Abstract
Selective autophagy is a quality control pathway through which cellular components are sequestered into double-membrane vesicles and delivered to specific intracellular compartments. This process requires autophagy receptors that link cargo to growing autophagosomal membranes. Selective autophagy is also implicated in various membrane trafficking events. Here we discuss the current view on how cargo selection and transport are achieved during selective autophagy, and point out molecular mechanisms that are congruent between autophagy and vesicle trafficking pathways.
Similar content being viewed by others
Main
Under basal conditions, autophagy is dedicated to the continuous renovation of the intracellular pool of proteins, carbohydrates, lipids, and organelles, and thus plays a critical role in cellular homeostasis. Following external or internal stress, autophagy is rapidly upregulated and exerts a cytoprotective function. During this process, cellular materials or invaders are sequestered by double-membrane vesicles known as autophagosomes, and delivered to the lysosome for degradation1,2,3. Autophagy was initially described as a non-selective bulk process induced in response to starvation; however, it is now clear that the turnover of damaged organelles, removal of protein aggregates, and elimination of intracellular pathogens, are highly selective and tightly regulated processes that require cargo recognition by the autophagy machinery2,3,4,5. Several targets of selective autophagy have been described, such as aggregated proteins (aggrephagy), mitochondria (mitophagy), peroxisomes (pexophagy), ribosomes (ribophagy), endoplasmic reticulum (reticulophagy) and pathogens (xenophagy). Our knowledge about the network of autophagy regulators has increased dramatically, bringing a new understanding of this highly selective and precisely regulated membrane-dependent process. Following autophagy induction, the rapid formation of numerous autophagosomes relies on different membrane sources. Membranes are recruited and recognized by autophagy receptors, which participate in cargo sequestration and degradation. Importantly, the autophagy machinery is also employed by non-degradative cellular pathways such as the secretion of immuno-modulatory molecules and cytokines, endosome transport, and the activation of hydrolytic enzymes by the cytoplasm-to-vacuole targeting (Cvt) pathway in yeast6.
The core autophagosomal machinery
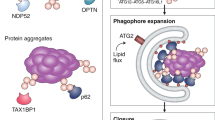
More than 30 autophagy-related (Atg) proteins have been identified in yeast7,8,9. Autophagy begins with the formation of a phagophore — a double-layered, crescent-shaped membrane — which expands to engulf cytoplasmic material, and then closes to form the autophagosome1 (Fig. 1a). Phagophore formation is primarily induced by the unc-51-like kinase (ULK1; Atg1 in yeast) complex, which can be regulated by the kinases AMPK (the cell energy sensor) and mTOR (the cell growth regulator)10,11,12,13,14. One of the crucial targets of ULK1 activity is the class III phosphatidylinositol 3-kinase (PI(3)K) complex, containing beclin 1 (Atg6 in yeast) and VPS34 (vacuolar protein sorting 34)15. When phosphorylated, beclin 1 promotes the local production of phosphatidylinositol 3-phosphate (PtdIns(3)P) by VPS34. PtdIns(3)P is recognized by early autophagic effector proteins that contain FYVE domains, such as Atg21 and WIPI1/2 (Atg18 in yeast)16,17. Although ULK1 is considered as the central autophagy regulator, there are ULK1-independent ways to trigger the autophagy cascade18,19. Moreover, ULK1-independent autophagy has been linked to non-canonical autophagic processes including LC3-associated phagocytosis (LAP; engulfment of extracellular particles in the form of cell debris or pathogens)20 and entosis (engulfment of a live epithelial cell by another epithelial cell)21.
(a) An LC3II-positive phagophore membrane forms around specific autophagic cargo based on the autophagy receptor. These receptors are characterized by their ability to simultaneously interact with autophagy modifiers (LC3/GABARAP/Atg8) and selective cargo, thus bridging the growing phagophore and targeted cargo. The phagophore then forms an intact, double-membrane autophagosome that encloses the cargo and subsequently fuses with the lysosome. Fusion allows for the lysosomal proteases to degrade the cargo together with its receptor and the inner autophagosomal membrane. (b,c) Several autophagy adaptors, that non-covalently bind autophagy modifiers, function in (b) autophagy initiation complex formation (ULK1, beclin 1, FIP200, VPS34) or in enzyme conjugation (ATG3, ATG7, ATG5, ATG12, ATG16) as well as in (c) fusion between autophagosomes and lysosome (SNX18) or microtubule transport machineries (FYCO1). The distinctive difference between autophagy receptors and autophagy adaptors is that adaptors are neither selective for cargos nor degraded within lysosomes with their respective cargos. Kinesin/dynein: motor proteins on the microtubule.
The core autophagic machinery in yeast also includes two ubiquitin (Ub)-like proteins, Atg12 and Atg8. In contrast to many other Ub-like proteins, the C-terminal glycine residue of Atg8 is not conjugated to the lysine residue of another protein but to the amino group of phosphatidyl ethanolamine (PE). In mammals, the autophagic machinery increases in complexity and the relevant Atg8 homologues cluster in two subfamilies: MAP-LC3 (microtubule-associated protein light chain 3; referred to as LC3 hereafter) and GABARAPs/GATE-16 (reviewed in ref. 22). Both subfamilies are essential for autophagosome biogenesis and act at different steps of phagophore assembly23,24. Importantly, these modifiers of autophagosomal membranes are also critical factors in cargo sequestration during selective autophagy and vesicle trafficking.
