Abstract
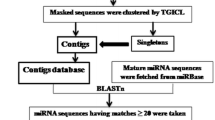
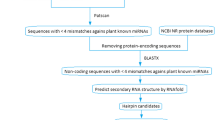
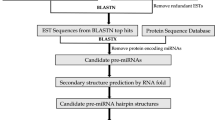
MicroRNAs (miRNAs) are a group of single-stranded noncoding RNAs with general size of 21–24 nucleotides, which negatively regulate gene expression posttranscriptionally by repressing gene translation or degrading targeted mRNAs. Despite the important functions of miRNAs in plants, little is known about miRNAs in citrus. Here we present a study of bioinformatics identification of microRNA precursors in citrus by comparing known plant miRNAs against 560,271 citrus ESTs. Seventy-eight ESTs from ten citrus species and three hybrids were identified as putative miRNA precursors, encoding 51 unique miRNA sequences, representing 28 miRNA families. Blastn search of the putative miRNAs against citrus unigenes and ESTs was performed to identify the target genes. As a result, 36 unigenes and 77 ESTs were identified as putative targets of 25 citrus miRNA families. The putative targets are mainly transcription factors and play important roles in development, metabolism, and stress resistance. To validate the predicted miRNAs, qRT-PCR was applied to detect the tissue-specific expression of nine putative miRNAs in Citrus sinensis cv. Valencia. The cleavage sites on five mRNAs targeted by three miRNA families were mapped by 5′RACE. This study provided information on citrus miRNA precursors, mature miRNAs, and miRNA targets and is helpful for future research of miRNA function in citrus.







Similar content being viewed by others
Abbreviations
- EST:
-
expressed sequence tag
- miRNA:
-
microRNA
- miRNA*:
-
passenger strand on precursor
- Pre-miRNA:
-
microRNA precursor
- MFE:
-
negative minimal free energies
- MFEi:
-
minimal free energy index
- qRT-PCR:
-
quantitative reverse transcription-polymerase chain reation
- 5′RACE:
-
5′ rapid amplification of cDNA ends
References
Achard P, Herr A, Baulcombe DC, Harberd NP (2004) Modulation of floral development by a gibberellin-regulated microRNA. Development 131:3357–3365
Adenot X, Elmayan T, Lauressergues D, Boutet S, Bouch N, Gasciolli V, Vaucheret H (2006) DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Current Biology 16:927–932
Arazi T, Talmor-Neiman M, Stav R, Riese M, Huijser P, Baulcombe DC (2005) Cloning and characterization of micro-RNAs from moss. Plant J 43:837–848
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Bao N, Lye KW, Barton MK (2004) MicroRNA binding sites in Arabidopsis class III HD-ZIP mRNAs are required for methylation of the template chromosome. Developmental Cell 7:653–662
Barakat A, Wall K, Leebens-Mack J, Wang YJ, Carlson JE, dePamphilis CW (2007) Large-scale identification of microRNAs from a basal eudicot (Eschscholzia californica) and conservation in flowering plants. Plant J 51:991–1003
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Baumberger N, Baulcombe DC (2005) Arabidopsis ARGONAUTE1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102:11928–11933
Brodersen P, Voinnet O (2009) Revisiting the principles of microRNA target recognition and mode of action. Nature Reviews Mol Cell Biology 10:141–148
Carra A, Mica E, Gambino G, Pindo M, Moser C, Enrico MP, Schubert A (2009) Cloning and characterization of small non-coding RNAs from grape. Plant J 59:750–63
Chen XM (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ (2005) Real-time quantification of microRNAs by stem-loop RT-PCR. Nucl Acids Res 33(20):e179
Chiou TJ, Aung K, Lin SI, Wu CC, Chiang SF, Cl Su (2006) Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 18:412–421
Chuck G, Cigan AM, Saeteurn K, Hake S (2007) The heterochronic maize mutant Corngrass1 results from overexpression of a tandem microRNA. Nat Genet 39:544–549
Chuck G, Candela H, Hake S (2009) Big impacts by small RNAs in plant development. Current Opinion in Plant Biology 12:81–86
Dezulian T, Palatnik J, Huson D, Weigel D (2005) Conservation and divergence of microRNA families in plants. Genome Biology 6:P13
