Abstract
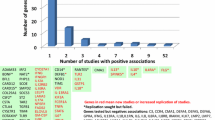
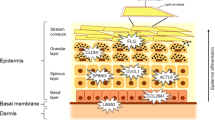
Atopic dermatitis (AD) is a chronic inflammatory skin disease caused by a combination of genetic and environmental factors. Genetic evidences depict a complex network comprising by epidermal barrier dysfunctions and dysregulation of innate and adaptive immunity in the pathogenesis of AD. Mutations in the human filaggrin gene (FLG) are the most significant and well-replicated genetic mutation associated with AD, and other mutations associated with epidermal barriers such as SPINK5, FLG-2, SPRR3, and CLDN1 have all been linked to AD. Gene variants may also contribute to the abnormal innate and adaptive responses found in AD, including mutations in PRRs and AMPs, TSLP and TSLPR, IL-1 family cytokines and receptors genes, vitamin D pathway genes, FCER1A, and Th2 and other cytokines genes. GWAS and Immunochip analysis have identified a total of 19 susceptibility loci for AD. Candidate genes at these susceptibility loci identified by GWAS and Immunochip analysis also suggest roles for epidermal barrier functions, innate and adaptive immunity, interleukin-1 family signaling, regulatory T cells, the vitamin D pathway, and the nerve growth factor pathway in the pathogenesis of AD. Increasing evidences show the modern lifestyle (i.e., the hygiene hypothesis, Western diet) and other environmental factors such as pollution and environmental tobacco smoke (ETS) lead to the increasing prevalence of AD with the development of industrialization. Epigenetic alterations in response to these environmental factors, including DNA methylation and microRNA related to immune system and skin barriers, have been found to contribute to the pathogenesis of AD. Genetic variants and epigenetic alteration might be the key tools for the molecular taxonomy of AD and provide the background for the personalized management.

Similar content being viewed by others
References
Bieber T (2008) Atopic dermatitis. N Engl J Med 358:1483–1494
Esparza-Gordillo J, Weidinger S, Folster-Holst R et al (2009) A common variant on chromosome 11q13 is associated with atopic dermatitis. Nat Genet 41:596–601
Sun LD, Xiao FL, Li Y et al (2011) Genome-wide association study identifies two new susceptibility loci for atopic dermatitis in the Chinese Han population. Nat Genet 43:690–694
Paternoster L, Standl M, Chen CM et al (2011) Meta-analysis of genome-wide association studies identifies three new risk loci for atopic dermatitis. Nat Genet 44:187–192
Hirota T, Takahashi A, Kubo M et al (2012) Genome-wide association study identifies eight new susceptibility loci for atopic dermatitis in the Japanese population. Nat Genet 44:1222–1226
Ellinghaus D, Baurecht H, Esparza-Gordillo J et al (2013) High-density genotyping study identifies four new susceptibility loci for atopic dermatitis. Nat Genet 45:808–812
Mu Z, Zhao Y, Liu X, Chang C, Zhang J (2014) Molecular biology of atopic dermatitis. Clin Rev Allergy Immunol 47:193–218
Morales Suarez-Varela M, Garcia-Marcos L, Kogan MD et al (2008) Parents’ smoking habit and prevalence of atopic eczema in 6-7 and 13-14 year-old schoolchildren in Spain. ISAAC phase III. Allergol Immunopathol (Madr) 36:336–342
Leung DY, Bieber T (2003) Atopic dermatitis. Lancet 361:151–160
Schafer T, Heinrich J, Wjst M et al (1999) Indoor risk factors for atopic eczema in school children from East Germany. Environ Res 81:151–158
Wang IJ, Chen SL, Lu TP, Chuang EY, Chen PC (2013) Prenatal smoke exposure, DNA methylation, and childhood atopic dermatitis. Clin Exp Allergy 43:535–543
Jirtle RL, Skinner MK (2007) Environmental epigenomics and disease susceptibility. Nat Rev Genet 8:253–262
Feil R, Fraga MF (2012) Epigenetics and the environment: emerging patterns and implications. Nat Rev Genet 13:97–109
Herberth G, Bauer M, Gasch M et al (2014) Maternal and cord blood miR-223 expression associates with prenatal tobacco smoke exposure and low regulatory T-cell numbers. J Allergy Clin Immunol 133:543–550
