Abstract
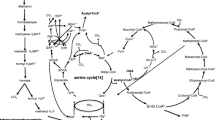
Pseudomonas sp. SMIC-3 grown on NMP was physiologically differentiated from that on glucose. Growth of SMIC-3 in an NMP-defined medium was approximately three times lower than that in a glucose-defined medium. Methylamine and 1-methyl succinimide were detected in culture fluid of SMIC-3 grown in an NMP-defined medium. Methylamine content in the culture fluid was very similar to NMP consumed by SMIC-3, but 1-methyl succinimide content was much less than the consumed NMP. Crude enzyme isolated from SMIC-3 grown on NMP catalyzed production of methylamine, 1-methyl succinimide, and succinate from NMP but that on glucose did not. Crude enzyme isolated from SMIC-3 grown on glucose and NMP commonly catalyzed dehydrogenation of pyruvate, isocitrate, and malate coupled to reduction of NAD+ to NADH. 2D-SDS-PAGE pattern of total soluble proteins isolated from SMIC-3 grown on glucose was significantly different from that on NMP. Physiological function of SMIC-3 for catabolizing NMP may be selectively induced and activated by NMP.
Similar content being viewed by others
References
M. Reisch, Chem. Eng. News, 86, 32 (2008).
J.R. Van der Meer, W. Roelofsen, G. Schraa and A. J. B. Zehner, FEMS Microbiol. Ecol., 45, 333 (1987).
W. J. Lee, J. K. Lee, J. Chung, Y. J. Cho and D. H. Park, J. Microbiol. Biotechnol., 20, 1230 (2010).
S. T. Chow and T. L. Ng, Water Res., 17, 117 (1983).
M.J. Huertas, V.M. Luque-Almagro, M. Martínez-Luque, R. Blasco, C. Moreno-Vivián, F. Castillo and M. D. Roldán, Biochem. Soc. Trans., 34, 152 (2006).
I. H. Nam, Y. S. Chang, H.B. Hong and Y. E. Lee, Appl. Microbiol. Biotechnol., 62, 284 (2003).
H. Nojiri, K. Maeda, H. Sekiguchi, M. Urata, M. Shintani, T. Yoshida, H. Habe and T. Omori, Biosci. Biotechnol. Biochem., 66, 897 (2002).
M.M. O’Mahony, A. D. Dobson, J. D. Barnes and I. Singleton, Chemosphere, 63, 307 (2006).
C. Onaca, M. Kieninger, K. H. Engesser and J. Altenbuchner, J. Bacteriol., 189, 3759 (2007).
K.M. Yen, M. R. Karl and L.M. Blatt, J. Bacteriol., 173, 5315 (1991).
H. Ochman and N. A. Moran, Science, 292, 1096 (2001).
A. Daddaoua, T. Krell and J. L. Ramos, J. Biol. Chem., 32, 21360 (2009).
H. S. Shin and D.G. Jung, Bull. Kor. Chem. Soc., 25, 806 (2004).
J. C. Wilkins, K. A. Homer and D. Beighton, Appl. Environ. Microbiol., 67, 3396 (2001).
J. K. Lee, W. J. Lee, Y. J. Cho, D. H. Park, Y.W. Lee and J.W. Chung, Korean J. Chem. Eng., 27, 1816 (2010).
T. Ferenci, Adv. Microb. Physiol., 53, 169 (2008).
A. Haibi and F. Vahabzadeh, J. Environ. Sci. Health A Tox. Hazard. Subst. Environ. Eng., 48, 279 (2013).
A. Niqam, P. S. Phale and P. P. Wanqikar, Biorersour. Technol., 114, 484 (2012).
F. Duffes, P. Jenoe and P. Boyaval, Appl. Environ. Microbiol., 66, 4318 (2000).
D. R. Lovley and M. J. Klug, Appl. Environ. Microbiol., 45, 1310 (1983).
J. J. Park, S.Y. Jung, C.G. Park, S. C. Lee, J. N. Kim and J. C. Kim, Korean J. Chem. Eng., 29, 489 (2012).
D. Zhang, Y. Ma, H. Feng, H. Luo, J. Chen and Y. Hao, Korean J. Chem. Eng., 29, 769 (2012).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jeon, B.Y., Yi, J.Y. & Park, D.H. Estimation on metabolic pathway of Pseudomonas sp. SMIC-3 for 1-methyl-2-pyrrolidinone based on physiological and biochemical analyses. Korean J. Chem. Eng. 31, 475–484 (2014). https://doi.org/10.1007/s11814-013-0231-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11814-013-0231-4