Abstract
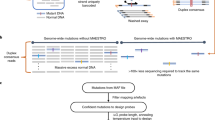
Array-based mutation detection methodology typically relies on direct hybridization of the fluorescently labeled query sequence to surface-bound oligonucleotide probes. These probes contain either small sequence variations or perfect-match sequence. The intensity of fluorescence bound to each oligonucleotide probe is intended to reveal which sequence is perfectly complementary to the query sequence1. However, these approaches have not always been successful, especially for detection of small frameshift mutations. Here we describe a multiplex assay to detect small insertions and deletions by using a modified PCR to evenly amplify each amplicon (PCR/PCR)2, followed by ligase detection reaction (LDR)3. Mutations were identified by screening reaction products with a universal DNA microarray4, which uncouples mutation detection from array hybridization and provides for high sensitivity. Using the three BRCA1 and BRCA2 founder mutations in the Ashkenazi Jewish population (BRCA1 185delAG; BRCA1 5382insC; BRCA2 6174delT)5 as a model system, the assay readily detected these mutations in multiplexed reactions. Our results demonstrate that universal microarray analysis of PCR/PCR/LDR2 products permits rapid identification of small insertion and deletion mutations in the context of both clinical diagnosis and population studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kozal, M.J. et al. Extensive polymorphisms observed in HIV-1 clade B protease gene using high-density oligonucleotide arrays. Nat. Med. 2, 753 –759 (1996).
Belgrader, P., Marino, M., Lubin, M. & Barany, F. A multiplex PCR-ligase detection reaction assay for human identity testing. Genome Sci. Technol. 1, 77–87 ( 1996).
Khanna, M. et al. Multiplex PCR/LDR for detection of K-ras mutations in primary colon tumors. Oncogene 18, 27– 38 (1999).
Gerry, N. et al. Universal DNA microarray method for multiplex detection of low abundance point mutations. J. Mol. Biol. 292, 251– 262 (1999).
Rahman, N. & Stratton, M. The genetics of breast cancer susceptibility. Annu. Rev. Genet. 32, 95– 121 (1998).
Hacia, J., Brody, L., Chee, M., Fodor, S. & Collins, F. Detection of heterozygous mutations in BRCA1 using high density oligonucleotide arrays and two-colour fluorescence analysis. Nat. Genet. 14, 441–447 (1996).
Ahrendt, S. et al. Rapid p53 sequence analysis in primary lung cancer using an oligonucleotide probe array. Proc. Natl. Acad. Sci. USA. 96, 7382–7387 (1999).
Hacia, J. Resequencing and mutational analysis using oligonucleotide microarrays. Nat. Genet. (Suppl.) 21, 42– 47 (1999).
Syvanen, A.C. & Landegren, U. Detection of point mutations by solid-phase methods. Hum. Mutat. 3, 172– 179 (1994).
Tong, J., Cao, W. & Barany, F. Biochemical properties of a high fidelity DNA ligase from Thermus species AK16D. Nucleic Acids Res. 27, 788– 794 (1999).
Luo, J., Bergstrom, D.E. & Barany, F. Improving the fidelity of Thermus thermophilus DNA ligase. Nucleic Acids Res. 24, 3071– 3078 (1996).
Zirvi, M. et al. Ligase-based detection of mononucleotide repeat sequences. Nucleic Acids Res. 27, e40 (1999).
Zirvi, M., Bergstrom, D.E., Saurage, A.S., Hammer, R.P. & Barany, F. Improved fidelity of thermostable ligases for detection of microsatellite repeat sequences using nucleoside analogs. Nucleic Acids Res. 27, e41 (1999).
Chakravarti, A. Population genetics—making sense out of sequence. Nat. Genet. 21, 56–60 ( 1999).
Luo, J. & Barany, F. Identification of essential residues in Thermus thermophilus DNA ligase. Nucleic Acids Res. 24, 3079–3085 (1996).
Barany, F. & Gelfand, D. Cloning, overexpression, and nucleotide sequence of a thermostable DNA ligase gene. Gene 109 , 1–11 (1991).
Acknowledgements
We thank Michael Osborne and Alvaro Monteiro for helpful discussion and members of the Barany lab for their suggestions and technical assistance. Support for this work was provided by the National Cancer Institute (P01-CA65930 and RO1-CA81467) and a grant from PE Biosystems.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Favis, R., Day, J., Gerry, N. et al. Universal DNA array detection of small insertions and deletions in BRCA1 and BRCA2. Nat Biotechnol 18, 561–564 (2000). https://doi.org/10.1038/75452
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/75452
This article is cited by
-
Genetic polymorphisms associated with nonalcoholic fatty liver disease in Uyghur population: a case-control study and meta-analysis
Lipids in Health and Disease (2019)
-
IL-2 gene polymorphisms affect tacrolimus response in myasthenia gravis
European Journal of Clinical Pharmacology (2019)
-
Association in a Chinese population of a genetic variation in the early B-cell factor 1 gene with coronary artery disease
BMC Cardiovascular Disorders (2017)
-
Systematic comparison of two whole-genome amplification methods for targeted next-generation sequencing using frozen and FFPE normal and cancer tissues
Scientific Reports (2017)
-
Association of DISC1 Polymorphisms with Late-Onset Alzheimer’s Disease in Northern Han Chinese
Molecular Neurobiology (2017)