Abstract
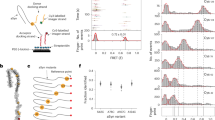
A practical high-throughput protein detection system is described, based on synthetic peptide arrays consisting of designed α-helical peptides, detected by fluorescence resonance energy transfer (FRET). Initially a model α-helical peptide known to interact with a structured protein, calmodulin, was selected to establish the strategy for high-throughput detection. In comparison to peptides with a single probe, a much higher FRET response has been observed with two fluorescent probes (7-diethylaminocoumarin-3-carboxylic acid and 5(6)-carboxy-fluorescein) at both termini of the synthetic peptides. To establish a reproducible high-throughput detection system, peptides were also immobilized onto a solid surface for detection of the target proteins. A small library of 112 different peptides was constructed, based on a model of the α-helical peptide with systematic replacement of residues carrying specific charges and/or hydrophobicities. The library was used to effectively characterize various proteins, giving their own `protein fingerprint' patterns. The resulting `protein fingerprints' correlate with the recognition properties of the proteins. The present microarray with designed synthetic peptides as the capturing agents is promising for the development of protein detection chips.
Similar content being viewed by others
References
Niemeyer, C. M. and Blohm, D., DNA microarrays, Angew. Chem. Int. Ed., 38 (1999) 2865-2869; Angew. Chem., 111 (1999) 3039-3043.
DeRisi, J. L., Iyer, V. R. and Brown, P. O., Exploring the metabolic and genetic control of gene expression on a genomic scale, Science, 278 (1997) 680-686.
Gygi, S. P., Rochon, Y., Franza, B. R. and Aebersold, R., Correlation between protein and mRNA abundance in yeast, Mol. Cell. Biol., 19 (1999) 1720-1730.
Anderson, N. L. and Anderson, N. G., Proteome and proteomics: New technologies, new concepts, and new words, Electrophoresis, 19 (1998) 1853-1861.
Shevchenko, A., Jensen, O. N., Podtelejnikov, A. V., Sagliocco, F., Wilm, M., Vorm, O., Mortensen, P., Shevchenko, A., Boucherie, H. and Mann, M., Linking genome and proteome by mass spectrometry: Large-scale identification of yeast proteins from two dimensional gels, Proc. Natl. Acad. Sci. USA, 93 (1996) 14440-14445.
Gygi, S. P., Rist, B., Gerber, S. A., Turecek, F., Gelb, M. H. and Aebersold, R., Quantitative analysis of complex protein mixtures using isotope-coded affinity tags, Nat. Biotech., 17 (1999) 994-999.
Natsume, T., Nakayama, H. and Isobe, T., BIA-MS-MS: Biomolecular interaction analysis for functional proteomics, Trends Biotech., 19 (Suppl.) (2001) S28–S33.
MacBeath, G. and Schreiber, S. L., Printing proteins as microarrays for high-throughput function determination, Science, 289 (2000) 1760–1763.
Arenkov, P., Kukhtin, A., Gemmell, A., Voloshchuk, S., Chupeeva, V. and Mirzabekov, A., Protein microchips: Use for immunoassay and enzymatic reactions, Anal. Biochem., 278 (2000) 123-131.
Zhu, H. and Snyder, M., Protein arrays and microarrays, Curr. Opin. Chem. Biol., 5 (2001) 40-45.
Zhu, H., Bilgin, M., Bangham, R., Hall, D., Casamayor, A., Bertone, P., Lan, N., Jansen, R., Bidlingmaier, S., Houfek, T., Mitchell, T., Miller, P., Dean, R. A., Gerstein, M. and Snyder, M., Global analysis of protein activities using proteome chips, Science, 293 (2001) 2101-2105.
Templin, M. F., Stoll, D., Schrenk, M., Traub, P. C., Vöhringer, C. F. and Joos, T. O., Protein microarray technology, Trends Biotech., 20 (2002) 160-166.
Wilson, D. S. and Nock, S., Recent developments in protein microarray technology, Angew. Chem. Int. Ed., 42 (2003) 494-500; Angew. Chem., 115 (2003) 510-517.
Mitchell, P., A perspective on protein microarrays, Nat. Biotech., 20 (2002) 225–229.
Takahashi, M., Nokihara, K. and Mihara, H., Construction of a protein-detection system using a loop peptide library with a fluorescence label, Chem. Biol., 10 (2003) 53–60.
Chan, W. C. and White, P. D., Fmoc solid phase peptide synthesis: A practical approach, Oxford University Press, New York, 2000.
Gregorius, K., Mouritsen, S. and Elsner, H. I., Hydrocoating: A new method for coupling biomolecules to solid phases, J. Immunol. Meth., 181 (1995) 65-73.
Wahler, D., Badalassi, F., Crotti, P. and Reymond, J. L., Enzyme fingerprints of activity, and stereo-and enantioselectivity from fluorogenic and chromogenic substrate arrays, Chem. Eur. J., 8 (2002) 3211-3228.
O'Neil, K. T. and DeGrado, W. F., How calmodulin binds its targets: Sequence independent recognition of amphiphilic a-helices, Trends Biol. Sci., 15 (1990) 59–64.
Maulet, Y. and Cox, J. A., Structural changes in melittin and calmodulin upon complex formation and their modulation by calcium, Biochemistry, 22 (1983) 5680-5686.
Cox, J. A., Comte, M., Fitton, J. E. and DeGrado, W. F., The interaction of calmodulin with amphiphilic peptides, J. Biol. Chem., 260 (1985) 2527-2534.
Greenfield, N. and Fasman, G. D., Computed circular dichroism spectra for the evaluation of protein conformation, Biochemistry, 8 (1969) 4108-4116.
Kuwabara, T., Nakamura, A., Ueno, A. and Toda, F., Inclusion complexes and guest-induced color changes of pH-indicator-modified β-cyclodextrins.J. Phys. Chem., 98 (1994) 6297-6303.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Usui, K., Takahashi, M., Nokihara, K. et al. Peptide arrays with designed α-helical structures for characterization of proteins from FRET fingerprint patterns. Mol Divers 8, 209–218 (2004). https://doi.org/10.1023/B:MODI.0000036237.82584.2d
Issue Date:
DOI: https://doi.org/10.1023/B:MODI.0000036237.82584.2d