Abstract
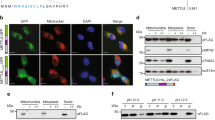
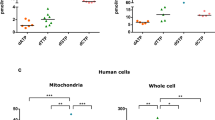
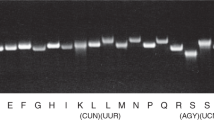
In human mitochondria, the AUA codon encodes methionine via a mitochondrial transfer RNA for methionine (mt-tRNAMet) that contains 5-formylcytidine (f5C) at the first position of the anticodon (position 34). f5C34 is required for deciphering the AUA codon during protein synthesis. Until now, the biogenesis and physiological role of f5C34 were unknown. We demonstrate that biogenesis of f5C34 is initiated by S-adenosylmethionine (AdoMet)-dependent methylation catalyzed by NSUN3, a putative methyltransferase in mitochondria. NSUN3-knockout cells showed strong reduction in mitochondrial protein synthesis and reduced oxygen consumption, leading to deficient mitochondrial activity. We reconstituted formation of 5-methylcytidine (m5C) at position 34 (m5C34) on mt-tRNAMet with recombinant NSUN3 in the presence of AdoMet, demonstrating that NSUN3-mediated m5C34 formation initiates f5C34 biogenesis. We also found two disease-associated point mutations in mt-tRNAMet that impaired m5C34 formation by NSUN3, indicating that a lack of f5C34 has pathological consequences.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Machnicka, M.A. et al. MODOMICS: a database of RNA modification pathways—2013 update. Nucleic Acids Res. 41, D262–D267 (2013).
Bjork, G. in tRNA: Structure, Biosynthesis, and Function (eds. Soll, D. & RajBhandary, U.L.) 165–205 (American Society for Microbiology, 1995).
Suzuki, T. in Fine-Tuning of RNA Functions by Modification and Editing vol. 12 (ed. Grosjean, H.) 23–69 (Springer, 2005).
Wallace, D.C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: a dawn for evolutionary medicine. Annu. Rev. Genet. 39, 359–407 (2005).
Suzuki, T., Nagao, A. & Suzuki, T. Human mitochondrial tRNAs: biogenesis, function, structural aspects, and diseases. Annu. Rev. Genet. 45, 299–329 (2011).
Hällberg, B.M. & Larsson, N.G. Making proteins in the powerhouse. Cell Metab. 20, 226–240 (2014).
Suzuki, T. & Suzuki, T. A complete landscape of post-transcriptional modifications in mammalian mitochondrial tRNAs. Nucleic Acids Res. 42, 7346–7357 (2014).
Suzuki, T., Suzuki, T., Wada, T., Saigo, K. & Watanabe, K. Taurine as a constituent of mitochondrial tRNAs: new insights into the functions of taurine and human mitochondrial diseases. EMBO J. 21, 6581–6589 (2002).
Kirino, Y. et al. Codon-specific translational defect caused by a wobble modification deficiency in mutant tRNA from a human mitochondrial disease. Proc. Natl. Acad. Sci. USA 101, 15070–15075 (2004).
Kurata, S. et al. Modified uridines with C5-methylene substituents at the first position of the tRNA anticodon stabilize U•G wobble pairing during decoding. J. Biol. Chem. 283, 18801–18811 (2008).
Moriya, J. et al. A novel modified nucleoside found at the first position of the anticodon of methionine tRNA from bovine liver mitochondria. Biochemistry 33, 2234–2239 (1994).
Takemoto, C. et al. Unconventional decoding of the AUA codon as methionine by mitochondrial tRNAMet with the anticodon f5CAU as revealed with a mitochondrial in vitro translation system. Nucleic Acids Res. 37, 1616–1627 (2009).
Cantara, W.A., Murphy, F.V. IV, Demirci, H. & Agris, P.F. Expanded use of sense codons is regulated by modified cytidines in tRNA. Proc. Natl. Acad. Sci. USA 110, 10964–10969 (2013).
Suzuki, T., Nagao, A. & Suzuki, T. Human mitochondrial diseases caused by lack of taurine modification in mitochondrial tRNAs. Wiley Interdiscip. Rev. RNA 2, 376–386 (2011).
Yasukawa, T., Suzuki, T., Ueda, T., Ohta, S. & Watanabe, K. Modification defect at anticodon wobble nucleotide of mitochondrial tRNAsLeu(UUR) with pathogenic mutations of mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. J. Biol. Chem. 275, 4251–4257 (2000).
Yasukawa, T. et al. Defect in modification at the anticodon wobble nucleotide of mitochondrial tRNALys with the MERRF encephalomyopathy pathogenic mutation. FEBS Lett. 467, 175–178 (2000).
