Abstract
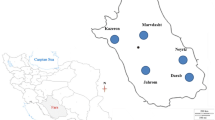
Citrus accessions including several undefined local or native genotypes originated from the regions of Caspian Sea (Iran) as well as some known varieties were analyzed by simple sequence repeat (SSR) markers. The aim of the present research work was to characterize 33 accessions of citrus, with unknown pedigrees, to assess their genetic diversity and to elucidate their possible relationships with 12 commercially important varieties. Sixteen SSR primer pairs with high levels of polymorphism, which have been previously developed for citrus, were used in this study. A total of 104 alleles were obtained for all the microsatellites studied, ranging from 2 to 13 alleles per locus, with a mean value of 6.5 alleles per locus. Polymorphic information content varied from 0.252 to 0.879 with mean of 0.610. The percentage of heterozygosity per marker detected in our samples ranged from 2.30 to 100 % with an average of 46.22 %. The genetic distance matrix was used to construct neighbor joining cluster which allowed the arrangement of all the genotypes according to their genetic relationships. The genetic relationships among known citrus varieties with uncharacterized citrus natural types are discussed. By analyzing dendrogram, it is confirmed that mandarin (Citrus reticulata), pummelo (C. grandis), and citron (C. medica) were clustered into three particular groups as major species of Citrus. The obtained data confirmed the usefulness of SSR markers for estimation of citrus genetic diversity.


Similar content being viewed by others
References
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a Citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Barret HC, Rhodes AM (1976) A numerical taxonomic study affinity relationships in cultivated Citrus and its close relatives. Syst Bot 1:105–136
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Breto MP, Ruiz C, Pina JA, Asins MJ (2001) The diversification of Citrus clementina Hort. ex Tan., a vegetatively propagated crop species. Mol Phylogenet Evol 21:285–293
Corazza-Nunes MJ, Machado MA, Nunes WMC, Cristofani M, Targon MLPN (2002) Assessment of genetic variability in grapefruits (C. paradisi Macf.) and pummelos (C. maxima (Burm.) Merr.) using RAPD and SSRs markers. Euphytica 126:169–176
Fang DQ, Roose ML (1997) Identification of closely related Citrus cultivars with inter- simple sequence repeat marker. Theor Appl Genet 95:408–417
FAO Commodities and Trade Division (2010) Citrus fruit, fresh and processed. Annual statistics. Food and Agriculture Organization of the United Nations, Rome (CCP: CI/ST/2007)
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 94:812–822
Gmitter FG, Grosse JW, Moore AG (1992) Citrus. In: Hammerschlag FA, Litz RE (eds) Biotechnology of perennial fruit crops. CAB International, Wallingford, pp 335–369
Golein B, Adouli B (2011) Citrus. Pouya Press, Tehran (in Persian)
Golein B, Bigonah M, Azadvar M, Golmohammadi M (2012) Analysis of genetic relationship between ‘Bakraee’ (Citrus sp.) and some known Citrus genotypes through SSR and PCR-RFLP markers. Sci Hortic 148:147–153
Green RM, Vardi A, Galun E (1986) The plastome of Citrus: physical map, variation among Citrus cultivars and species and comparison with related genera. Theor Appl Genet 72:170–177
Gulsen O, Roose ML (2001) Lemons: diversity and relationship with selected Citrus genotypes as measured with nuclear genome markers. J Am Soc Hortic Sci 126:309–317
Herrero R, Asins MJ, Carbonell AE, Navarro L (1996) Genetic diversity in the orange subfamily Aurantioideae. I. Intraspecies and intragenus genetic variability. Theor Appl Genet 92:599–609
