Abstract
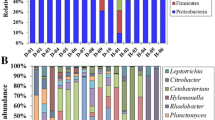
The gut microbiota is closely related to health and disease. Grass carp (Ctenopharyngodon idella) is an important food fish in China. We aimed to investigate the effect of a chicken faeces diet on the gut microbiota composition of grass carp reared in an integrated farming system in China. Gut microbiota compositions of grass carps fed chicken faeces, a commercial diet, and grass were compared based on 16S rRNA gene sequencing. The major intestinal phyla in grass carps fed chicken faeces were Firmicutes, Proteobacteria, and Actinobacteria. The untreated chicken faeces diet altered the gut microbiota composition and increased the number of potential pathogens and antibiotic-resistant bacteria in the gut to varying degrees. To reduce the risk of diseases, it is necessary to remove residual antibiotics and antibiotic-resistant bacteria in chicken faeces by fermentation or other techniques, before it can be used as a fish feed for grass carp.





Similar content being viewed by others
References
Awasthi MK, Chen H, Duan Y, Liu T, Awasthi SK, Wang Q, Pandey A, Zhang Z (2019) An assessment of the persistence of pathogenic bacteria removal in chicken manure compost employing clay as additive via meta-genomic analysis. J Hazard Mater 366:184–191
Byard RW, Moore L, Bourne AJ, Lawrence AJ, Goldwater PN (2010) Clostridium botulinum and sudden infant death syndrome: a 10-year prospective study. J Paediatr Child Health 28:156–157
Carlson K, Yang S (2003) Evolution of antibiotic occurrence in a river through pristine, urban and agricultural landscapes. Water Res 37:4645–4656
Chen Z, Jiang X (2014) Microbiological safety of chicken litter or chicken litter-based organic fertilizers: a review. Agriculture 4:1–29
Desai AR, Links MG, Collins SA, Mansfield GS, Drew MD, Kessel AGV, Hill JE (2012) Effects of plant-based diets on the distal gut microbiome of rainbow trout (Oncorhynchus mykiss). Aquaculture 350-353:134–142
Duan D-X, Shu-Yun L, Jing-Sheng S, Guo-Si Z, and Gui-Ping Z (1998) Fish farming integrated with livestock and poultry in China. Integr Fish Farm 73–82
Goldman E (2004) Antibiotic abuse in animal agriculture: exacerbating drug resistance in human pathogens. Hum Ecol Risk Assess 10:121–134
Guerrero Iii RD, Guerrero L, Alamar M (1988) Polyculture of Nile tilapia, grass carp and common carp in freshwater ponds. Trans Nat Acad Sci Tech (Phils) 10:365–368
Guinane CM, Cotter PD (2013) Role of the gut microbiota in health and chronic gastrointestinal disease: understanding a hidden metabolic organ. Ther Adv Gastroenterol 6:295–308
Hale CN, Wilkie JP (1972) A comparative study of Pseudomonas species pathogenic to sorghum. N Z J Agric Res 15:448–456
Han S, Liu Y, Zhou Z, He S, Cao Y, Shi P, Yao B, Ring E (2010) Analysis of bacterial diversity in the intestine of grass carp (Ctenopharyngodon idellus) based on 16S rDNA gene sequences. Aquac Res 42:47–56
Huang H, Shi P, Wang Y, Luo H, Shao N, Wang G, Yang P, Yao B (2009) Diversity of beta-propeller phytase genes in the intestinal contents of grass carp provides insight into the release of major phosphorus from phytate in nature. Appl Environ Microbiol 75:1508–1516
Icaza-Chávez ME (2013) Gut microbiota in health and disease. Rev Gastroenterol Mex 78:240–248
Jorgensen JH, Hindler JF, Reller LB, Weinstein MP (2007) New consensus guidelines from the Clinical and Laboratory Standards Institute for antimicrobial susceptibility testing of infrequently isolated or fastidious bacteria. Clin Infect Dis 44:280–286
Levine UY, Stanton T (2015) Poultry intestinal microbiota: animal health and food safety perspectives. In: Highlander SK, Rodriguez-Valera F, White B (eds) Encyclopedia of Metagenomics. Springer, Boston, MA, pp 1–8
Li J, Ni J, Li J, Wang C, Li X, Wu S, Zhang T, Yu Y, Yan Q (2014) Comparative study on gastrointestinal microbiota of eight fish species with different feeding habits. J Appl Microbiol 117:1750–1760
Li T, Long M, Gatesoupe FJ, Zhang Q, Li A, Gong X (2015) Comparative analysis of the intestinal bacterial communities in different species of carp by pyrosequencing. Microb Ecol 69:25–36
Mitzner L (1978) Evaluation of biological control of nuisance aquatic vegetation by grass carp. Trans Am Fish Soc 107:135–145
Ni J, Yu Y, Zhang T, Gao L, Ni J, Yu Y, Zhang T, Gao L (2012) Comparison of intestinal bacterial communities in grass carp, Ctenopharyngodon idellus, from two different habitats. Chin J Oceanol Limnol 30:757–765
Ni J, Yan Q, Yu Y, Zhang T (2014) Factors influencing the grass carp gut microbiome and its effect on metabolism. FEMS Microbiol Ecol 87:704–714