Hinz D, Bauer M, Roder S et al (2012) Cord blood Tregs with stable FOXP3 expression are influenced by prenatal environment and associated with atopic dermatitis at the age of one year. Allergy 67:380–389
Rodriguez E, Baurecht H, Wahn AF et al (2014) An integrated epigenetic and transcriptomic analysis reveals distinct tissue-specific patterns of DNA methylation associated with atopic dermatitis. J Investig Dermatol 134:1873–1883
Tezza G, Mazzei F, Boner A (2013) Epigenetics of allergy. Early Hum Dev 89(Suppl 1):S20–S21
Sharma S, Litonjua A (2014) Asthma, allergy, and responses to methyl donor supplements and nutrients. J Allergy Clin Immunol 133:1246–1254
Sandilands A, Sutherland C, Irvine AD, McLean WH (2009) Filaggrin in the frontline: role in skin barrier function and disease. J Cell Sci 122:1285–1294
Brown SJ, McLean WH (2012) One remarkable molecule: filaggrin. J Investig Dermatol 132:751–762
McAleer MA, Irvine AD (2013) The multifunctional role of filaggrin in allergic skin disease. J Allergy Clin Immunol 131:280–291
Irvine AD, McLean WH, Leung DY (2011) Filaggrin mutations associated with skin and allergic diseases. N Engl J Med 365:1315–1327
Elias PM, Wakefield JS (2014) Mechanisms of abnormal lamellar body secretion and the dysfunctional skin barrier in patients with atopic dermatitis. J Allergy Clin Immunol 134:781–791
Palmer CN, Irvine AD, Terron-Kwiatkowski A et al (2006) Common loss-of-function variants of the epidermal barrier protein filaggrin are a major predisposing factor for atopic dermatitis. Nat Genet 38:441–446
Rodriguez E, Baurecht H, Herberich E et al (2009) Meta-analysis of filaggrin polymorphisms in eczema and asthma: robust risk factors in atopic disease. J Allergy Clin Immunol 123:1361–1370
van den Oord RA, Sheikh A (2009) Filaggrin gene defects and risk of developing allergic sensitisation and allergic disorders: systematic review and meta-analysis. BMJ 339:b2433
Brown SJ, McLean WH (2009) Eczema genetics: current state of knowledge and future goals. J Investig Dermatol 129:543–552
Brown SJ, Sandilands A, Zhao Y et al (2008) Prevalent and low-frequency null mutations in the filaggrin gene are associated with early-onset and persistent atopic eczema. J Investig Dermatol 128:1591–1594
Margolis DJ, Apter AJ, Gupta J et al (2012) The persistence of atopic dermatitis and filaggrin (FLG) mutations in a US longitudinal cohort. J Allergy Clin Immunol 130:912–917
Weidinger S, Rodriguez E, Stahl C et al (2007) Filaggrin mutations strongly predispose to early-onset and extrinsic atopic dermatitis. J Investig Dermatol 127:724–726
Gao PS, Rafaels NM, Hand T et al (2009) Filaggrin mutations that confer risk of atopic dermatitis confer greater risk for eczema herpeticum. J Allergy Clin Immunol 124:507–513, 513 e501-507
Weidinger S, O’Sullivan M, Illig T et al (2008) Filaggrin mutations, atopic eczema, hay fever, and asthma in children. J Allergy Clin Immunol 121:1203–1209
Palmer CN, Ismail T, Lee SP et al (2007) Filaggrin null mutations are associated with increased asthma severity in children and young adults. J Allergy Clin Immunol 120:64–68
Brown SJ, Asai Y, Cordell HJ et al (2011) Loss-of-function variants in the filaggrin gene are a significant risk factor for peanut allergy. J Allergy Clin Immunol 127:661–667
Henderson J, Northstone K, Lee SP et al (2008) The burden of disease associated with filaggrin mutations: a population-based, longitudinal birth cohort study. J Allergy Clin Immunol 121:872–877
O’Regan GM, Kemperman PM, Sandilands A et al (2008) Raman profiles of the stratum corneum define 3 filaggrin genotype-determined atopic dermatitis endophenotypes. J Allergy Clin Immunol 126:574–580
Kezic S, O’Regan GM, Lutter R et al (2012) Filaggrin loss-of-function mutations are associated with enhanced expression of IL-1 cytokines in the stratum corneum of patients with atopic dermatitis and in a murine model of filaggrin deficiency. J Allergy Clin Immunol 129:1031–1039
Murphy JE, Robert C, Kupper TS (2000) Interleukin-1 and cutaneous inflammation: a crucial link between innate and acquired immunity. J Investig Dermatol 114:602–608
