Abstract
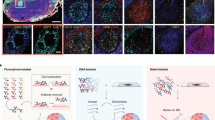
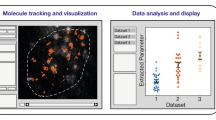
Temporal and spatial regulation of proteins contributes to function. We describe a multidimensional microscopic robot technology for high-throughput protein colocalization studies that runs cycles of fluorescence tagging, imaging and bleaching in situ. This technology combines three advances: a fluorescence technique capable of mapping hundreds of different proteins in one tissue section or cell sample; a method selecting the most prominent combinatorial molecular patterns by representing the data as binary vectors; and a system for imaging the distribution of these protein clusters in a so-called toponome map. By analyzing many cell and tissue types, we show that this approach reveals rules of hierarchical protein network organization, in which the frequency distribution of different protein clusters obeys Zipf's law, and state-specific lead proteins appear to control protein network topology and function. The technology may facilitate the development of diagnostics and targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Collins, F.S., Green, E.D., Guttmacher, A.E. & Guyer, M.S. US National Human Genome Research Institute. A vision for the future of genomics research. Nature 422, 835–847 (2003).
Schubert, W. Topological proteomics, toponomics, MELK technology. Adv. Biochem. Engin. Biotechnol. 83, 189–209 (2003).
Herman, B., Krishnan, R.V. & Centonze, V.E. Microscopic analysis of fluorescence resonance energy transfer (FRET). Methods Mol. Biol. 261, 351–370 (2004).
Jares-Erijman, E.A. & Jovin, T.M. FRET imaging. Nat. Biotechnol. 21, 1387–1395 (2003).
Ganesan, S., Ameer-Berg, S.M., Ng, T.T., Vojnovic, B. & Wouters, F.S. A dark yellow fluorescent protein (YFP)-based resonance energy-accepting chromoprotein (REACh) for Forster resonance energy transfer with GFP. Proc. Natl. Acad. Sci. USA 103, 4089–4094 (2006).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Perfetto, S.P., Chattopadhyay, P.K. & Roederer, M. Seventeen-colour flow cytometry: unravelling the immune system. Nat. Rev. Immunol. 8, 648–655 (2004).
Pacholski, M.L. & Winograd, N. Imaging with mass spectrometry. Chem. Rev. 99, 2977–3006 (1999).
Stoeckli, M., Chaurand, P., Hallahan, D.E. & Caprioli, R.M. Imaging mass spectrometry: a new technology for the analysis of protein expression in mammalian tissues. Nat. Med. 7, 493–496 (2001).
Bray, D. Molecular networks: the top-down view. Science 301, 1864–1865 (2003).
Han, J.-D.J. et al. Evidence for dynamically organized modularity in the yeast protein-protein interaction network. Nature 430, 88–93 (2004).
Dress, A.W.M., Lokot, T., Pustylnikov, L.D. & Schubert, W. Poisson numbers and poisson distributions in subset surprisology. Ann. Combinatorics 8, 475–485 (2004).
Gottlieb, A.B. et al. TNF inhibition rapidly down-regulates multiple proinflammatory pathways in psoriasis plaques. J. Immunol. 175, 2721–2729 (2005).
Boguniewicz, M. & Leung, D.Y. Atopic dermatitis. J. Allergy Clin. Immunol. 117, S475–480 (2006).
Böckelmann, R. et al. Suprabasal overexpression of the hsRPB7 gene in psoriatic epidermis as identified by a reverse transcriptase-polymerase chain reaction differential display model comparing psoriasis plaque tissue with peritonsillar mucosa. Am. J. Pathol. 158, 367–372 (2001).
Zipf, G.K. . Human Behaviour and the Principle of Least Effort (Addison-Wesley, Cambridge MA, 1949).
Hoyle, D.C., Rattray, M., Jupp, R. & Brass, A. Making sense of microarray data distributions. Bioinformatics 18, 576–584 (2002).
Ferrer i Cancho, R. & Sole, R.V. Least effort and the origins of scaling in human language. Proc. Natl. Acad. Sci. U.S.A. 100, 788–791 (2003).
Newman, M.E.J. Power laws, Pareto distributions and Zipf's law. Contemp. Phys. 46, 323–351 (2005).
Bennett, G.J. & Xie, Y.K. A peripheral mononeuropathy in rat that produces disorders of pain sensation like those seen in man. Pain 33, 87–107 (1988).
Garry, E.M. et al. Neuropathic sensitization of behavioral reflexes and spinal NMDA receptor/CaM kinase II interactions are disrupted in PSD-95 mutant mice. Curr. Biol. 13, 321–328 (2003).
Chen, H., Roques, B.P. & Fournie-Zaluski, M.C. Design of the first highly potent and selective aminopeptidase N (EC 3.4.11.2) inhibitor. Bioorg. Med. Chem. Lett. 9, 1511–1516 (1999).
Kehlen, A., Lendeckel, U., Dralle, H., Langner, J. & Hoang-Vu, C. Biological significance of aminopeptidase N/CD13 in thyroid carcinomas. Cancer Res. 63, 8500–8506 (2003).
Conrad, C. et al. Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 14, 1130–1136 (2004).
Chen, X. & Murphy, R.F. Objective clustering of proteins based on subcellular location patterns. J. Biomed. Biotechol. 2, 87–95 (2005).
Uhlen, M. et al. A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol. Cell Proteomics 4, 1920–1932 (2005).
Koonin, E.V., Wolf, Y.I. & Karev, G.P. The structure of the protein universe and genome evolution. Nature 420, 218–223 (2002).
Jeong, H., Mason, S.P., Barabasi, A.L. & Oltvai, Z.N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001).