Nicholson FA, Chambers BJ, Smith KA (1996) Nutrient composition of poultry faeces in England and Wales. Bioresour Technol 58:279–284
Ouyang WY, Huang FY, Zhao Y, Li H, Su JQ (2015) Increased levels of antibiotic resistance in urban stream of Jiulongjiang River, China. Appl Microbiol Biotechnol 99:5697–5707
Pond MJ, Stone DM, Alderman DJ (2006) Comparison of conventional and molecular techniques to investigate the intestinal microflora of rainbow trout (Oncorhynchus mykiss). Aquaculture 261:194–203
Russo P, Iturria I, Mohedano ML, Caggiaaniello G, Rainieri S, Fiocco D, Pardo MA, Lopez P, Spano G (2015) Zebrafish gut colonization by mCherry-labelled lactic acid bacteria. Appl Microbiol Biotechnol 99:3479–3490
Sahoo U, Singh S (2015) Integrated fish-pig and fish-poultry farming in East Kalcho, Saiha District of Mizoram, North-East India: an economic analysis. Inte J Agric For 5:281–286
Sanz Y, De Palma G (2009) Gut microbiota and probiotics in modulation of epithelium and gut-associated lymphoid tissue function. Int Rev Immunol 28:397–413
Schloss PD, Westcott SL, Thomas R, Hall JR, Martin H, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schloss PD, Gevers D, and Westcott SL (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PloS One 6
Sekirov I, Russell SL, Antunes LCM, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90:859–904
Tran NT, Wang GT, Wu SG (2017) A review of intestinal microbes in grass carp Ctenopharyngodon idellus (Valenciennes). Aquac Res 48:3287–3297
Tran NT, Zhang J, Xiong F, Wang G-T, Li W-X, Wu S-G (2018) Altered gut microbiota associated with intestinal disease in grass carp (Ctenopharyngodon idellus). World J Microbio Biotech 34:71
Wang X (2013) Analysis on nutrient contents of chicken faeces and its utilization and development. Hubei Agric Sci 52:5314–5316
Wang AR, Ran C, Ring E, Zhou ZG (2018) Progress in fish gastrointestinal microbiota research. Rev Aquac 10:626–640
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H, Wu S, Wang G, Angert ER, Wang W (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PLoS One 7:e30440
Wu S, Gao T, Zheng Y, Wang W, Cheng Y, Wang G (2016) Microbial diversity of intestinal contents and mucus in yellow catfish (Pelteobagrus fulvidraco). Aquaculture 303:1–7
Yang Q, Ren S, Niu T, Guo Y, Qi S, Han X, Liu D, Pan F (2014) Distribution of antibiotic-resistant bacteria in chicken faeces and faeces-fertilized vegetables. Environ Sci Pollut Res Int 21:1231–1241
Yang Q, Wang R, Ren S, Szoboszlay M, Moe LA (2016) Practical survey on antibiotic-resistant bacterial communities in livestock faeces and faeces-amended soil. J Environ Sci Health B 51:14–23
Yang Q, Tian T, Niu T, Wang P (2017) Molecular characterization of antibiotic resistance in cultivable multidrug-resistant bacteria from livestock manure. Environ Pollut 229:188–198
Ye L, Amberg J, Chapman D, Gaikowski M, Liu WT (2014) Fish gut microbiota analysis differentiates physiology and behavior of invasive Asian carp and indigenous American fish. ISME J 8:541–551
Zarkasi KZ, Abell GC, Taylor RS, Neuman C, Hatje E, Tamplin ML, Katouli M, Bowman JP (2014) Pyrosequencing-based characterization of gastrointestinal bacteria of Atlantic salmon (Salmo salar L.) within a commercial mariculture system. J Appl Microbiol 117:18–27
Zhang X (2018) China fishery statistical yearbook. China Agriculture Press, Beijing
Zhang QL, Yan Y, Shen JY, Hao GJ, Shi CY, Wang QT, Liu H, Huang J (2013) Development of a reverse transcription loop-mediated isothermal amplification assay for rapid detection of grass carp reovirus. J Virol Methods 187:384–389
Funding
This work was supported by the Scientific Research Foundation for Talent Introduction of Southwest Medical University (0903-00040071) and Science Fund Project of Southwest Medical University (0903-00031532).
Author information
Authors and Affiliations
Contributions
XG, CZ, and LL designed the experiments. KF, YG, XG conducted the experiments. XG and QW wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
No protected species were involved in this study, and thus, this study did not require specific permissions. The Southwest Medical University Ethics Committee reviewed and approved this study.
Additional information
Responsible Editor: Philippe Garrigues
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 732 kb)
Rights and permissions
About this article
Cite this article
Ke, F., Gao, Y., Liu, L. et al. Comparative analysis of the gut microbiota of grass carp fed with chicken faeces. Environ Sci Pollut Res 27, 32888–32898 (2020). https://doi.org/10.1007/s11356-020-09012-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-020-09012-8