Abstract
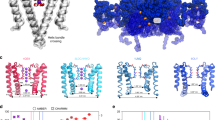
Ion-selective channels enable the specific permeation of ions through cell membranes and provide the basis of several important biological functions; for example, electric signalling in the nervous system1. Although a large amount of electrophysiological data is available1,2, the molecular mechanisms by which these channels can mediate ion transport remain a significant unsolved problem. With the recently determined crystal structure of the representative K+ channel (KcsA) from Streptomyces lividans3, it becomes possible to examine ion conduction pathways on a microscopic level. K+ channels utilize multi-ion conduction mechanisms1,2,4,5,6, and the three-dimensional structure also shows several ions present in the channel. Here we report results from molecular dynamics free energy perturbation calculations that both establish the nature of the multiple ion conduction mechanism and yield the correct ion selectivity of the channel. By evaluating the energetics of all relevant occupancy states of the selectivity filter, we find that the favoured conduction pathway involves transitions only between two main states with a free difference of about 5 kcal mol-1. Other putative permeation pathways can be excluded because they would involve states that are too high in energy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hille,B. Ionic Channels of Excitable Membranes (Sinauer, Sunderland, Massachusetts, 1992).
Latorre,R. & Miller,C. Conduction and selectivity in potassium channels. J. Membr. Biol. 71, 11– 30 (1983).
Doyle,D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Hodgkin,A. L. & Keynes,R. D. The potassium permeability of a giant nerve fibre. J. Physiol. 128, 61– 88 (1955).
Hille,B. & Schwarz,W. Potassium channels as multi-ion single-file pores. J. Gen. Physiol. 72, 409– 442 (1978).
Begenisich,T. & de Weer,P. Potassium flux ratio in voltage-clamped squid giant axons. J. Gen. Physiol. 76, 83–98 (1980).
Schrempf,H. et al. A prokaryotic potassium ion channel with two predicted transmembrane segments from Streptomyces lividans. EMBO J. 14, 5170–5178 (1995).
Bamberg,E. & Läuger,P. Temperature-dependent properties of gramicidin A channels. Biochim. Biophys. Acta 367 , 127–133 (1974).
Hladky,S. B. & Haydon,D. A. Ion transfer across lipid membranes in the presence of gramicidin A. Biochim. Biophys. Acta 274, 294–312 (1972).
Eisenman,G. & Horn,R. Ion selectivity revisited: the role of kinetic and equilibrium processes in ion permeation through channels. J. Membr. Biol. 76, 197–225 (1983).
Roux,B. & Karplus,M. Ion transport in a gramicidin-like channel: dynamics and mobility. J. Phys. Chem. 95, 4856–4868 (1991).
McCleskey,E. W. Calcium channel permeation: a field in flux. J. Gen. Physiol. 113, 765–772.
Miller,C. Ionic hopping defended. J. Gen. Physiol. 113, 783–787 (1999).
van Gunsteren,W. F. & Berendsen,H. J. C. Computer simulation of molecular dynamics: methodology, applications and perspectives in chemistry. Angew. Chem. Int. Edn Engl. 29, 992–1023 (1990).
Karplus,M. & Petsko,G. A. Molecular dynamics simulations in biology. Nature 347, 631– 639 (1990).
Kollman,P. Free energy calculations: applications to chemical and biochemical phenomena. Chem. Rev. 93, 2395–2417 (1993).
Roux,B. & MacKinnon,R. The cavity and pore helices in the KcsA K+ channel: electrostatic stabilization of monovalent cations. Science 285, 100– 102 (1999).
Åqvist,J. Calculation of absolute binding free energies for charged ligands and effects of long-range electrostatic interactions. J. Comput. Chem. 17, 1587–1597 (1996).
Marelius,J., Kolmodin,K., Feierberg,I. & Åqvist,J. Q: a molecular dynamics program for free energy calculations and empirical valence bond simulations in biomolecular systems. J. Mol. Graph. Model. 16, 213–225 ( 1998).
van Gunsteren,W. F. & Berendsen,H. J. C. Groningen Molecular Simulation (GROMOS) Library Manual (Biomos B.V., Groningen, The Netherlands, 1987).
Åqvist,J. Ion-water interaction potential derived from free energy perturbation simulations. J. Phys. Chem. 94, 8021– 8024 (1990).
Lee,F. S. & Warshel,A. A local reaction field method for fast evaluation of long-range electrostatic interactions in molecular simulations. J. Chem. Phys. 97, 3100– 3107 (1992).
Ryckaert,J. P., Ciccotti,G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints. J. Comput. Phys. 23, 327–341 (1977).
Acknowledgements
We thank T. A. Jones for comments and M. R. Harris for graphics. This work was supported by the Wenner–Gren Foundation and the Swedish Natural Science Research Council (NFR).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Åqvist, J., Luzhkov, V. Ion permeation mechanism of the potassium channel. Nature 404, 881–884 (2000). https://doi.org/10.1038/35009114
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35009114
This article is cited by
-
The Donnan potential revealed
Nature Communications (2022)
-
THz trapped ion model and THz spectroscopy detection of potassium channels
Nano Research (2022)
-
The Ca2+ permeation mechanism of the ryanodine receptor revealed by a multi-site ion model
Nature Communications (2020)
-
Potassium channel selectivity filter dynamics revealed by single-molecule FRET
Nature Chemical Biology (2019)
-
A quantum mechanical computational method for modeling electrostatic and solvation effects of protein
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.