Membrane sources and their potential role in selectivity
Formation of autophagosomes is a rapid process that mobilizes a large pool of intracellular membranes during engulfment of the substrate. In mammals, the endoplasmic reticulum (ER), the Golgi and mitochondria are suggested sources for phagophore initiation. In yeast, the formation of phagophores occurs at a defined place in close proximity to the vacuole, termed the pre-autophagosomal structure (PAS)25,26. The phagophore membrane is then further propagated through the fusion of vesicles originating from different membrane sources including the plasma membrane, the Golgi, recycling endosomes, and probably others (reviewed in refs 6,27). Despite intensive research, it is still unresolved how these different membrane sources feed into autophagosomal membranes, and whether they have distinct functions in autophagy pathways. Dependent on the stress source, selective autophagy may require the use of spatially restricted membrane pools, and thus the origin of the membrane might contribute to the selectivity of cargo recognition. Assuming an economic management of cellular resources, membrane-containing cargos such as depolarized mitochondria or pieces of ER destined for lysosomal destruction could donate membranes for growing phagophores. Although no evidence for such a scenario yet exists, a recent study on lipid droplets suggests that the activity of autophagy also depends on the local availability of lipids28.
Inter-organelle contact sites have emerged as dynamic spots for lipid transition between membranes and for the growth of nascent phagophores. Mitochondria-associated ER membranes (MAMs) are claimed to be the origin of phagophores in mammalian cells29,30. These organelles are the major sites of PE production31 and harbour key Atg proteins and their regulators. Moreover, microscopic and tomographic studies have provided direct evidence for the presence of initial stages of phagophore formation on these organelles29,32,33. Proteins responsible for tethering ER to mitochondria, mitofusin 2 (MFN2) and PACS-2, and ER resident protein syntaxin 17, are important for subsequent ATG5-dependent LC3 lipidation and the recruitment of early autophagy effectors, such as ATG14 (refs 29,30,34). MAMs spatially overlap with specialized membrane compartments called omegasomes (omega-shaped DFCP1-positive structures), which are supposed to originate from the ER and have been claimed to serve as a cradle of phagophore formation and autophagosomal vesicles close to the ER (ref. 35). PtdIns(3)P, the initial membrane mark for autophagosomal membranes, is abundant in omegasomes and recruits FYVE-domain containing effector proteins such as WIPI2 (yeast Atg18)36, which are required for the formation of LC3II-positive autophagosomes. All these findings strongly support that MAMs/omegasomes are an important initiation site for phagophores. How ER-associated omegasomes participate in the clearance of damaged mitochondria was recently visualized by microscopy37. This study unveiled that mitophagy occurs in a bit-by-bit manner at the intersection of parkin-labelled mitochondrial tubules and the ER (ref. 37). The ER also contacts the plasma membrane38 through tethering by extended synaptotagmins (E-Syts) via PtdIns(4,5)P2 interactions. These contact sites allow for crosstalk between the organelles independently of membrane trafficking (reviewed in ref. 39) and may, similarly to MAMs, serve as phagophore initiation sites under specific conditions.
Whereas a large body of evidence indicates that the ER and mitochondria might be the main source for phagophore initiation, components of the endocytic compartment can also contribute to the expansion of the autophagosome. Several key autophagic proteins such as ATG9, ULK1 or ATG16 as well as the PI(3)K complex localize to the different kinds of endosomes defined by present subsets of Rab GTPases. Furthermore, a recent study in mammalian cells demonstrated that the plasma membrane contributes to autophagosome biogenesis via two distinct routes: ATG9- and ATG16-positive vesicles. Vesicles generated in both pathways are LC3-negative and converge in the recycling endosome40. In yeast, a subfraction of Atg9-containing single-membrane vesicles fuse with autophagosomal membranes33,41. However, the majority of them only transiently associate with and recruit autophagic regulatory proteins — for example, Atg1, the TRAPPIII complex and Ypt1 — to the pre-autophagosomal membrane, rather than providing large amount of lipids42,43,44. Clearly, more molecular studies are needed to elucidate the entangled pathways that regulate membrane formation during different autophagy pathways.
Autophagy receptors are key players in selective autophagy
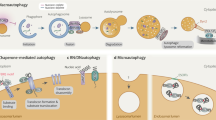
Whereas the autophagy response to starvation is bulk degradation of cytosolic material, other types of stresses such as damaged organelles or aggregated proteins require selective sequestration of the specific cargo into autophagosomal membranes. Selectivity is achieved through autophagy receptors, which recognize cargos tagged with degradation signals and the autophagosomal membrane through their LC3-interacting regions (LIR). LIR motifs in general interact with autophagy modifier proteins of the LC3/GABARAP family22. In yeast, five receptors have been described so far to mediate cargo selection: Atg19 and Atg34 (Cvt pathway), Atg32 (mitophagy), Atg36 (pexophagy), and Atg30 (pexophagy), several of which have homologues in higher eukaryotes9. In mammalian cells, more than two dozen autophagy receptors were identified by the yeast two-hybrid system and proteomic approaches45,46. Not all proteins interacting with Atg8, LC3s or GABARAPs are autophagy receptors. An autophagy receptor is defined by its ability to bridge cargo and autophagosomal membrane, leading to the engulfment of cargo by the autophagic membrane (Fig. 2a). There is an additional class of proteins — autophagy adaptors — that also bind to members of the Atg8 family. However, they do not facilitate engulfment and subsequent degradation of cargo, but serve as anchor points for the autophagy machinery and regulate initiation, conjugation, transport and fusion of autophagosomes (Fig. 1b,c).
(a) In mitophagy, p62 recognizes poly-ubiquitylated (Ubn, purple) mitochondria destined for autophagosome degradation. Alternatively, mitochondria can also be bound by NIX directly without necessitating poly-ubiquitylation. In aggrephagy, poly-ubiquitylated protein aggregates are engulfed by the phagophore via p62 and NBR1. In xenophagy, the autophagic receptor proteins NDP52, p62 and OPTN recognize poly-ubiquitylated bacteria. In this case, the association of the LIR motif with LC3II can be enhanced by TBK1-dependent OPTN phosphorylation. Lastly, for pexophagy, p62 and NBR1 recruits the LC3II-positive phagophore to mono-ubiquitylated peroxisomes. (b) Domain structure of autophagy receptors involved in selective autophagy pathways, containing distinct ubiquitin-binding domains (blue) and LC3-interacting motifs (LIR, red).