Kirino, Y., Goto, Y., Campos, Y., Arenas, J. & Suzuki, T. Specific correlation between the wobble modification deficiency in mutant tRNAs and the clinical features of a human mitochondrial disease. Proc. Natl. Acad. Sci. USA 102, 7127–7132 (2005).
Yasukawa, T. et al. Wobble modification deficiency in mutant tRNAs in patients with mitochondrial diseases. FEBS Lett. 579, 2948–2952 (2005).
Umeda, N. et al. Mitochondria-specific RNA-modifying enzymes responsible for the biosynthesis of the wobble base in mitochondrial tRNAs. Implications for the molecular pathogenesis of human mitochondrial diseases. J. Biol. Chem. 280, 1613–1624 (2005).
Takeuchi, N. et al. Mammalian mitochondrial methionyl-tRNA transformylase from bovine liver. Purification, characterization, and gene structure. J. Biol. Chem. 273, 15085–15090 (1998).
Momb, J. & Appling, D.R. Mitochondrial one-carbon metabolism and neural tube defects. Birth Defects Res. A Clin. Mol. Teratol. 100, 576–583 (2014).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Miyauchi, K., Ohara, T. & Suzuki, T. Automated parallel isolation of multiple species of non-coding RNAs by the reciprocal circulating chromatography method. Nucleic Acids Res. 35, e24 (2007).
Rhee, H.W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Robinson, B.H., Petrova-Benedict, R., Buncic, J.R. & Wallace, D.C. Nonviability of cells with oxidative defects in galactose medium: a screening test for affected patient fibroblasts. Biochem. Med. Metab. Biol. 48, 122–126 (1992).
Lott, M.T. et al. mtDNA variation and analysis using MITOMAP and MITOMASTER. Curr. Protoc. Bioinformatics 1, 1.23.1–1.23.26 (2013).
Yue, Y., Liu, J. & He, C. RNA N6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev. 29, 1343–1355 (2015).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Païs de Barros, J.P. et al. 2′-O-methyl-5-formylcytidine (f5Cm), a new modified nucleotide at the 'wobble' of two cytoplasmic tRNAs Leu (NAA) from bovine liver. Nucleic Acids Res. 24, 1489–1496 (1996).
Huber, S.M. et al. Formation and abundance of 5-hydroxymethylcytosine in RNA. ChemBioChem 16, 752–755 (2015).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981).
Rooijers, K., Loayza-Puch, F., Nijtmans, L.G. & Agami, R. Ribosome profiling reveals features of normal and disease-associated mitochondrial translation. Nat. Commun. 4, 2886 (2013).
Spencer, A.C., Heck, A., Takeuchi, N., Watanabe, K. & Spremulli, L.L. Characterization of the human mitochondrial methionyl-tRNA synthetase. Biochemistry 43, 9743–9754 (2004).
Chi, L. & Delgado-Olguín, P. Expression of NOL1/NOP2/Sun domain (Nsun) RNA methyltransferase family genes in early mouse embryogenesis. Gene Expr. Patterns 13, 319–327 (2013).
Sharma, S., Yang, J., Watzinger, P., Kötter, P. & Entian, K.D. Yeast Nop2 and Rcm1 methylate C2870 and C2278 of the 25S rRNA, respectively. Nucleic Acids Res. 41, 9062–9076 (2013).
Brzezicha, B. et al. Identification of human tRNA:m5C methyltransferase catalysing intron-dependent m5C formation in the first position of the anticodon of the pre-tRNALeu(CAA). Nucleic Acids Res. 34, 6034–6043 (2006).
Hussain, S. et al. NSun2-mediated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs. Cell Rep. 4, 255–261 (2013).
Squires, J.E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Haag, S. et al. NSUN6 is a human RNA methyltransferase that catalyzes formation of m5C72 in specific tRNAs. RNA 21, 1532–1543 (2015).
Metodiev, M.D. et al. NSUN4 is a dual function mitochondrial protein required for both methylation of 12S rRNA and coordination of mitoribosomal assembly. PLoS Genet. 10, e1004110 (2014).
Spåhr, H., Habermann, B., Gustafsson, C.M., Larsson, N.G. & Hallberg, B.M. Structure of the human MTERF4-NSUN4 protein complex that regulates mitochondrial ribosome biogenesis. Proc. Natl. Acad. Sci. USA 109, 15253–15258 (2012).
Harris, T., Marquez, B., Suarez, S. & Schimenti, J. Sperm motility defects and infertility in male mice with a mutation in Nsun7, a member of the Sun domain-containing family of putative RNA methyltransferases. Biol. Reprod. 77, 376–382 (2007).