Schubert, W. Method of blocking cytotoxic activity in patients with amyotrophic lateral sclerosis using antibodies to FcγRIII. US patent no. 6,638,506 (first published as international patent application WO 99/29731, 1999).
Mohamed, H.A. et al. Immunoglobulin Fc gamma receptor promotes immunoglobulin uptake, immunoglobulin-mediated calcium increase, and neurotransmitter release in motor neurons. J. Neurosci. Res. 69, 110–116 (2002).
Conchello, J.A. & McNally, J.G. Fast regularization technique for expectation maximization algorithm for computational optical sectioning microscopy. in Three-Dimensional Microscopy: Image Acquisition and Processing Cogswell, C.J., Kino, G.S. & Wilson, T. (eds.) Proc. SPIE 2655, 199–208 (1996).
Acknowledgements
We thank K. Neubert for supplying bis-ala-rhodamine 110, B.P. Roques for RB3014, and A. Borissenko and P. Karcher for computational help. The study was supported by the Bundesministerium für Bildung und Forschung (NBL-3 FKZ: 01ZZ0107, 01ZZ0407; NGFN2 01 GR 0446, CELLECT 0312844, BioChance 0312452), the DFG (627/1-8; INK “Bildinformation”) and Land Saxony-Anhalt. A.W.M. Dress thanks the Center for Combinatorics at Nankai University for hospitality during the preparation of this manuscript. We are grateful for animal tissue provided by F. Hucho/O. Bogen and fruitful discussions, and appreciate critical reading by A. Leech. We thank Anja Bastian, Katrin Brennecke and Franziska Böckelmann for technical assistance.
Author information
Authors and Affiliations
Contributions
W.S. invented MELC and designed the study. M.F. and M.B. closely cooperated with L.P., A.J.P. and W.S. to develop robots and corresponding biological assays. A.W.M.D. performed mathematical image analyses. W.S., M.F. and M.B. designed, performed and interpreted data of multidimensional analyses of protein locations. CCI tissue material was analyzed and interpreted by W.S. and corresponding statistical motif analyses were done by L.P. Experiments with rhabdomyosarcoma cells were performed by M.F. and analyzed and interpreted in close cooperation with M.B., L.P. and W.S. 3D-images were generated by M.B. in close cooperation with W.S. W.S., M.B. and M.F. prepared the corresponding parts of the manuscript. B.B., R.B. and H.G. closely cooperated with A.J.P., Y.M. and L.P. in the dermatological and skin tissue–related investigations. In detail, B.B., A.J.P., R.B. and H.G. designed these clinico-experimental studies. B.B. and H.G. performed the clinical study part. R.B. did most of the accompanying conventional immunohistology. A.J.P., Y.M. and L.P. did the MELC experiments. R.B. binarized most of the MELC images and performed statistics. L.P. contributed all computer analyses and visualizations. B.B., A.J.P., L.P., R.B., Y.M. and H.G. interpreted these data and prepared the corresponding parts of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that their competing financial interests in this work are too numerous to itemize.
Supplementary information
Supplementary Fig. 1
Principle of the MELC Procedure. (PDF 713 kb)
Supplementary Fig. 2
Control Experiment. (PDF 467 kb)
Supplementary Fig. 3
Performance and Validation Tests for MELC. (PDF 147 kb)
Supplementary Fig. 4
Toponome Map of the Upper Dermis. (PDF 80 kb)
Supplementary Fig. 5
Colocalization Map of Keratinocyte Stem Cells. (PDF 851 kb)
Supplementary Fig. 6
Efalizumab Binding Sites in Skin. (PDF 13577 kb)
Supplementary Table 1
MELC Tag Libraries. (PDF 22 kb)
Supplementary Table 2
CMP and Motif Lists. (PDF 1523 kb)
Supplementary Table 3
MELC Toponome Data Base. (PDF 10 kb)
Supplementary Table 4
CD11a as Lead Protein in Inflammatory Skin Disease. (PDF 24 kb)
Supplementary Table 5
Demographic Data of Individuals Subjected to MELC Analysis of Skin Biopsies. (PDF 17 kb)
Supplementary Video 1
Exploring a Human Hepatocyte at 3D. (MOV 8902 kb)
Supplementary Video 2
3D Rocking Images of Nervous and Muscle Tissue. (MOV 9781 kb)
Supplementary Video 3
100 MELC Cycles on Skin Tissue and 3D Skin Imaging. (MOV 9064 kb)
Supplementary Video 4
Lead Protein Inhibition and Time-resolved Toponomics of Rhabdomyosarcoma Cells. (MOV 8291 kb)
Rights and permissions
About this article
Cite this article
Schubert, W., Bonnekoh, B., Pommer, A. et al. Analyzing proteome topology and function by automated multidimensional fluorescence microscopy. Nat Biotechnol 24, 1270–1278 (2006). https://doi.org/10.1038/nbt1250
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1250
This article is cited by
-
MarShie: a clearing protocol for 3D analysis of single cells throughout the bone marrow at subcellular resolution
Nature Communications (2024)
-
Quantitative multiplex immunohistochemistry with colorimetric staining (QUIVER) may still benefit from MILAN
Acta Neuropathologica Communications (2023)
-
Quantitative multiplexed imaging technologies for single-cell analysis to assess predictive markers for immunotherapy in thoracic immuno-oncology: promises and challenges
British Journal of Cancer (2023)
-
DNA-barcoded signal amplification for imaging mass cytometry enables sensitive and highly multiplexed tissue imaging
Nature Methods (2023)
-
CellSighter: a neural network to classify cells in highly multiplexed images
Nature Communications (2023)