Brown SJ, Kroboth K, Sandilands A et al (2012) Intragenic copy number variation within filaggrin contributes to the risk of atopic dermatitis with a dose-dependent effect. J Investig Dermatol 132:98–104
Walley AJ, Chavanas S, Moffatt MF et al (2001) Gene polymorphism in Netherton and common atopic disease. Nat Genet 29:175–178
Hachem JP, Wagberg F, Schmuth M et al (2006) Serine protease activity and residual LEKTI expression determine phenotype in Netherton syndrome. J Investig Dermatol 126:1609–1621
Nishio Y, Noguchi E, Shibasaki M et al (2003) Association between polymorphisms in the SPINK5 gene and atopic dermatitis in the Japanese. Genes Immun 4:515–517
Kusunoki T, Okafuji I, Yoshioka T et al (2005) SPINK5 polymorphism is associated with disease severity and food allergy in children with atopic dermatitis. J Allergy Clin Immunol 115:636–638
Lan CC, Tu HP, Wu CS et al (2011) Distinct SPINK5 and IL-31 polymorphisms are associated with atopic eczema and non-atopic hand dermatitis in Taiwanese nursing population. Exp Dermatol 20:975–979
Kato A, Fukai K, Oiso N, Hosomi N, Murakami T, Ishii M (2003) Association of SPINK5 gene polymorphisms with atopic dermatitis in the Japanese population. Br J Dermatol 148:665–669
Zhao LP, Di Z, Zhang L et al (2012) Association of SPINK5 gene polymorphisms with atopic dermatitis in Northeast China. J Eur Acad Dermatol Venereol 26:572–577
Fortugno P, Furio L, Teson M et al (2012) The 420K LEKTI variant alters LEKTI proteolytic activation and results in protease deregulation: implications for atopic dermatitis. Hum Mol Genet 21:4187–4200
Briot A, Deraison C, Lacroix M et al (2009) Kallikrein 5 induces atopic dermatitis-like lesions through PAR2-mediated thymic stromal lymphopoietin expression in Netherton syndrome. J Exp Med 206:1135–1147
Pellerin L, Henry J, Hsu CY et al (2012) Defects of filaggrin-like proteins in both lesional and nonlesional atopic skin. J Allergy Clin Immunol 131:1094–1102
Makino T, Mizawa M, Yamakoshi T, Takaishi M, Shimizu T (2014) Expression of filaggrin-2 protein in the epidermis of human skin diseases: a comparative analysis with filaggrin. Biochem Biophys Res Commun 449:100–106
Wu Z, Hansmann B, Meyer-Hoffert U, Glaser R, Schroder JM (2009) Molecular identification and expression analysis of filaggrin-2, a member of the S100 fused-type protein family. PLoS One 4:e5227
Margolis DJ, Gupta J, Apter AJ et al (2014) Filaggrin-2 variation is associated with more persistent atopic dermatitis in African American subjects. J Allergy Clin Immunol 133:784–789
Marenholz I, Rivera VA, Esparza-Gordillo J et al (2011) Association screening in the Epidermal Differentiation Complex (EDC) identifies an SPRR3 repeat number variant as a risk factor for eczema. J Investig Dermatol 131:1644–1649
Kelsell DP, Byrne C (2011) SNPing at the epidermal barrier. J Investig Dermatol 131:1593–1595
Saunders SP, Goh CS, Brown SJ et al (2013) Tmem79/Matt is the matted mouse gene and is a predisposing gene for atopic dermatitis in human subjects. J Allergy Clin Immunol 132:1121–1129
Sasaki T, Shiohama A, Kubo A et al (2013) A homozygous nonsense mutation in the gene for Tmem79, a component for the lamellar granule secretory system, produces spontaneous eczema in an experimental model of atopic dermatitis. J Allergy Clin Immunol 132:1111–1120
De Benedetto A, Slifka MK, Rafaels NM et al (2011) Reductions in claudin-1 may enhance susceptibility to herpes simplex virus 1 infections in atopic dermatitis. J Allergy Clin Immunol 128:242–246
Mu Z, Zhao Y, Liu X, Chang C, Zhang J (2012) Molecular biology of atopic dermatitis. Clin Rev Allergy Immunol 47:193–218
Eyerich K, Novak N (2013) Immunology of atopic eczema: overcoming the Th1/Th2 paradigm. Allergy 68:974–982
Wollenberg A, Rawer HC, Schauber J (2008) Innate immunity in atopic dermatitis. Clin Rev Allergy Immunol 41:272–281
Potaczek DP, Nastalek M, Okumura K, Wojas-Pelc A, Undas A, Nishiyama C (2011) An association of TLR2-16934A >T polymorphism and severity/phenotype of atopic dermatitis. J Eur Acad Dermatol Venereol 25:715–721