Takemoto, C., Ueda, T., Miura, K. & Watanabe, K. Nucleotide sequences of animal mitochondrial tRNAsMet possibly recognizing both AUG and AUA codons. Nucleic Acids Symp. Ser. 42, 77–78 (1999).
Watanabe, Y. et al. Primary and higher order structures of nematode (Ascaris suum) mitochondrial tRNAs lacking either the T or D stem. J. Biol. Chem. 269, 22902–22906 (1994).
Lu, Z. et al. The tRNAMet 4435A>G mutation in the mitochondrial haplogroup G2a1 is responsible for maternally inherited hypertension in a Chinese pedigree. Eur. J. Hum. Genet. 19, 1181–1186 (2011).
Qu, J. et al. The novel A4435G mutation in the mitochondrial tRNAMet may modulate the phenotypic expression of the LHON-associated ND4 G11778A mutation. Invest. Ophthalmol. Vis. Sci. 47, 475–483 (2006).
Tang, S. et al. Transition to next generation analysis of the whole mitochondrial genome: a summary of molecular defects. Hum. Mutat. 34, 882–893 (2013).
Pyzocha, N.K., Ran, F.A., Hsu, P.D. & Zhang, F. RNA-guided genome editing of mammalian cells. Methods Mol. Biol. 1114, 269–277 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Gao, Y., Yang, J., Cancilla, M.T., Meng, F. & McLuckey, S.A. Top-down interrogation of chemically modified oligonucleotides by negative electron transfer and collision induced dissociation. Anal. Chem. 85, 4713–4720 (2013).
Kato, K. et al. Structural and functional analyses of DNA-sensing and immune activation by human cGAS. PLoS One 8, e76983 (2013).
Reed, S.E., Staley, E.M., Mayginnes, J.P., Pintel, D.J. & Tullis, G.E. Transfection of mammalian cells using linear polyethylenimine is a simple and effective means of producing recombinant adeno-associated virus vectors. J. Virol. Methods 138, 85–98 (2006).
Arai, T. et al. Single methylation of 23S rRNA triggers late steps of 50S ribosomal subunit assembly. Proc. Natl. Acad. Sci. USA 112, E4707–E4716 (2015).
Sampson, J.R. & Uhlenbeck, O.C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro. Proc. Natl. Acad. Sci. USA 85, 1033–1037 (1988).
Stiernagle, T. in WormBook (ed. The C. elegans Research Community) doi:10.1895/wormbook.1.101.1 (11 February 2006).
Acknowledgements
We are grateful to Y. Sakaguchi, K. Miyauchi, T. Ohira, S. Kimura, H. Hojo, H. Lin, T. Chujo, Y. Ikeuchi and other members of Tsutomu Suzuki's laboratory for technical support and insightful discussions. Special thanks to O. Nureki, I. Koyama-Honda, N. Mizushima (University of Tokyo), C. Ushida (Hirosaki University) and Primetech Inc. (Japan) for materials and technical support. This work was supported by Grants-in-Aid for Scientific Research on Priority Areas from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT) and the Japan Society for the Promotion of Science (JSPS) (to Tsutomu Suzuki and Takeo Suzuki).
Author information
Authors and Affiliations
Contributions
Tsutomu Suzuki and Takeo Suzuki designed this study. S.N., L.K. and H.I. performed experiments. K.A. assisted with biochemical work. All authors discussed the results. Tsutomu Suzuki, Takeo Suzuki. and S.N. wrote the paper. Tsutomu Suzuki supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1 – 5 and Supplementary Tables 1 and 2. (PDF 1382 kb)
Rights and permissions
About this article
Cite this article
Nakano, S., Suzuki, T., Kawarada, L. et al. NSUN3 methylase initiates 5-formylcytidine biogenesis in human mitochondrial tRNAMet. Nat Chem Biol 12, 546–551 (2016). https://doi.org/10.1038/nchembio.2099
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2099
This article is cited by
-
RNA methylations in hepatic fibrosis, a gradually emerging new treatment strategy
Cell & Bioscience (2023)
-
Integrated analysis of RNA methylation regulators crosstalk and immune infiltration for predictive and personalized therapy of diabetic nephropathy
Human Genomics (2023)
-
Insights into the regulatory role of RNA methylation modifications in glioma
Journal of Translational Medicine (2023)
-
RNA modifications in cancer
British Journal of Cancer (2023)
-
NSUN3-mediated mitochondrial tRNA 5-formylcytidine modification is essential for embryonic development and respiratory complexes in mice
Communications Biology (2023)