Abstract
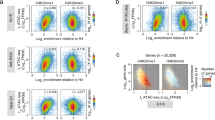
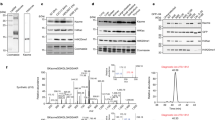
Lysine methylation of histones in vivo occurs in three states: mono-, di- and tri-methyl1. Histone H3 has been found to be di-methylated at lysine 4 (K4) in active euchromatic regions but not in silent heterochromatic sites2. Here we show that the Saccharomyces cerevisiae Set1 protein can catalyse di- and tri-methylation of K4 and stimulate the activity of many genes. Using antibodies that discriminate between the di- and tri-methylated state of K4 we show that di-methylation occurs at both inactive and active euchromatic genes, whereas tri-methylation is present exclusively at active genes. It is therefore the presence of a tri-methylated K4 that defines an active state of gene expression. These findings establish the concept of methyl status as a determinant for gene activity and thus extend considerably the complexity of histone modifications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Strahl, B. D., Ohba, R., Cook, R. G. & Allis, C. D. Methylation of histone H3 at lysine 4 is highly conserved and correlates with transcriptionally active nuclei in Tetrahymena. Proc. Natl Acad. Sci. USA 96, 14967–14972 (1999)
Noma, K., Allis, C. D. & Grewal, S. I. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293, 1150–1155 (2001)
Cheung, P., Allis, C. D. & Sassone-Corsi, P. Signaling to chromatin through histone modifications. Cell 103, 263–271 (2000)
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000)
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 579–580 (2000)
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001)
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001)
Roguev, A. et al. The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J. 20, 7137–7148 (2001)
Briggs, S. D. et al. Histone H3 lysine 4 methylation is mediated by Set1 and required for cell growth and rDNA silencing in Saccharomyces cerevisiae. Genes Dev. 15, 3286–3295 (2001)
Nislow, C., Ray, E. & Pillus, L. SET1, a yeast member of the trithorax family, functions in transcriptional silencing and diverse cellular processes. Mol. Biol. Chem. 8, 2421–2436 (1997)
Krogan, N. J. et al. COMPASS: a complex of proteins associated with a trithorax-related SET domain protein. Proc. Natl Acad. Sci. 98, 12902–12907 (2001)
Bryk, M. et al. Evidence that Set1, a factor required for methylation of histone H3, regulates rDNA silencing in S. cerevisiae by a Sir2-independent mechanism. Curr. Biol. 12, 165–170 (2002)
Krogan, N. J. et al. COMPASS, a histone H3 (Lysine 4) methyltransferase required for telomeric silencing of gene expression. J. Biol. Chem. 13, 10753–10755 (2002)
Litt, M. D., Simpson, M., Gaszner, M., Allis, C. D. & Felsenfeld, G. Correlation between histone lysine methylation and developmental changes at the chicken β-globin locus. Science 293, 2453–2455 (2001)
Bernstein, B. E. et al. Methylation of histone H3 Lys 4 in coding regions of active genes. Proc. Natl Acad. Sci. USA 99, 8695–8697 (2002)
Siniossoglou, S. et al. A novel complex of nucleoporins, which includes Sec13p and a Sec13p homolog, is essential for normal nuclear pores. Cell 84, 265–275 (1996)
Carroll, A. S., Bishop, A. C., DeRisi, J. L., Shokat, K. M. & O'Shea, E. K. Chemical inhibition of the Pho85 cyclin-dependent kinase reveals a role in the environmental stress response. Proc. Natl Acad. Sci. USA 98, 12578–12583 (2001)
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997)
Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000)
Kent, N. A., Karabetsou, N., Politis, P. K. & Mellor, J. In vivo chromatin remodeling by yeast ISWI homologs Isw1p and Isw2p. Genes Dev. 15, 619–626 (2001)
Acknowledgements
We thank V. Geli for the gift of the UCC1001 wild-type strain, set1::HIS3 and set1::kan strains, and K. Nightingale for the gift of recombinant wild-type and tail-less H3 histone. This work was funded by an EU grant to H.S.R., an EMBO long-term fellowship to R.S. and a Cancer Research UK grant to T.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
T.K. is a founding member of Abcam, the company that has made the tri-methylated K4 histone H3 antibody.
Rights and permissions
About this article
Cite this article
Santos-Rosa, H., Schneider, R., Bannister, A. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002). https://doi.org/10.1038/nature01080
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature01080
This article is cited by
-
Transcriptionally active chromatin loops contain both ‘active’ and ‘inactive’ histone modifications that exhibit exclusivity at the level of nucleosome clusters
Epigenetics & Chromatin (2024)
-
Temporally-coordinated bivalent histone modifications of BCG1 enable fungal invasion and immune evasion
Nature Communications (2024)
-
Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Nature Communications (2024)
-
DPY30 promotes colorectal carcinoma metastasis by upregulating ZEB1 transcriptional expression
Cancer Cell International (2023)
-
Epigenetic regulation in metabolic diseases: mechanisms and advances in clinical study
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.