The most prevalent autophagy targeting signal in mammals is the modification of cargos with Ub (ref. 4). The efficiency of ubiquitylation as a cargo signal was shown by ubiquitylation of endogenous proteins on the surface of mitochondria or pathogens such as Salmonella enterica, and by the artificial attachment of monoubiquitin to the cytosolic side of peroxisomes or long-lived cytosolic proteins47, which is both required and sufficient to induce their autophagic clearance. Indeed, most of the currently known autophagy receptors harbour both Ub-binding domains (UBDs) and LIRs (ref. 48) (Fig. 2b). On the other hand, selective autophagy in yeast does not use Ub modifications as a degradation signal, similarly to the mammalian autophagy receptors NIX or BNIP3, which are located at the outer mitochondrial membrane (reviewed in ref. 49; see below).
In higher organisms, autophagy receptors and mechanisms involved in mitophagy are probably the best studied so far. Depolarization of mitochondria leads to activation of PINK1 (PTEN-induced putative kinase protein 1) that accumulates at the outer mitochondrial membrane and phosphorylates numerous proteins50. Subsequent recruitment of the E3 ligase Parkin leads to multiple poly-Ub proteins51,52,53,54,55, which are recognized and clustered through polymerization by the autophagy receptor p62/SQSTM1 (sequestosome-1)56. Unexpectedly, p62 localizes to the phagophore even in the absence of LC3 interaction57. There is some evidence that self-oligomerization of p62 via the PB1 domain rather than binding to LC3 is critical for the translocation of p62 to the autophagosomal membrane57. Whereas the autophagic degradation of p62 loaded with selective cargo is dependent on the LIR–LC3 interaction, the initial targeting of p62 to autophagosomal membranes might not require the LIR domain.
These mechanisms for mitophagy have mostly been discovered under rather artificial conditions such as the treatment of cells with the mitochondrial uncoupling reagent CCCP and overexpression of Parkin, and so it has been questioned whether they take place in a physiological context. Whereas p62 is required for Parkin-mediated mitochondrial clustering as described, it may not be essential for efficient mitophagy under certain conditions56. In addition, the Ub ligase Parkin is dispensable for the clearance of dysfunctional mitochondria in mice58, indicating the presence of redundant, Ub-independent mitophagy pathways and/or additional autophagy receptors playing a redundant role for p62-mediated mitophagy56. In fact, during erythrocyte differentiation or in response to hypoxia, the removal of mitochondria can be mediated by NIX, BNIP3 and FUNDC1 (refs 59,60,61). All three receptors are mitochondrial outer-membrane proteins that can directly link mitochondria to autophagosomal membranes via their LIR domain in a Ub-independent way. Two reports hypothesize a regulatory function of these receptors: BNIP3 and NIX can trigger mitochondrial depolarization62 and NIX might also promote the mitochondrial translocation of Parkin, thereby contributing to Parkin–Ub–p62-mediated mitochondrial priming63. In addition, NIX regulates undamaged mitochondria in response to their energetic activity: high oxidative phosphorylation activity leads to the accumulation of the small GTPase Rheb on the mitochondrial outer membrane, where it physically interacts with NIX and enhances the autophagic removal of the undamaged organelle64.
An autophagy-receptor-independent mechanism of mitophagy was recently discovered in neuronal cells, and involves the inner mitochondrial membrane phospholipid cardiolipin. Mitophagy-stimulating substances caused cardiolipin externalization to the outer mitochondrial membrane, where it was directly recognized by LC3 (ref. 65) and facilitated mitophagy. In yeast, the removal of damaged mitochondria via autophagy generally happens independently of Ub. Comparable to NIX, the mitochondrial-anchored receptor Atg32 links damaged mitochondria to autophagosomal membranes66. It might be that Ub-independent selective autophagy pathways developed first, and then Ub-dependent pathways were developed by higher eukaryotes to speed up the autophagic process and introduce additional regulatory layers, such as recruitment of the Ub ligase Parkin by NIX (ref. 63) and phosphorylation of p62 and optineurin (OPTN), leading to increased Ub- and LC3-binding affinities, respectively67,68,69 (see below).
Another intriguing feature of autophagy receptors is their tendency to oligomerize, which further facilitates sequestration and clustering of the autophagic cargo. This formation of larger aggregates or inclusions (also called 'sequestosomes') could constitute a crucial step towards degradation by autophagy, as has been observed during the removal of defective mitochondria in mammals via p62 (refs 56,57) and during aggrephagy in mammalian cells45 and Drosophila brain70. Impairment in aggrephagy leads to formation of inclusion bodies containing poly-Ub protein aggregates that are hallmarks of neurodegenerative diseases such as Parkinson's or Alzheimer's disease. Genetic inactivation of p62 in mice or Ref(2)P (the Drosophila homologue of p62) in Drosophila models of neurodegenerative diseases revealed that p62 plays an important role in the formation of these inclusion bodies70,71 and contributes to their autolysosomal elimination72.
Selective autophagy receptors lack a clear specialization but often cooperate with each other in selecting a specific cargo: the autophagic receptor NBR1 (neighbour of BRCA1) interacts with p62 and plays an essential role in p62-dependent sequestration and degradation of aggregated proteins45, peroxisomes73, and mid body rings74. On the other hand, during xenophagy, p62 teams up with OPTN and NDP52 (refs 67,75,76) to facilitate the removal of invading bacteria. All of the receptors seem to have certain specificities, as they are recruited independently, and depletion of only one of them impairs bacterial clearance77. Notably, NDP52 is able to recognize invading bacteria not only through binding to Ub but also through binding to cytosolic lectin galectin-8 (ref. 75).
Little is known about the mechanisms governing selectivity in ribophagy and reticulophagy. Yeast ribophagy is dependent on the Ubp3/Bre5 deubiquitylation complex78 that also regulates the Cvt pathway through its action on Atg19 (ref. 79). Recent work indicates that there is an interplay between the Ubp3/Bre5 deubiquitylation and the E3 ligase Ltn1 in the regulation of ribophagy80.