Ahmad-Nejad P, Mrabet-Dahbi S, Breuer K et al (2004) The toll-like receptor 2 R753Q polymorphism defines a subgroup of patients with atopic dermatitis having severe phenotype. J Allergy Clin Immunol 113:565–567
Mrabet-Dahbi S, Dalpke AH, Niebuhr M et al (2008) The Toll-like receptor 2 R753Q mutation modifies cytokine production and Toll-like receptor expression in atopic dermatitis. J Allergy Clin Immunol 121:1013–1019
Niebuhr M, Lutat C, Sigel S, Werfel T (2009) Impaired TLR-2 expression and TLR-2-mediated cytokine secretion in macrophages from patients with atopic dermatitis. Allergy 64:1580–1587
Niebuhr M, Langnickel J, Draing C, Renz H, Kapp A, Werfel T (2008) Dysregulation of toll-like receptor-2 (TLR-2)-induced effects in monocytes from patients with atopic dermatitis: impact of the TLR-2 R753Q polymorphism. Allergy 63:728–734
Levchenko L, Izmailova OV, Shlykova OA, Kaidashev IP (2013) Polymorphism 896A/G of TLR4 gene rather than 1196C/T and 2258G/A of TLR2 gene determines severe and complicated course of atopic dermatitis in children. Tsitol Genet 47:46–53
Miedema KG, Tissing WJ, Te Poele EM et al (2005) Polymorphisms in the TLR6 gene associated with the inverse association between childhood acute lymphoblastic leukemia and atopic disease. Leukemia 26:1203–1210
Hoffjan S, Stemmler S, Parwez Q et al (2005) Evaluation of the toll-like receptor 6 Ser249Pro polymorphism in patients with asthma, atopic dermatitis and chronic obstructive pulmonary disease. BMC Med Genet 6:34
Novak N, Yu CF, Bussmann C et al (2007) Putative association of a TLR9 promoter polymorphism with atopic eczema. Allergy 62:766–772
Weidinger S, Klopp N, Rummler L et al (2005) Association of CARD15 polymorphisms with atopy-related traits in a population-based cohort of Caucasian adults. Clin Exp Allergy 35:866–872
Macaluso F, Nothnagel M, Parwez Q et al (2007) Polymorphisms in NACHT-LRR (NLR) genes in atopic dermatitis. Exp Dermatol 16:692–698
Weidinger S, Klopp N, Rummler L et al (2005) Association of NOD1 polymorphisms with atopic eczema and related phenotypes. J Allergy Clin Immunol 116:177–184
Prado-Montes de Oca E, Garcia-Vargas A, Lozano-Inocencio R et al (2007) Association of beta-defensin 1 single nucleotide polymorphisms with atopic dermatitis. Int Arch Allergy Immunol 142:211–218
Mohamed HG, Abbas A, El-Kabarity RH, Diab HM (2009) Association of beta-defensin 1 single nucleotide polymorphism with atopic dermatitis. Egypt J Immunol 16:125–138
Kim E, Lee JE, Namkung JH et al (2009) Single nucleotide polymorphisms and the haplotype in the DEFB1 gene are associated with atopic dermatitis in a Korean population. J Dermatol Sci 54:25–30
Montes P, de Oca E, Li W (2013) Human beta-defensin 1 (DEFB1) allele and genotype frequencies probably impact on ethnic susceptibility to atopic dermatitis. Int J Dermatol 52:115–117
Kou K, Aihara M, Matsunaga T et al (2012) Association of serum interleukin-18 and other biomarkers with disease severity in adults with atopic dermatitis. Arch Dermatol Res 304:305–312
Savinko T, Matikainen S, Saarialho-Kere U et al (2012) IL-33 and ST2 in atopic dermatitis: expression profiles and modulation by triggering factors. J Investig Dermatol 132:1392–1400
Novak N, Kruse S, Potreck J et al (2005) Single nucleotide polymorphisms of the IL18 gene are associated with atopic eczema. J Allergy Clin Immunol 115:828–833
Ibrahim GH, ElTabbakh MT, Gomaa AH, Mohamed EA (2012) Interleukin-18 gene polymorphisms in Egyptian patients with allergic diseases. Am J Rhinol Allergy 26:385–389
Jariwala SP, Abrams E, Benson A, Fodeman J, Zheng T (2011) The role of thymic stromal lymphopoietin in the immunopathogenesis of atopic dermatitis. Clin Exp Allergy 41:1515–1520
Liu YJ (2007) Thymic stromal lymphopoietin and OX40 ligand pathway in the initiation of dendritic cell-mediated allergic inflammation. J Allergy Clin Immunol 120:238–244, quiz 245-236
Hoffjan S, Beygo J, Akkad DA, Parwez Q, Petrasch-Parwez E, Epplen JT (2009) Analysis of variation in the IL7RA and IL2RA genes in atopic dermatitis. J Dermatol Sci 55:138–140
Gao PS, Rafaels NM, Mu D et al (2010) Genetic variants in thymic stromal lymphopoietin are associated with atopic dermatitis and eczema herpeticum. J Allergy Clin Immunol 125:1403–1407
