. 71, 1433–1444 (2005).
Rompler, H. et al. Multiplex amplification of ancient DNA. Nat. Prot. 1, 720–728 (2006).
Acknowledgements
We thank Loic Nazaries and Stacey Munro for initial technical support on this method. We also thank Mrs Laura Price (Unilever, Safety & Environmental Assurance Centre, Bedfordshire), Dr. Catriona Macdonald (Environmental Science Research, Porirua, New Zealand), Dr. Oliver Price (Cambridge Environmental Assessment, Boxworth, Cambridge) and Dr. Ian Anderson and Dr. Colin Campbell (Macaulay Institute), three anonymous referees and Dr. K. Barnes for their valuable suggestions to improve the quality of the manuscript. A patent application for commercial use of this method is under consideration by the British Patent Office.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Author declares competing financial interests: a patent application is under consideration which has been mentioned in the Acknowledgements section
Supplementary information
Supplementary Table 1
List of primers used in the present study. (PDF 50 kb)
Rights and permissions
About this article
Cite this article
Singh, B., Thomas, N. Multiplex-terminal restriction fragment length polymorphism. Nat Protoc 1, 2428–2433 (2006). https://doi.org/10.1038/nprot.2006.392
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.392
This article is cited by
-
Frequency of disturbance alters diversity, function, and underlying assembly mechanisms of complex bacterial communities
npj Biofilms and Microbiomes (2019)
-
Response of ammonia oxidizers and denitrifiers to repeated applications of a nitrification inhibitor and a urease inhibitor in two pasture soils
Journal of Soils and Sediments (2017)
-
Effects of 3,4-dimethylpyrazole phosphate (DMPP) on nitrification and the abundance and community composition of soil ammonia oxidizers in three land uses
Biology and Fertility of Soils (2016)
-
Deterministic processes vary during community assembly for ecologically dissimilar taxa
Nature Communications (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.