We are only beginning to understand how cargos for selective autophagy are recognized and how this process is regulated. Further specific degradation signals and novel receptors are expected to be unveiled, such as the adaptor protein AP2 that was recently identified as a cargo receptor for aggregation-prone proteins involved in Alzheimer's disease81. Other examples of newly discovered regulators of autophagy are the proto-oncogene c-Cbl, which targets active Src in cancer cells via a newly identified LC3-interacting region to autophagosomes82, and cardiolipin, the only lipid-based autophagy linker between cargo and autophagosomes identified so far65.
Regulation of autophagy receptors
The activity of autophagy receptors is tightly regulated by inducible expression, spatial organization and cellular localization, and multiple post-translational modifications. To maintain a low basic protein level without accumulation in times of low autophagy, most receptors and scaffold proteins undergo constant degradation by autophagy even in an unloaded state.
More specific regulation occurs by post-translational modifications. As mentioned, selective autophagy in yeast does not use ubiquitylation to initiate degradation and therefore putative cargos need protection from constant undesired removal. In yeast, the known five receptors involved in cargo selection are localized in an inactive form on the cargo organelle and, following autophagy induction, interact with Atg8 and the basic autophagic scaffold Atg11 in a phosphorylation-dependent manner66,83,84. Atg11 can bind to cargo receptors as well as to components of the autophagy machinery. Thus the recruitment of cargo into autophagosomes not only depends on Atg8 but on a tripartite interaction of the receptor–cargo complex with the autophagosomal membrane (via Atg8) and core components of the autophagosomal machinery (via the scaffold Atg11). Both interactions are controlled by phosphorylation of the Atg11 and Atg8 binding sites of autophagy receptors by as-yet unknown kinases85. Studies in higher eukaryotes also identified scaffold proteins involved in specific types of autophagy, such as mammalian ALFY (ref. 86) and EPG-7 in C. elegans87. Moreover, phosphorylation as an inducing event of autophagy is conserved from yeast to mammals: phosphorylation of the LIR domain of BNIP3 activates its interaction with LC3s (ref. 88), and specific phosphorylation of the autophagy receptors p62 and OPTN results in increased affinity to Ub chains and LC3, shown to be essential for autophagic clearance of cytosolic salmonella67,68,69,89. It will be a challenge to identify the multiple post-translational modifications that govern autophagy selectivity and spatiotemporal regulation.
Secretory pathways use the autophagy machinery
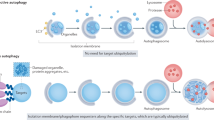
Processes involving molecules and mechanisms of the autophagic machinery in controlling secretory pathways have been recently identified. Such autophagy-dependent secretion allows the cell to secret proteins that lack a conventional secretory signal sequence. In yeast this is implicated by the formation of a 'compartment for unconventional protein secretion' (CUPS). Initiation of CUPS seems to occur independently of the PAS and autophagosomes dedicated for the Cvt pathway, but CUPS shares several features with omegasomes in mammalian cells: CUPS is enriched in PtdIns(3)P and Atg8, and forms close to the ER (ref. 90). Besides the involvement of specific Atg factors, CUPS is characterized by the dependence on Golgi re-assembly and stacking proteins (GRASPs). Illustrating the interdependency of membrane trafficking and unconventional autophagosomal pathways, IL-1β secretion in mammalian cells is inhibited during basal autophagy91,92 and upregulated following autophagy induction93. Additionally, the autophagosome contributes effector molecules (hydrolases) for the correct processing of IL-1 family members. The role of selective autophagy during the secretion of effector molecules can also be observed in osteoclasts (Fig. 3a). In these specialized cells, secretory lysosomes fuse with the plasma membrane juxtaposed to bone to create the ruffled border, a cellular organelle responsible for resorbing the osseous tissue in osteoclasts. The formation of the ruffled border relies on several autophagic proteins including ATG5, ATG7, ATG4B and LC3 (ref. 94).
(a) Formation of the ruffled border in osteoclasts and secretion of lysosomal proteins lead to resorption of bone tissue. In this process, cathepsin-K-containing lysosomes are recruited to and subsequently fuse with the plasma membrane in an LC3II-dependent manner. (b) The formation of a LC3II-positive single-layer membrane around bacteria or dead cells results in phagocytosis. Subsequent fusion with a lysosome forms the phagolysosome, where the cargo is then degraded.
Autophagy-dependent secretion has also been implicated in the pathology of Crohn's disease, a polygenic inflammatory disease involving the small intestine. Mutations in several autophagy-related genes, including ATG16L1 and NOD2, are associated with Crohn's disease. NOD2 is a pattern-recognition-receptor (PRR) that senses invading pathogens and induces their autophagic elimination at the plasma membrane95 (Fig. 3b). Moreover, ATG16L1 as well as ATG5 and ATG7 are crucial for the secretion of granule contents of the Paneth cell, a specialized secretory epithelial cell96. It remains unclear how the interplay between the secretory system and autophagy machinery affects the final fate of the transport cargo — particularly in terms of what determines the fusion events between different intracellular membranes along these pathways, and what mechanisms engage the autophagy components in secretion pathways rather than promoting cargo transport to the lysosome for degradation.
Overlapping roles of autophagy and intracellular trafficking
Autophagy shares many central features with membrane trafficking pathways, including membrane remodelling, the budding of vesicles, their directed movement along microtubule tracks, and the homo- as well as heterotypic fusion of vesicles6. As both types of transport systems rely on the same functional modules, membrane trafficking pathways can be regulated by autophagy and, vice versa, several proteins controlling intracellular transport can modulate autophagy.