Fahlgren N, Howell MD, Kasschau KD, Chapman EJ, Sullivan CM, Cumbie JS, Givan SA, Law TF, Grant SR, Dangl JL, Carrington JC (2007) High-throughput sequencing of Arabidopsis microRNAs: evidence for frequent birth and death of MIRNA genes. PLoS ONE 2:e219
Fujii H, Chiou TJ, Lin SI, Aung K, Zhu JK (2005) A miRNA involved in phosphate-starvation response in Arabidopsis. Current Biology 15:2038–2043
Fujioka T, Kaneko F, Kazama T, Suwabe K, Suzuki G, Makino A, Mae T, Endo M, Kawagishi-Kobayashi M, Watanabe M (2008) Identification of small RNAs in late developmental stage of rice anthers. Genes & Genetic Systems 83:281–284
Gandikota M, Birkenbihl RP, Höhmann SH, Cardon GH, Heinz S, Huijser P (2007) The miRNA156/157 recognition element in the 3′ UTR of the Arabidopsis SBP box gene SPL3 prevents early flowering by translational inhibition in seedlings. Plant J 49:683–693
Garcia D, Collier SA, Byrne ME, Martienssen RA (2006) Specification of leaf polarity in Arabidopsis via the trans-acting siRNA pathway. Current Biology 16:933–938
Gleave A, Ampomah-Dwamena C, Berthold S, Dejnoprat S, Karunairetnam S, Nain B, Wang YY, Crowhurst R, MacDiarmid R (2008) Identification and characterisation of primary microRNAs from apple (Malus domestica cv. Royal Gala) expressed sequence tags. Tree Genetics & Genomes 4:343–358
Goudeau D, Uratsu SL, Inoue K, daSilva FG, Leslie A, Cook D, Reagan R, Dandekar AM (2008) Turning the orchestra: selective gene regulation and orange fruit quality. Plant Sci 174:310–320
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucl Acids Res 36:D154–158
Guo WW, Li DL, Duan YX (2007) Citrus transgenics: current status and prospects. Transgenic Plant J 1:202–209
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Juarez MT, Kui JS, Thomas J, Heller BA, Timmermans MCP (2004) MicroRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature 428:84–88
Jung JH, Park CM (2007) MIR166/165 genes exhibit dynamic expression patterns in regulating shoot apical meristem and floral development in Arabidopsis. Planta 225:1327–1338
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim VN, Chua NH, Park CM (2005) MicroRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42:84–94
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20:2238–2251
Liu Y, Liu Q, Tao NG, Deng XX (2006) Efficient isolation of RNA from fruit peel and pulp of ripening navel orange (Citrus sinensis Osbeck). J Huazhong Agric Univ 25:300–304
Liu Q, Xu J, Liu Y, Zhao X, Deng X, Guo L, Gu J (2007) A novel bud mutation that confers abnormal patterns of lycopene accumulation in sweet orange fruit (Citrus sinensis L. Osbeck). J Exp Bot 58:4161–4171
Liu N, Okamura K, Tyler DM, Phillips MD, Chung WJ, Lai EC (2008) The evolution and functional diversification of animal microRNA genes. Cell Res 18:985–996
Llave C, Xie ZX, Kasschau KD, Carrington JC (2002) Cleavage of scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 297:2053–2056
Lu C, Tej SS, Luo S, Haudenschild CD, Meyers BC, Green PJ (2005a) Elucidation of the small RNA component of the transcriptome. Science 309:1567–1569
Lu S, Sun YH, Shi R, Clark C, Li L, Chiang VL (2005b) Novel and mechanical stress-responsive microRNAs in Populus trichocarpa that are absent from Arabidopsis. Plant Cell 17:2186–2203
Lu C, Kulkarni K, Souret FF, MuthuValliappan R, Tej SS, Poethig RS, Henderson IR, Jacobsen SE, Wang WZ, Green PJ, Meyers BC (2006) MicroRNAs and other small RNAs enriched in the Arabidopsis RNA-dependent RNA polymerase-2 mutant. Genome Res 16:1276–1288
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17:1360–1375
McHale NA, Koning RE (2004) MicroRNA-directed cleavage of Nicotiana sylvestris PHAVOLUTA mRNA regulates the vascular cambium and structure of apical meristems. Plant Cell 16:1730–1740
Meyers BC, Axtell MJ, Bartel B, Bartel DP, Baulcombe D, Bowman JL, Cao X, Carrington JC, Chen X, Green PJ, Griffiths-Jones S, Jacobsen SE, Mallory AC, Martienssen RA, Poethig RS, Qi Y, Vaucheret H, Voinnet O, Watanabe Y, Weigel D, Zhu J-K (2008) Criteria for annotation of plant microRNAs. Plant Cell 20:3186–3190
