Abstract
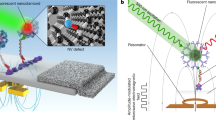
Rapid and highly sensitive detection of DNA is critical in diagnosing genetic diseases. Conventional approaches often rely on cumbersome, semi-quantitative amplification of target DNA to improve detection sensitivity. In addition, most DNA detection systems (microarrays, for example), regardless of their need for target amplification, require separation of unhybridized DNA strands from hybridized stands immobilized on a solid substrate, and are thereby complicated by solution–surface binding kinetics1,2. Here, we report an ultrasensitive nanosensor based on fluorescence resonance energy transfer (FRET) capable of detecting low concentrations of DNA in a separation-free format. This system uses quantum dots (QDs)3,4,5 linked to DNA probes to capture DNA targets. The target strand binds to a dye-labelled reporter strand thus forming a FRET donor–acceptor ensemble. The QD also functions as a concentrator that amplifies the target signal by confining several targets in a nanoscale domain. Unbound nanosensors produce near-zero background fluorescence, but on binding to even a small amount of target DNA (∼50 copies or less) they generate a very distinct FRET signal. A nanosensor-based oligonucleotide ligation assay has been demonstrated to successfully detect a point mutation6 typical of some ovarian tumours in clinical samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Southern, E., Mir, K. & Shchepinov, M. Molecular interactions on microarrays. Nature Genet. 21, 5–9 (1999).
Taton, T. A., Mirkin, C. A. & Letsinger, R. L. Scanometric DNA array detection with nanoparticle probes. Science 289, 1757–1760 (2000).
Chan, W. C. W. & Nie, S. M. Quantum dot bioconjugates for ultrasensitive nonisotopic detection. Science 281, 2016–2018 (1998).
Bruchez, M., Moronne, M., Gin, P., Weiss, S. & Alivisatos, A. P. Semiconductor nanocrystals as fluorescent biological labels. Science 281, 2013–2016 (1998).
Medintz, I. L., Uyeda, H. T., Goldman, E. R. & Mattoussi, H. Quantum dot bioconjugates for imaging, labelling and sensing. Nature Mater. 4, 435–446 (2005).
Ho, C. L., Karman, R. J., Dehari, R., Wang, T. L. & Shih, I. M. Mutations of BRAF and KRAS precede the development of ovarian serous borderline tumors. Cancer Res. 64, 6915–6918 (2004).
Cardullo, R. A., Agrawal, S., Flores, C., Zamecnik, P. C. & Wolf, D. E. Detection of nucleic-acid hybridization by nonradiative fluorescence resonance energy-transfer. Proc. Natl Acad. Sci. USA 85, 8790–8794 (1988).
Holland, P. M., Abramson, R. D., Watson, R. & Gelfand, D. H. Detection of specific polymerase chain-reaction product by utilizing the 5′→3′ exonuclease activity of thermus-aquaticus dna-polymerase. Proc. Natl Acad. Sci. USA 88, 7276–7280 (1991).
Tyagi, S. & Kramer, F. R. Molecular beacons: Probes that fluoresce upon hybridization. Nature Biotechnol. 14, 303–308 (1996).
Dubertret, B., Calame, M. & Libchaber, A. J. Single-mismatch detection using gold-quenched fluorescent oligonucleotides. Nature Biotechnol. 19, 365–370 (2001).
Knemeyer, J. P., Marmé, N. & Sauer, M. Probes for detection of specific DNA sequences at the single-molecule level. Anal. Chem. 72, 3717–3724 (2000).
Barnes, M. D., Ng, K. C., Whitten, W. B. & Ramsey, J. M. Detection of single rhodamine-6g molecules in levitated microdroplets. Anal. Chem. 65, 2360–2365 (1993).
Shera, E. B., Seitzinger, N. K., Davis, L. M., Keller, R. A. & Soper, S. A. Detection of single fluorescent molecules. Chem. Phys. Lett. 174, 553–557 (1990).
Nie, S. M., Chiu, D. T. & Zare, R. N. Probing individual molecules with confocal fluorescence microscopy. Science 266, 1018–1021 (1994).
Eigen, M. & Rigler, R. Sorting single molecules—application to diagnostics and evolutionary biotechnology. Proc. Natl Acad. Sci. USA 91, 5740–5747 (1994).
Castro, A. & Williams, J. G. K. Single-molecule detection of specific nucleic acid sequences in unamplified genomic DNA. Anal. Chem. 69, 3915–3920 (1997).