A family of important regulators of endocytic trafficking are the RabGAPs, which control the GTPase activity of Rab proteins (reviewed in ref. 97). Several RabGAPs have been recently reported to play an important role in autophagy — such as TBC1D14, which mediates autophagosome growth by regulating Rab11 and recycling endosomes, and interacting at the same time with ULK1 and TBC1D25, which mediates fusion of autophagosome and lysosomes98,99. In addition, TBC1D5 regulates both retrograde transport (from endosomes to the trans-Golgi network, TGN) and autophagy flux100,101. The late-endosome marker TBC1D15, which is necessary for the recruitment of the tethering complex HOPS, associates with mitochondria via the fission protein Fis1 and contributes to their selective removal by autophagy102. In addition, the transport between the Golgi complex and ER depends on coatomer subunits α-, β- and ε-COP (refs. 103,104,105), which are at the same time critical for autophagy, as depletion of β′-, β- or α-COP results in an accumulation of autophagosomes and amphisomes106.
On the other hand, mammalian Atg8 family members are implicated in the regulation of the membrane fusion machinery107, which mediates the recognition and fusion of a transport vesicle and its target membrane independently of autophagy. Soluble (not lipidated) GATE-16 was shown to modulate intra-Golgi transport as well as post-mitotic Golgi re-assembly by coupling N-ethylmaleimide-sensitive factor (NSF) activity and SNAREs activation, and by regulating NSF function108,109. GATE-16 preferentially binds the unpaired form of a mitochondrial v-SNARE and interferes with the binding to its cognate t-SNARE (refs 109,110), thus preventing the assembly of SNARE complexes at inappropriate times109. Also, an interaction of GABARAP with NSF is required for the ability of GABARAP to regulate trafficking of the GABAA receptor, the predominant inhibitory neurotransmitter receptor in the brain111,112.
Finally, selective autophagy and the endolysosomal sorting of plasma membrane proteins employ the same strategy of cargo delivery. Both systems use Ub as a degradation signal and Ub-binding proteins as specific receptors that link cargo destined for lysosomal degradation. The plasma membrane can also directly contribute to the autophagy pathway by promoting internalization and trafficking of ATG9- and ATG16L1-positive vesicles needed for autophagosome formation40,100. The mechanisms governing endocytosis and lysosomal sorting have been extensively studied, and more principles that also apply to selective autophagy might be unravelled in the future.
Concluding remarks and future directions
The plethora of processes regulated by selective autophagy is indicative of its unique importance for cellular homeostasis in health and disease. Selective autophagy clearly goes beyond a simple 'eating' process involving double membranes, as it also directly or indirectly controls secretion and other intracellular transport processes. Apparently, lipid-conjugated Ub-like modifiers such as Atg8 in yeast and LC3/GABARAPs in mammals provide a regulatory network that combines the characteristics of the Ub system with the specialized functions of cellular membranes, making it an extremely versatile and generally applicable regulatory process. We have just begun to understand the molecular basis of how these adaptive networks cooperate to achieve the tremendous diversity observed in cells. Although several key findings have unravelled the basic principles of autophagosome formation and cargo selection by autophagic receptors, many questions remain. These are related to the molecular events that govern the very early steps of selective autophagy induction, the source of autophagic membranes, and their interplay with other endosomal membranous systems. Selective autophagy pathways can also positively and negatively impact human disorders including inflammation, infectious and neurodegenerative diseases, and cancer. Consequently, this evidence has attracted increased attention from the pharmaceutical industry for the development of the next generation of drugs.
References
Mizushima, N., Yoshimori, T. & Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell Dev. Biol. 27, 107–132 (2011).
Levine, B., Mizushima, N. & Virgin, H. W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011).
Yang, Z. & Klionsky, D. J. Eaten alive: a history of macroautophagy. Nat. Cell Biol. 12, 814–822 (2010).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Kraft, C., Peter, M. & Hofmann, K. Selective autophagy: ubiquitin-mediated recognition and beyond. Nat. Cell Biol. 12, 836–841 (2010).
Lamb, C. A., Yoshimori, T. & Tooze, S. A. The autophagosome: origins unknown, biogenesis complex. Nat. Rev. Mol. Cell Biol. 14, 759–774 (2013).
Thumm, M. et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 349, 275–280 (1994).
Tsukada, M. & Ohsumi, Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333, 169–174 (1993).
Meijer, W. H., van der Klei, I. J., Veenhuis, M. & Kiel, J. A. ATG genes involved in non-selective autophagy are conserved from yeast to man, but the selective Cvt and pexophagy pathways also require organism-specific genes. Autophagy 3, 106–116 (2007).
Budovskaya, Y. V., Stephan, J. S., Deminoff, S. J. & Herman, P. K. An evolutionary proteomics approach identifies substrates of the cAMP-dependent protein kinase. Proc. Natl Acad. Sci. USA 102, 13933–13938 (2005).
Chang, Y. Y. & Neufeld, T. P. An Atg1/Atg13 complex with multiple roles in TOR-mediated autophagy regulation. Mol. Biol. Cell 20, 2004–2014 (2009).
Ganley, I. G. et al. ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J. Biol. Chem. 284, 12297–12305 (2009).
Hoyer-Hansen, M. & Jaattela, M. AMP-activated protein kinase: a universal regulator of autophagy? Autophagy 3, 381–383 (2007).
Samari, H. R. & Seglen, P. O. Inhibition of hepatocytic autophagy by adenosine, aminoimidazole-4-carboxamide riboside, and N6-mercaptopurine riboside. Evidence for involvement of amp-activated protein kinase. J. Biol. Chem. 273, 23758–23763 (1998).
Russell, R. C. et al. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat. Cell Biol. 15, 741–750 (2013).
Krick, R., Tolstrup, J., Appelles, A., Henke, S. & Thumm, M. The relevance of the phosphatidylinositolphosphat-binding motif FRRGT of Atg18 and Atg21 for the Cvt pathway and autophagy. FEBS Lett. 580, 4632–4638 (2006).