International Plant Genetic Resources Institute (1999) Descriptors for Citrus. International Plant Genetic Resources Institute, Rome. Available at http://www.ipgri.cgiar.org/
Iwamasa M, Nito N, Ling JT (1988) Intra and inter generic hybridization in the orange subfamily Aurantioideae. Proc Int Soc Citric 1:123–130
Kumar S, Nair KN (2013) Genetic variation and phylogenetic relationships among Indian Citrus taxa revealed by DAMD-PCR markers. Genet Resour Crop Evol 60:1777–1800
Luro F, Laigret F, Bove JM, Ollitrault P (1995) DNA amplified fingerprinting, a useful tool for determination of genetic origin and diversity analysis in Citrus. HortScience 30:1063–1067
Malik SK, Rohini, Kumar S, Choudhary R, Pal D, Chaudhury R (2012) Assessment of genetic diversity in sweet orange [Citrus sinensis (L.) Osbeck] cultivars of India using morphological and RAPD markers. Agric Res 1(4):317–324
Moore GA (2001) Oranges and lemons: clues to the taxonomy of Citrus from molecular markers. Trends Genet 17:536–540
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Naghavi MR, Ghariazi B, Hossini Salekdeh G (2005) Molecular markers. Tehran University Press, Tehran (In Persian)
Nicolosi E, Deng Z, Gentile A, LaMalfa S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Novelli VM, Machado MA, Lopes CR (2006) Isoenzymatic polymorphism in Citrus spp. and P. trifoliata (L.) Raf. (Rutaceae). Genet Mol Biol 23:163–168
Pal D, Malik SK, Kumar S, Choudhary R, Sharma KC, Chaudhury R (2013) Genetic variability and relationship studies of mandarin (Citrus reticulate Blanco) using morphological and molecular markers. Agric Res 2(3):236–245
Rouhi Ghorabaie HR, Fotouhi Ghazvini R, Golein B, Nabipour AR (2010) Identification of Some Citrus Accessions in a Citrus Germplasm Utilizing Simple Sequence Repeat Markers (SSRs). Hortic Environ Biotechnol 51(4):343–347
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Phyl Evol 4:477–491
Scora RW (1975) On the history and origin of Citrus. Bull Torrey Bot Club 102:369–375
Scora RW (1988) Biochemistry, taxonomy and evolution of modern cultivated Citrus. Proc Int Soc Citric 1:277–289
Shahsavar AR, Izadpanah K, Tafazoli E, Tabatabaei BES (2007) Characterization of Citrus germplasm including unknown variants by inter-simple sequence repeat (ISSR) markers. Sci Hortic 112:310–314
Swingle WT, Reece PC (1967) The botany of Citrus and its wild relatives. In: Reuther W, Webber HJ, Bachelor LD (eds) The Citrus industry, vol 1. University of California Press, Berkeley, pp 190–430
Torres AM, Soost RK, Diedenhofen U (1978) Leaf isosymes as genetic markers in Citrus. Am J Bot 65:869–881
Uzuan A, Yesiloglu T, Aka-Kacar Y, Tuzcu O, Gulsen O (2009) Genetic diversity and relationships within Citrus and related genera based on sequence related amplified polymorphism markers (SRAPs). Sci Hortic 121:306–312
Wright S (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19:395–420
Yamamoto M, Kobayashi S, Nakamura Y, Yamada Y (1993) Phylogenetic relationships of Citrus revealed by diversity of cytoplasmic genomes. In: Hayashi T, Omura M, Scott NS (eds) Techniques on gene diagnosis and breeding in fruit trees. Fruit Trees Research Station Press, Okitsu, pp 39–46
Zane L, Bargelloni L, Patarnello T (2002) Strategies for microsatellite isolation: a review. Mol Ecol 11:1–16
Acknowledgments
The help and cooperation received from Citrus Research Institute, I.R. Iran is fully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bakhshipour, H., Golein, B., Mehregan, I. et al. Simple Sequence Repeat Polymorphism in Iranian Citrus Germplasm Including Unknown Variants. Agric Res 4, 152–159 (2015). https://doi.org/10.1007/s40003-015-0159-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40003-015-0159-5