Margolis DJ, Kim B, Apter AJ et al (2014) Thymic stromal lymphopoietin variation, filaggrin loss of function, and the persistence of atopic dermatitis. JAMA Dermatol 150:254–259
Benson AA, Toh JA, Vernon N, Jariwala SP (2012) The role of vitamin D in the immunopathogenesis of allergic skin diseases. Allergy 67:296–301
Wang SS, Hon KL, Kong AP et al (2013) Eczema phenotypes are associated with multiple vitamin D pathway genes in Chinese children. Allergy 69:118–124
He JQ, Chan-Yeung M, Becker AB et al (2003) Genetic variants of the IL13 and IL4 genes and atopic diseases in at-risk children. Genes Immun 4:385–389
Iizuka M, Katsuyama Y, Mabuchi T et al (2002) Genetic association analysis using microsatellite markers in atopic dermatitis. Tokai J Exp Clin Med 27:51–56
Kawashima T, Noguchi E, Arinami T et al (1998) Linkage and association of an interleukin 4 gene polymorphism with atopic dermatitis in Japanese families. J Med Genet 35:502–504
Namkung JH, Lee JE, Kim E et al (2011) Association of polymorphisms in genes encoding IL-4, IL-13 and their receptors with atopic dermatitis in a Korean population. Exp Dermatol 20:915–919
Hershey GK, Friedrich MF, Esswein LA, Thomas ML, Chatila TA (1997) The association of atopy with a gain-of-function mutation in the alpha subunit of the interleukin-4 receptor. N Engl J Med 337:1720–1725
Oiso N, Fukai K, Ishii M (2000) Interleukin 4 receptor alpha chain polymorphism Gln551Arg is associated with adult atopic dermatitis in Japan. Br J Dermatol 142:1003–1006
Isidoro-Garcia M, Davila I, Moreno E, Laffond E, Lorente F, Gonzalez-Sarmiento R (2005) IL4RA gene polymorphism (Q576R) is associated with higher total IgE levels in Spanish patients with family history of atopy. Med Clin (Barc) 124:211–212
Yamamoto N, Sugiura H, Tanaka K, Uehara M (2003) Heterogeneity of interleukin 5 genetic background in atopic dermatitis patients: significant difference between those with blood eosinophilia and normal eosinophil levels. J Dermatol Sci 33:121–126
Namkung JH, Lee JE, Kim E et al (2007) IL-5 and IL-5 receptor alpha polymorphisms are associated with atopic dermatitis in Koreans. Allergy 62:934–942
Vladich FD, Brazille SM, Stern D, Peck ML, Ghittoni R, Vercelli D (2005) IL-13 R130Q, a common variant associated with allergy and asthma, enhances effector mechanisms essential for human allergic inflammation. J Clin Invest 115:747–754
Tamura K, Arakawa H, Suzuki M et al (2001) Novel dinucleotide repeat polymorphism in the first exon of the STAT-6 gene is associated with allergic diseases. Clin Exp Allergy 31:1509–1514
Tamura K, Suzuki M, Arakawa H, Tokuyama K, Morikawa A (2003) Linkage and association studies of STAT6 gene polymorphisms and allergic diseases. Int Arch Allergy Immunol 131:33–38
Schulz F, Marenholz I, Folster-Holst R et al (2007) A common haplotype of the IL-31 gene influencing gene expression is associated with nonatopic eczema. J Allergy Clin Immunol 120:1097–1102
Nagy N, Tanaka A, Techanukul T, McGrath JA (2010) Common IL-31 gene haplotype associated with non-atopic eczema is not implicated in epidermolysis bullosa pruriginosa. Acta Derm Venereol 90:631–632
Tsunemi Y, Saeki H, Nakamura K et al (2002) Interleukin-12 p40 gene (IL12B) 3′-untranslated region polymorphism is associated with susceptibility to atopic dermatitis and psoriasis vulgaris. J Dermatol Sci 30:161–166
Takahashi N, Akahoshi M, Matsuda A et al (2005) Association of the IL12RB1 promoter polymorphisms with increased risk of atopic dermatitis and other allergic phenotypes. Hum Mol Genet 14:3149–3159
Leung DY, Gao PS, Grigoryev DN et al (2011) Human atopic dermatitis complicated by eczema herpeticum is associated with abnormalities in IFN-gamma response. J Allergy Clin Immunol 127:965–973
Gao PS, Leung DY, Rafaels NM et al (2012) Genetic variants in interferon regulatory factor 2 (IRF2) are associated with atopic dermatitis and eczema herpeticum. J Investig Dermatol 132:650–657