Taguchi-Atarashi, N. et al. Modulation of local PtdIns3P levels by the PI phosphatase MTMR3 regulates constitutive autophagy. Traffic 11, 468–478 (2010).
Kundu, M. et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 112, 1493–1502 (2008).
Alers, S. et al. Atg13 and FIP200 act independently of Ulk1 and Ulk2 in autophagy induction. Autophagy 7, 1423–1433 (2011).
Martinez, J. et al. Microtubule-associated protein 1 light chain 3 alpha (LC3)-associated phagocytosis is required for the efficient clearance of dead cells. Proc. Natl Acad. Sci. USA 108, 17396–17401 (2011).
Florey, O., Kim, S. E., Sandoval, C. P., Haynes, C. M. & Overholtzer, M. Autophagy machinery mediates macroendocytic processing and entotic cell death by targeting single membranes. Nat. Cell Biol. 13, 1335–1343 (2011).
Slobodkin, M. R. & Elazar, Z. The Atg8 family: multifunctional ubiquitin-like key regulators of autophagy. Essays Biochem. 55, 51–64 (2013).
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178 (2007).
Weidberg, H. et al. LC3 and GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis. EMBO J. 29, 1792–1802 (2010).
Kim, J., Huang, W. P., Stromhaug, P. E. & Klionsky, D. J. Convergence of multiple autophagy and cytoplasm to vacuole targeting components to a perivacuolar membrane compartment prior to de novo vesicle formation. J. Biol. Chem. 277, 763–773 (2002).
Suzuki, K. et al. The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO J. 20, 5971–5981 (2001).
Mari, M., Tooze, S. A. & Reggiori, F. The puzzling origin of the autophagosomal membrane. F1000 Biol. Rep. 3, 25 (2011).
Dupont, N. et al. Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Curr. Biol. 24, 609–620 (2014).
Hailey, D. W. et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141, 656–667 (2010).
Hamasaki, M. et al. Autophagosomes form at ER–mitochondria contact sites. Nature 495, 389–393 (2013).
Vance, J. E. & Tasseva, G. Formation and function of phosphatidylserine and phosphatidylethanolamine in mammalian cells. Biochim. Biophys. Acta 1831, 543–554 (2013).
Hayashi-Nishino, M. et al. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat. Cell Biol. 11, 1433–1437 (2009).
Yla-Anttila, P., Vihinen, H., Jokitalo, E. & Eskelinen, E. L. 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy 5, 1180–1185 (2009).
De Brito, O. M. & Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456, 605–610 (2008).
Axe, E. L. et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, 685–701 (2008).
Polson, H. E. et al. Mammalian Atg18 (WIPI2) localizes to omegasome-anchored phagophores and positively regulates LC3 lipidation. Autophagy 6, 506–522 (2010).
Yang, J. Y. & Yang, W. Y. Bit-by-bit autophagic removal of parkin-labelled mitochondria. Nat. Commun. 4, 2428 (2013).
Giordano, F. et al. PI(4,5)P2-dependent and Ca2+-regulated ER–PM interactions mediated by the extended synaptotagmins. Cell 153, 1494–1509 (2013).
Stefan, C. J., Manford, A. G. & Emr, S. D. ER–PM connections: sites of information transfer and inter-organelle communication. Curr. Opin. Cell Biol. 25, 434–442 (2013).
Puri, C., Renna, M., Bento, C. F., Moreau, K. & Rubinsztein, D. C. Diverse autophagosome membrane sources coalesce in recycling endosomes. Cell 154, 1285–1299 (2013).
Yamamoto, H. et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J. Cell Biol. 198, 219–233 (2012).
Wang, J. et al. Ypt1 recruits the Atg1 kinase to the preautophagosomal structure. Proc. Natl Acad. Sci. USA 110, 9800–9805 (2013).
Kakuta, S. et al. Atg9 vesicles recruit vesicle-tethering proteins Trs85 and Ypt1 to the autophagosome formation site. J. Biol. Chem. 287, 44261–44269 (2012).
Lynch-Day, M. A. et al. Trs85 directs a Ypt1 GEF, TRAPPIII, to the phagophore to promote autophagy. Proc. Natl Acad. Sci. USA 107, 7811–7816 (2010).
Kirkin, V. et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol. Cell 33, 505–516 (2009).
Behrends, C., Sowa, M. E., Gygi, S. P. & Harper, J. W. Network organization of the human autophagy system. Nature 466, 68–76 (2010).
Kim, P. K., Hailey, D. W., Mullen, R. T. & Lippincott-Schwartz, J. Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl Acad. Sci. USA 105, 20567–20574 (2008).
Wild, P. M., D. McEwan & Dikic, I. The LC3 interactome at a glance. J. Cell Sci. 127, 3–9 (2014).
Zhang, J. & Ney, P. A. Role of BNIP3 and NIX in cell death, autophagy, and mitophagy. Cell Death Differ. 16, 939–946 (2009).
Chen, Y. & Dorn, G. W. 2nd PINK1-phosphorylated mitofusin 2 is a Parkin receptor for culling damaged mitochondria. Science 340, 471–475 (2013).
Sarraf, S. A. et al. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496, 372–376 (2013).
van Wijk, S. J. et al. Fluorescence-based sensors to monitor localization and functions of linear and K63-linked ubiquitin chains in cells. Mol. Cell 47, 797–809 (2012).
Narendra, D., Tanaka, A., Suen, D. F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 12, 119–131 (2010).
Gegg, M. E. et al. Mitofusin 1 and mitofusin 2 are ubiquitinated in a PINK1/parkin-dependent manner upon induction of mitophagy. Hum. Mol. Genet. 19, 4861–4870 (2010).
Narendra, D., Kane, L. A., Hauser, D. N., Fearnley, I. M. & Youle, R. J. p62/SQSTM1 is required for Parkin-induced mitochondrial clustering but not mitophagy; VDAC1 is dispensable for both. Autophagy 6, 1090–1106 (2010).