Wang, T. H., Peng, Y. H., Zhang, C. Y., Wong, P. K. & Ho, C. M. Single-molecule tracing on a fluidic microchip for quantitative detection of low-abundance nucleic acids. J. Am. Chem. Soc. 127, 5354–5359 (2005).
Zhang, C. Y., Chao, S. Y. & Wang, T. H. Comparative quantification of nucleic acids using single-molecule detection and molecular beacons. Analyst 130, 483–488 (2005).
Wabuyele, M. B. et al. Approaching real-time molecular diagnostics: Single-pair fluorescence resonance energy transfer (spFRET) detection for the analysis of low abundant point mutations in K-ras oncogenes. J. Am. Chem. Soc. 125, 6937–6945 (2003).
Medintz, I. L. et al. A fluorescence resonance energy transfer-derived structure of a quantum dot-protein bioconjugate nanoassembly. Proc. Natl Acad. Sci. USA 101, 9612–9617 (2004).
Medintz, I. L. et al. Self-assembled nanoscale biosensors based on quantum dot FRET donors. Nature Mater. 2, 630–638 (2003).
Willard, D. M., Carillo, L. L., Jung, J. & Van Orden, A. CdSe-ZnS quantum dots as resonance energy transfer donors in a model protein-protein binding assay. Nano Lett. 1, 469–474 (2001).
Mattoussi, H. et al. Self-assembly of CdSe-ZnS quantum dot bioconjugates using an engineered recombinant protein. J. Am. Chem. Soc. 122, 12142–12150 (2000).
Clapp, A. R. et al. Fluorescence resonance energy transfer between quantum dot donors and dye-labeled protein acceptors. J. Am. Chem. Soc. 126, 301–310 (2004).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy (Kluwer Academic/Plenum, New York, 1999).
Ha, T. et al. Ligand-induced conformational changes observed in single RNA molecules. Proc. Natl Acad. Sci. USA 96, 9077–9082 (1999).
Deniz, A. A. et al. Single-pair fluorescence resonance energy transfer on freely diffusing molecules: Observation of Forster distance dependence and subpopulations. Proc. Natl Acad. Sci. USA 96, 3670–3675 (1999).
Bonnet, G., Krichevsky, O. & Libchaber, A. Kinetics of conformational fluctuations in DNA hairpin-loops. Proc. Natl Acad. Sci. USA 95, 8602–8606 (1998).
Landegren, U., Kaiser, R., Sanders, J. & Hood, L. A ligase-mediated gene detection technique. Science 241, 1077–1080 (1988).
Nickerson, D. A. et al. Automated DNA diagnostics using an elisa-based oligonucleotide ligation assay. Proc. Natl Acad. Sci. USA 87, 8923–8927 (1990).
Kwok, P. Y. Single Nucleotide Polymorphisms Methods and Protocols (Human Press, Totowa, New Jersey, 2003).
Acknowledgements
The authors thank Y. Peng, S. Yang, S. Lin and Y. P. Ho for valuable discussions, I. M. Shih for providing us with PCR products from clinical samples for Kras point mutation detections, and L. Brand and D. Toptygin for providing support with the TCSPC fluorescence lifetime measurements. This work was supported primarily by NSF under award no. DBI-0352407 and also by the Whitaker Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary figure S1-5 (PDF 310 kb)
Rights and permissions
About this article
Cite this article
Zhang, CY., Yeh, HC., Kuroki, M. et al. Single-quantum-dot-based DNA nanosensor. Nature Mater 4, 826–831 (2005). https://doi.org/10.1038/nmat1508
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat1508
This article is cited by
-
Single-molecule fluorescence methods for protein biomarker analysis
Analytical and Bioanalytical Chemistry (2023)
-
Temporal evolution of optical absorption and emission spectra of thiol capped CdTe quantum dots
Applied Physics A (2022)
-
Energy Transfer Interactions and Sensing Characteristics of Gain-Assisted and Graphene-Coated Plasmonic Nanomatryoshka
Plasmonics (2021)
-
Synthesis and application of stationary phase for DNA-affinity chromatographic analysis of unmodified and antisense oligonucleotide
Analytical and Bioanalytical Chemistry (2021)
-
Single-molecule photoreaction quantitation through intraparticle-surface energy transfer (i-SET) spectroscopy
Nature Communications (2020)