Esparza-Gordillo J, Schaarschmidt H, Liang L et al (2013) A functional IL-6 receptor (IL6R) variant is a risk factor for persistent atopic dermatitis. J Allergy Clin Immunol 132:371–377
Gharagozlou M, Farhadi E, Khaledi M et al (2013) Association between the interleukin 6 genotype at position -174 and atopic dermatitis. J Investig Allergol Clin Immunol 23:89–93
Kayserova J, Sismova K, Zentsova-Jaresova I et al (2012) A prospective study in children with a severe form of atopic dermatitis: clinical outcome in relation to cytokine gene polymorphisms. J Investig Allergol Clin Immunol 22:92–101
Shin HD, Park BL, Kim LH, Kim JS, Kim JW (2005) Interleukin-10 haplotype associated with total serum IgE in atopic dermatitis patients. Allergy 60:1146–1151
Namkung JH, Lee JE, Kim E et al (2011) An association between IL-9 and IL-9 receptor gene polymorphisms and atopic dermatitis in a Korean population. J Dermatol Sci 62:16–21
Niwa Y, Potaczek DP, Kanada S et al (2010) FcepsilonRIalpha gene (FCER1A) promoter polymorphisms and total serum IgE levels in Japanese atopic dermatitis patients. Int J Immunogenet 37:139–141
Zhou J, Zhou Y, Lin LH et al (2012) Association of polymorphisms in the promoter region of FCER1A gene with atopic dermatitis, chronic uticaria, asthma, and serum immunoglobulin E levels in a Han Chinese population. Hum Immunol 73:301–305
Potaczek DP, Nastalek M, Wojas-Pelc A, Okumura K, Undas A, Nishiyama C (2010) Naturally occurring FCER1A N222K mutation - its ethnicity-dependent distribution and a role in atopic dermatitis. Mol Immunol 48:979–980
Mahachie John JM, Baurecht H, Rodriguez E et al (2010) Analysis of the high affinity IgE receptor genes reveals epistatic effects of FCER1A variants on eczema risk. Allergy 65:875–882
Weidinger S, Gieger C, Rodriguez E et al (2008) Genome-wide scan on total serum IgE levels identifies FCER1A as novel susceptibility locus. PLoS Genet 4:e1000166
Manolio TA (2010) Genomewide association studies and assessment of the risk of disease. N Engl J Med 363:166–176
Trynka G, Hunt KA, Bockett NA et al (2011) Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat Genet 43:1193–1201
Tran DQ, Andersson J, Wang R, Ramsey H, Unutmaz D, Shevach EM (2009) GARP (LRRC32) is essential for the surface expression of latent TGF-beta on platelets and activated FOXP3+ regulatory T cells. Proc Natl Acad Sci U S A 106:13445–13450
Doherty TA, Soroosh P, Khorram N et al (2011) The tumor necrosis factor family member LIGHT is a target for asthmatic airway remodeling. Nat Med 17:596–603
Nair M, Teng A, Bilanchone V, Agrawal A, Li B, Dai X (2006) Ovol1 regulates the growth arrest of embryonic epidermal progenitor cells and represses c-myc transcription. J Cell Biol 173:253–264
Hulin A, Deroanne CF, Lambert CA et al (2012) Metallothionein-dependent up-regulation of TGF-beta2 participates in the remodelling of the myxomatous mitral valve. Cardiovasc Res 93:480–489
Horikawa T, Nakayama T, Hikita I et al (2002) IFN-gamma-inducible expression of thymus and activation-regulated chemokine/CCL17 and macrophage-derived chemokine/CCL22 in epidermal keratinocytes and their roles in atopic dermatitis. Int Immunol 14:767–773
Sugawara N, Yamashita T, Ote Y, Miura M, Terada N, Kurosawa M (2002) TARC in allergic disease. Allergy 57:180–181
Fujita H (2013) The role of IL-22 and Th22 cells in human skin diseases. J Dermatol Sci 72:3–8
Tremblay F, Revett T, Huard C et al (2009) Bidirectional modulation of adipogenesis by the secreted protein Ccdc80/DRO1/URB. J Biol Chem 284:8136–8147
Hsiao HW, Liu WH, Wang CJ et al (2009) Deltex1 is a target of the transcription factor NFAT that promotes T cell anergy. Immunity 31:72–83
Damm A, Lautz K, Kufer TA (2013) Roles of NLRP10 in innate and adaptive immunity. Microbes Infect 15:516–523
Jones G, Prosser DE, Kaufmann M (2014) Cytochrome P450-mediated metabolism of vitamin D. J Lipid Res 55:13–31
Karadag B, Ege MJ, Scheynius A et al (2007) Environmental determinants of atopic eczema phenotypes in relation to asthma and atopic sensitization. Allergy 62:1387–1393
Purvis DJ, Thompson JM, Clark PM et al (2005) Risk factors for atopic dermatitis in New Zealand children at 3.5 years of age. Br J Dermatol 152:742–749