Itakura, E. & Mizushima, N. p62 Targeting to the autophagosome formation site requires self-oligomerization but not LC3 binding. J. Cell Biol. 192, 17–27 (2011).
Sterky, F. H., Lee, S., Wibom, R., Olson, L. & Larsson, N. G. Impaired mitochondrial transport and Parkin-independent degeneration of respiratory chain-deficient dopamine neurons in vivo. Proc. Natl Acad. Sci. USA 108, 12937–12942 (2011).
Mazure, N. M. & Pouyssegur, J. Hypoxia-induced autophagy: cell death or cell survival? Curr. Opin. Cell Biol. 22, 177–180 (2010).
Novak, I. et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 11, 45–51 (2010).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Sandoval, H. et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 454, 232–235 (2008).
Ding, W. X. et al. Nix is critical to two distinct phases of mitophagy, reactive oxygen species-mediated autophagy induction and Parkin–ubiquitin–p62-mediated mitochondrial priming. J. Biol. Chem. 285, 27879–27890 (2010).
Melser, S. et al. Rheb regulates mitophagy induced by mitochondrial energetic status. Cell Metab. 17, 719–730 (2013).
Chu, C. T., Bayir, H. & Kagan, V. E. LC3 binds externalized cardiolipin on injured mitochondria to signal mitophagy in neurons: Implications for Parkinson disease. Autophagy 10, 376–378 (2014).
Okamoto, K., Kondo-Okamoto, N. & Ohsumi, Y. Mitochondria-anchored receptor Atg32 mediates degradation of mitochondria via selective autophagy. Dev. Cell 17, 87–97 (2009).
Wild, P. et al. Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333, 228–233 (2011).
Matsumoto, G., Wada, K., Okuno, M., Kurosawa, M. & Nukina, N. Serine 403 phosphorylation of p62/SQSTM1 regulates selective autophagic clearance of ubiquitinated proteins. Mol. Cell 44, 279–289 (2011).
Pilli, M. et al. TBK-1 promotes autophagy-mediated antimicrobial defense by controlling autophagosome maturation. Immunity 37, 223–234 (2012).
Nezis, I. P. et al. Ref(2)P, the Drosophila melanogaster homologue of mammalian p62, is required for the formation of protein aggregates in adult brain. J. Cell Biol. 180, 1065–1071 (2008).
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163 (2007).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Deosaran, E. et al. NBR1 acts as an autophagy receptor for peroxisomes. J. Cell Sci. 126, 939–952 (2013).
Isakson, P. et al. TRAF6 mediates ubiquitination of KIF23/MKLP1 and is required for midbody ring degradation by selective autophagy. Autophagy 9, 1955–1964 (2013).
Thurston, T. L., Wandel, M. P., von Muhlinen, N., Foeglein, A. & Randow, F. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature 482, 414–418 (2012).
Mostowy, S. et al. p62 and NDP52 proteins target intracytosolic Shigella and Listeria to different autophagy pathways. J. Biol. Chem. 286, 26987–26995 (2011).
Cemma, M., Kim, P. K. & Brumell, J. H. The ubiquitin-binding adaptor proteins p62/SQSTM1 and NDP52 are recruited independently to bacteria-associated microdomains to target Salmonella to the autophagy pathway. Autophagy 7, 341–345 (2011).
Kraft, C., Deplazes, A., Sohrmann, M. & Peter, M. Mature ribosomes are selectively degraded upon starvation by an autophagy pathway requiring the Ubp3p/Bre5p ubiquitin protease. Nat. Cell Biol. 10, 602–610 (2008).
Baxter, B. K. et al. Atg19p ubiquitination and the cytoplasm to vacuole trafficking pathway in yeast. J. Biol. Chem. 280, 39067–39076 (2005).
Ossareh-Nazari, B. et al. Ubiquitylation by the Ltn1 E3 ligase protects 60S ribosomes from starvation-induced selective autophagy. J. Cell Biol. 204, 909–917 (2014).
Tian, Y., Chang, J. C., Fan, E. Y., Flajolet, M. & Greengard, P. Adaptor complex AP2/PICALM, through interaction with LC3, targets Alzheimer's APP-CTF for terminal degradation via autophagy. Proc. Natl Acad. Sci. USA 110, 17071–17076 (2013).
Sandilands, E. et al. Autophagic targeting of Src promotes cancer cell survival following reduced FAK signalling. Nat. Cell Biol. 14, 51–60 (2012).
Kanki, T., Wang, K., Cao, Y., Baba, M. & Klionsky, D. J. Atg32 is a mitochondrial protein that confers selectivity during mitophagy. Dev. Cell 17, 98–109 (2009).
Farre, J. C., Manjithaya, R., Mathewson, R. D. & Subramani, S. PpAtg30 tags peroxisomes for turnover by selective autophagy. Dev. Cell 14, 365–376 (2008).
Farre, J. C., Burkenroad, A., Burnett, S. F. & Subramani, S. Phosphorylation of mitophagy and pexophagy receptors coordinates their interaction with Atg8 and Atg11. EMBO Rep. 14, 441–449 (2013).
Filimonenko, M. et al. The selective macroautophagic degradation of aggregated proteins requires the PI3P-binding protein Alfy. Mol. Cell 38, 265–279 (2010).
Lin, L. et al. The scaffold protein EPG-7 links cargo-receptor complexes with the autophagic assembly machinery. J. Cell Biol. 201, 113–129 (2013).
Zhu, Y. et al. Modulation of serines 17 and 24 in the LC3-interacting region of Bnip3 determines pro-survival mitophagy versus apoptosis. J. Biol. Chem. 288, 1099–1113 (2013).
Rogov, V. V. et al. Structural basis for phosphorylation-triggered autophagic clearance of Salmonella. Biochem. J. 454, 459–466 (2013).
Bruns, C., McCaffery, J. M., Curwin, A. J., Duran, J. M. & Malhotra, V. Biogenesis of a novel compartment for autophagosome-mediated unconventional protein secretion. J. Cell Biol. 195, 979–992 (2011).
