Abstract
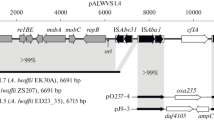
The carbazole-degrading (car) operon on the chromosome of Pseudomonas stutzeri strain OM1 showed >99% identity to that in the 72.8 kb catabolic transposon, Tn4676, on plasmid pCAR1. Southern hybridization using probes prepared from the pCAR1 sequence and sequencing analyses showed that the OM1 chromosome contained the 55 kb DNA region, almost all of which was a part of Tn4676, flanked by two copies of novel insertion sequence, ISPst3, and included the car gene.
Similar content being viewed by others
References
Berger B and Haas D (2001) Transposase and cointegrase: specialized transposition proteins of the bacterial insertion sequence IS21 and related elements. Cell Mol. Life Sci. 58: 403–419.
Dyda F, Hickman AB, Jenkins TM, Engelman A, Craigie R, Davies DR (1994) Crystal structure of the catalytic domain of HIV-1 integrase: similarity to other polynucleotidyl transferases. Science 266: 1981–1986.
Ka JO and Tiedje JM (1994) Integration and excision of a 2,4-dichlorophenoxyacetic acid-degradative plasmid in Alcaligenes paradoxus and evidence of its natural intergeneric transfer. J. Bacteriol. 176: 5284–5289.
Maeda K, Nojiri H, Shintani M, Yoshida T, Habe H, Omori T (2003) Complete nucleotide sequence of carbazole/dioxindegrading plasmid pCAR1 in Pseudomonas resinovorans strain CA10 indicates its mosaicity and the presence of large catabolic transposon Tn4676. J. Mol. Biol. 326: 21–33.
Nojiri H, Sekiguchi H, Maeda K, Urata M, Nakai S, Yoshida T, Habe H, Omori T (2001) Genetic characterization and evolutionary implications of car gene cluster in the carbazole degrader Pseudomonas sp. strain CA10. J. Bacteriol. 183: 3663–3679.
Ouchiyama N, Miyachi S, Omori T (1998) Cloning and nucleotide sequence of carbazole catabolic genes from Pseudomonas stutzeri strain OM1, isolated from activated sludge. J. Gen. Appl. Microbiol. 44: 57–63.
Ouchiyama N, Zhang Y, Omori T, and Kodama T (1993) Biodegradation of carbazole by Pseudomonas spp. CA06 and CA10. Biosci. Biotechnol. Biochem. 57: 455–460.
Sambrook J, Russell DW (2001) Molecular Cloning. A Laboratory Manual, 3rd edn. Cold Spring Harbour, NY: Cold Spring Harbour Laboratory Press.
Sato S, Nam J-W, Kasuga K, Nojiri H, Yamane H, Omori T (1997a) Identification and characterization of genes encoding carbazole 1,9a-dioxygenase in Pseudomonas sp. strain CA10. J. Bacteriol. 179: 4850–4858.
Sato S, Ouchiyama N, Kimura T, Nojiri H, Yamane H Omori T (1997b) Cloning of genes involved in carbazole degradation of Pseudomonas sp. strain CA10: nucleotide sequences of genes and characterization of meta-cleavage enzymes and hydrolase. J. Bacteriol. 179: 4841–4849.
Soby S, Kirkpatrick B, Kosuge T (1993) Characterization of an insertion sequence (IS53) located within IS51 on the iaacontaining plasmid of Pseudomonas syringae pv. savastanoi. Plasmid 29: 135–141.
Solinas F, Marconi AM, Ruzzi M, Zennaro E (1995) Characterization of a novel insertion sequence, IS1162, from Pseudomonas fluorescens. Gene 155: 77–82.
Tsuda M, Genka H (2001) Identification and characterization of Tn4656, a novel class II transposon carrying a set of toluenedegrading genes from TOL plasmid pWW53. J. Bacteriol. 183: 6215–6224.
Tsuda M, Iino T (1987) Genetic analysis of a transposon carrying toluene degrading genes on a TOL plasmid pWW0. Mol. Gen. Genet. 210: 270–276.
Tsuda M, Iino T (1988) Identification and characterization of Tn4653, a transposon covering the toluene transposon Tn4651 on TOL plasmid pWW0. Mol. Gen. Genet. 213: 72–77.
Yeo CC, Poh CL (1997) Characterization of IS1474, an insertion sequence of the IS21 family isolated from Pseudomonas alcaligenes NCIB9867. FEMS Microbiol. Lett. 149: 257–263.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shintani, M., Nojiri, H., Yoshida, T. et al. Carbazole/dioxin-degrading car gene cluster is located on the chromosome of Pseudomonas stutzeri strain OM1 in a form different from the simple transposition of Tn4676 . Biotechnology Letters 25, 1255–1261 (2003). https://doi.org/10.1023/A:1025079027730
Issue Date:
DOI: https://doi.org/10.1023/A:1025079027730