Miyake Y, Okubo H, Sasaki S, Tanaka K, Hirota Y (2011) Maternal dietary patterns during pregnancy and risk of wheeze and eczema in Japanese infants aged 16-24 months: the Osaka Maternal and Child Health Study. Pediatr Allergy Immunol 22:734–741
Wang Y, Liang Y, Lu Q (2008) MicroRNA epigenetic alterations: predicting biomarkers and therapeutic targets in human diseases. Clin Genet 74:307–315
Martino D, Kesper DA, Amarasekera M, Harb H, Renz H, Prescott S (2014) Epigenetics in immune development and in allergic and autoimmune diseases. J Reprod Immunol 104–105:43–48
Prescott S, Saffery R (2011) The role of epigenetic dysregulation in the epidemic of allergic disease. Clin Epigenetics 2:223–232
Fontenot JD, Rasmussen JP, Williams LM, Dooley JL, Farr AG, Rudensky AY (2005) Regulatory T cell lineage specification by the forkhead transcription factor foxp3. Immunity 22:329–341
Liu J, Lluis A, Illi S et al (2010) T regulatory cells in cord blood--FOXP3 demethylation as reliable quantitative marker. PLoS One 5:e13267
Polansky JK, Kretschmer K, Freyer J et al (2008) DNA methylation controls Foxp3 gene expression. Eur J Immunol 38:1654–1663
Leung PS, Shu SA, Chang C (2014) The changing geoepidemiology of food allergies. Clin Rev Allergy Immunol 46:169–179
Ho MH, Wong WH, Chang C (2014) Clinical spectrum of food allergies: a comprehensive review. Clin Rev Allergy Immunol 46:225–240
Shu SA, Chang C, Leung PS (2014) Common methodologies in the evaluation of food allergy: pitfalls and prospects of food allergy prevalence studies. Clin Rev Allergy Immunol 46:198–210
Liang Y, Wang P, Zhao M et al (2012) Demethylation of the FCER1G promoter leads to FcepsilonRI overexpression on monocytes of patients with atopic dermatitis. Allergy 67:424–430
White GP, Watt PM, Holt BJ, Holt PG (2002) Differential patterns of methylation of the IFN-gamma promoter at CpG and non-CpG sites underlie differences in IFN-gamma gene expression between human neonatal and adult CD45RO- T cells. J Immunol 168:2820–2827
Kaminuma O, Kitamura F, Miyatake S et al (2009) T-box 21 transcription factor is responsible for distorted T(H)2 differentiation in human peripheral CD4+ T cells. J Allergy Clin Immunol 123:813–823
Brand S, Teich R, Dicke T et al (2011) Epigenetic regulation in murine offspring as a novel mechanism for transmaternal asthma protection induced by microbes. J Allergy Clin Immunol 128:618–625
Ly NP, Litonjua A, Gold DR, Celedon JC (2011) Gut microbiota, probiotics, and vitamin D: interrelated exposures influencing allergy, asthma, and obesity? J Allergy Clin Immunol 127:1087–1094, quiz 1095-1086
Cao S, Feehley TJ, Nagler CR (2014) The role of commensal bacteria in the regulation of sensitization to food allergens. FEBS Lett
Amberbir A, Medhin G, Erku W et al (2011) Effects of Helicobacter pylori, geohelminth infection and selected commensal bacteria on the risk of allergic disease and sensitization in 3-year-old Ethiopian children. Clin Exp Allergy 41:1422–1430
Kummeling I, Stelma FF, Dagnelie PC et al (2007) Early life exposure to antibiotics and the subsequent development of eczema, wheeze, and allergic sensitization in the first 2 years of life: the KOALA Birth Cohort Study. Pediatrics 119:e225–e231
Ochoa-Reparaz J, Mielcarz DW, Haque-Begum S, Kasper LH (2010) Induction of a regulatory B cell population in experimental allergic encephalomyelitis by alteration of the gut commensal microflora. Gut Microbes 1:103–108
Hill DA, Siracusa MC, Abt MC et al (2012) Commensal bacteria-derived signals regulate basophil hematopoiesis and allergic inflammation. Nat Med 18:538–546
Perera F, Tang WY, Herbstman J et al (2009) Relation of DNA methylation of 5′-CpG island of ACSL3 to transplacental exposure to airborne polycyclic aromatic hydrocarbons and childhood asthma. PLoS One 4:e4488
Tang WY, Levin L, Talaska G et al (2012) Maternal exposure to polycyclic aromatic hydrocarbons and 5′-CpG methylation of interferon-gamma in cord white blood cells. Environ Health Perspect 120:1195–1200