Nakahira, K. et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 12, 222–230 (2011).
Zhou, R., Yazdi, A. S., Menu, P. & Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 469, 221–225 (2011).
Dupont, N. et al. Autophagy-based unconventional secretory pathway for extracellular delivery of IL-1beta. EMBO J. 30, 4701–4711 (2011).
DeSelm, C. J. et al. Autophagy proteins regulate the secretory component of osteoclastic bone resorption. Dev. Cell 21, 966–974 (2011).
Travassos, L. H. et al. Nod1 and Nod2 direct autophagy by recruiting ATG16L1 to the plasma membrane at the site of bacterial entry. Nat. Immunol. 11, 55–62 (2010).
Cadwell, K. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008).
Langemeyer, L. & Barr, F. A. Analysis of Rab GTPases. Curr. Prot. Cell Biol. 57, 15.18 (2012).
Itoh, T., Kanno, E., Uemura, T., Waguri, S. & Fukuda, M. OATL1, a novel autophagosome-resident Rab33B-GAP, regulates autophagosomal maturation. J. Cell Biol. 192, 839–853 (2011).
Longatti, A. et al. TBC1D14 regulates autophagosome formation via Rab11- and ULK1-positive recycling endosomes. J. Cell Biol. 197, 659–675 (2012).
Popovic, D. et al. Rab GTPase-activating proteins in autophagy: regulation of endocytic and autophagy pathways by direct binding to human ATG8 modifiers. Mol. Cell. Biol. 32, 1733–1744 (2012).
Popovic, D. & Dikic, I. TBC1D5 and the AP2 complex regulate ATG9 trafficking and initiation of autophagy. EMBO Rep. http://dx/doi.org/10.1002/embr.201337995 (2014).
Yamano, K., Fogel, A. I., Wang, C., van der Bliek, A. M. & Youle, R. J. Mitochondrial Rab GAPs govern autophagosome biogenesis during mitophagy. eLife 3, e01612 (2014).
Gu, F., Aniento, F., Parton, R. G. & Gruenberg, J. Functional dissection of COP-I subunits in the biogenesis of multivesicular endosomes. J. Cell Biol. 139, 1183–1195 (1997).
Daro, E., Sheff, D., Gomez, M., Kreis, T. & Mellman, I. Inhibition of endosome function in CHO cells bearing a temperature-sensitive defect in the coatomer (COPI) component epsilon-COP. J. Cell Biol. 139, 1747–1759 (1997).
Nickel, W., Brugger, B. & Wieland, F. T. Vesicular transport: the core machinery of COPI recruitment and budding. J. Cell Sci. 115, 3235–3240 (2002).
Razi, M., Chan, E. Y. & Tooze, S. A. Early endosomes and endosomal coatomer are required for autophagy. J. Cell Biol. 185, 305–321 (2009).
Elazar, Z., Scherz-Shouval, R. & Shorer, H. Involvement of LMA1 and GATE-16 family members in intracellular membrane dynamics. Biochim. Biophys. Acta 1641, 145–156 (2003).
Sagiv, Y., Legesse-Miller, A., Porat, A. & Elazar, Z. GATE-16, a membrane transport modulator, interacts with NSF and the Golgi v-SNARE GOS-28. EMBO J. 19, 1494–1504 (2000).
Muller, J. M. et al. Sequential SNARE disassembly and GATE-16–GOS-28 complex assembly mediated by distinct NSF activities drives Golgi membrane fusion. J. Cell Biol. 157, 1161–1173 (2002).
Subramaniam, V. N., Loh, E. & Hong, W. N-Ethylmaleimide-sensitive factor (NSF) and alpha-soluble NSF attachment proteins (SNAP) mediate dissociation of GS28-syntaxin 5 Golgi SNAP receptors (SNARE) complex. J. Biol. Chem. 272, 25441–25444 (1997).
Kittler, J. T. et al. The subcellular distribution of GABARAP and its ability to interact with NSF suggest a role for this protein in the intracellular transport of GABA(A) receptors. Mol. Cell. Neurosci. 18, 13–25 (2001).
Wang, H., Bedford, F. K., Brandon, N. J., Moss, S. J. & Olsen, R. W. GABA(A)-receptor-associated protein links GABA(A) receptors and the cytoskeleton. Nature 397, 69–72 (1999).
Acknowledgements
We apologize to all scientists whose important contribution was not referenced in this Review due to space limitations. We would like to thank Daniela Höller, Kerstin Koch, Doris Popovic, David McEwan, Ligia Carinha Gomes and Jaime Lopez Mosqueda for critical reading and comments on the manuscript. Research in the Ernst laboratory is supported by the LOEWE Ub-Net grant and the Cluster of Excellence 'Macromolecular Complexes' of the Goethe University of Frankfurt, and in the Dikic laboratory by grants from the Deutsche Forschungsgemeinschaft (DI 931/3-1), the LOEWE Ub-Net and CGT, and the European Research Council (ERC) grant agreement no. 250241-LineUb.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Stolz, A., Ernst, A. & Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat Cell Biol 16, 495–501 (2014). https://doi.org/10.1038/ncb2979
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2979
This article is cited by
-
Arginyltransferase 1 modulates p62-driven autophagy via mTORC1/AMPk signaling
Cell Communication and Signaling (2024)
-
Carnosine regulation of intracellular pH homeostasis promotes lysosome-dependent tumor immunoevasion
Nature Immunology (2024)
-
PRKAA2, MTOR, and TFEB in the regulation of lysosomal damage response and autophagy
Journal of Molecular Medicine (2024)
-
The role of NEDD4 related HECT-type E3 ubiquitin ligases in defective autophagy in cancer cells: molecular mechanisms and therapeutic perspectives
Molecular Medicine (2023)
-
Deficiency of cancer/testis antigen gene CT55 causes male infertility in humans and mice
Cell Death & Differentiation (2023)