Nurmatov U, Devereux G, Sheikh A (2011) Nutrients and foods for the primary prevention of asthma and allergy: systematic review and meta-analysis. J Allergy Clin Immunol 127:724–733
Joseph CL, Ownby DR, Havstad SL et al (2011) Early complementary feeding and risk of food sensitization in a birth cohort. J Allergy Clin Immunol 127:1203–1210
Hollingsworth JW, Maruoka S, Boon K et al (2008) In utero supplementation with methyl donors enhances allergic airway disease in mice. J Clin Invest 118:3462–3469
Brown SB, Reeves KW, Bertone-Johnson ER (2014) Maternal folate exposure in pregnancy and childhood asthma and allergy: a systematic review. Nutr Rev 72:55–64
Bekkers MB, Elstgeest LE, Scholtens S et al (2012) Maternal use of folic acid supplements during pregnancy, and childhood respiratory health and atopy. Eur Respir J 39:1468–1474
Matsui EC, Matsui W (2009) Higher serum folate levels are associated with a lower risk of atopy and wheeze. J Allergy Clin Immunol 123:1253–1259
Whitrow MJ, Moore VM, Rumbold AR, Davies MJ (2009) Effect of supplemental folic acid in pregnancy on childhood asthma: a prospective birth cohort study. Am J Epidemiol 170:1486–1493
Husemoen LL, Toft U, Fenger M, Jorgensen T, Johansen N, Linneberg A (2006) The association between atopy and factors influencing folate metabolism: is low folate status causally related to the development of atopy? Int J Epidemiol 35:954–961
Kiefte-de Jong JC, Timmermans S, Jaddoe VW et al (2012) High circulating folate and vitamin B-12 concentrations in women during pregnancy are associated with increased prevalence of atopic dermatitis in their offspring. J Nutr 142:731–738
Thuesen BH, Husemoen LL, Ovesen L et al (2010) Atopy, asthma, and lung function in relation to folate and vitamin B(12) in adults. Allergy 65:1446–1454
Sugiura H, Ebise H, Tazawa T et al (2005) Large-scale DNA microarray analysis of atopic skin lesions shows overexpression of an epidermal differentiation gene cluster in the alternative pathway and lack of protective gene expression in the cornified envelope. Br J Dermatol 152:146–149
Jarzab J, Filipowska B, Zebracka J et al (2010) Locus 1q21 Gene expression changes in atopic dermatitis skin lesions: deregulation of small proline-rich region 1A. Int Arch Allergy Immunol 151:28–37
Bin L, Howell MD, Kim BE, Streib JE, Hall CF, Leung DY (2011) Specificity protein 1 is pivotal in the skin’s antiviral response. J Allergy Clin Immunol 127:430–438
Soumelis V, Reche PA, Kanzler H et al (2002) Human epithelial cells trigger dendritic cell mediated allergic inflammation by producing TSLP. Nat Immunol 3:673–680
Luo Y, Zhou B, Zhao M, Tang J, Lu Q (2014) Promoter demethylation contributes to TSLP overexpression in skin lesions of patients with atopic dermatitis. Clin Exp Dermatol 39:48–53
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Makeyev EV, Maniatis T (2008) Multilevel regulation of gene expression by microRNAs. Science 319:1789–1790
Djuranovic S, Nahvi A, Green R (2011) A parsimonious model for gene regulation by miRNAs. Science 331:550–553
Janssen HL, Reesink HW, Lawitz EJ et al (2013) Treatment of HCV infection by targeting microRNA. N Engl J Med 368:1685–1694
Sonkoly E, Wei T, Janson PC et al (2007) MicroRNAs: novel regulators involved in the pathogenesis of psoriasis? PLoS One 2:e610
Sonkoly E, Janson P, Majuri ML et al (2010) MiR-155 is overexpressed in patients with atopic dermatitis and modulates T-cell proliferative responses by targeting cytotoxic T lymphocyte-associated antigen 4. J Allergy Clin Immunol 126:581–589
Bieber T (2012) Atopic dermatitis 2.0: from the clinical phenotype to the molecular taxonomy and stratified medicine. Allergy 67:1475–1482
Acknowledgments
This study was supported by the National Natural Science Foundation of China (Nos. 30800993, 81371742, and 81573052).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liang, Y., Chang, C. & Lu, Q. The Genetics and Epigenetics of Atopic Dermatitis—Filaggrin and Other Polymorphisms. Clinic Rev Allerg Immunol 51, 315–328 (2016). https://doi.org/10.1007/s12016-015-8508-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-015-8508-5