Millar AA, Gubler F (2005) The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 17:705–721
Montgomery TA, Howell MD, Cuperus JT, Li D, Hansen JE, Alexander AL, Chapman EJ, Fahlgren N, Allen E, Carrington JC (2008) Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133:128–141
Ori N, Cohen AR, Etzioni A, Brand A, Yanai O, Shleizer S, Menda N, Amsellem Z, Efroni I, Pekker I, Alvarez JP, Blum E, Zamir D, Eshed Y (2007) Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet 39:787–791
Palatnik JF, Allen E, Wu XL, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263
Palatnik JF, Wollmann H, Schommer C, Schwab R, Boisbouvier J, Rodriguez R, Warthmann N, Allen E, Dezulian T, Huson D, Carrington J, Weigel D (2007) Sequence and expression differences underlie functional specialization of Arabidopsis microRNAs miR159 and miR319. Developmental Cell 13:115–125
Qiu CX, Xie FL, Zhu YY, Guo K, Huang SQ, Nie L, Yang ZM (2007) Computational identification of microRNAs and their targets in Gossypium hirsutum expressed sequence tags. Gene 395:49–61
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP (2002) MicroRNAs in plants. Genes & Development 16:1616–1626
Roose ML, Close TJ (2008) Genomics of citrus, a major fruit crop of tropical and subtropical regions. In: Moore PH, Ming R (eds) Genomics of tropical crop plants, Springer, pp. 187–201
Ru P, Xu L, Ma H, Huang H (2006) Plant fertility defects induced by the enhanced expression of microRNA167. Cell Res 16:457–465
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Developmental Cell 8:517–527
Sieber P, Wellmer F, Gheyselinck J, Riechmann JL, Meyerowitz EM (2007) Redundancy and specialization among plant microRNAs: role of the MIR164 family in developmental robustness. Development 134:1051–1060
Smita R, Thomas G, Alexis P, Thomas B, Patrick L, Klaus T (2008) Interplay of miR164, CUP-SHAPED COTYLEDON genes and LATERAL SUPPRESSOR controls axillary meristem formation in Arabidopsis thaliana. Plant J 55:65–76
Song C, Fang J, Li X, Liu H, Chao CT (2009) Identification and characterization of 27 conserved microRNAs in citrus. Planta 230:671–685
Sunkar R, Jagadeeswaran G (2008) In silico identification of conserved microRNAs in large number of diverse plant species. BMC Plant Biology 8:37
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Sunkar R, Girke T, Jain PK, Zhu JK (2005) Cloning and characterization of microRNAs from rice. Plant Cell 17:1397–1411
Talon M, Gmitter FG Jr (2008) Citrus genomics. International Journal of Plant Genomics. 1687–5370
Varkonyi-Gasic E, Wu R, Wood M, Walton E, Hellens R (2007) Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 3:12
Vaucheret H, Mallory AC, Bartel DP (2006) AGO1 homeostasis entails coexpression of MIR168 and AGO1 and preferential stabilization of miR168 by AGO1. Mol Cell 22:129–136
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17:2204–2216
Wang L, Wang MB, Tu JX, Helliwell CA, Waterhouse PM, Dennis ES, Fu TD, Fan YL (2007) Cloning and characterization of microRNAs from Brassica napus. FEBS Lett 581:3848–3856
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132:3657–3668
Wu G, Poethig RS (2006) Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 133:3539–3547
Wu MF, Tian Q, Reed JW (2006) Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133:4211–4218
Xie Z, Kasschau KD, Carrington JC (2003) Negative feedback regulation of Dicer-like1 in Arabidopsis by microRNA-guided mRNA degradation. Current Biology 13:784–789
Xie K, Wu C, Xiong L (2006) Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol 142:280–293
Yin Z, Li C, Han X, Shen F (2008) Identification of conserved microRNAs and their target genes in tomato (Lycopersicon esculentum). Gene 414:60–66
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2005) Identification and characterization of new plant microRNAs using EST analysis. Cell Res 15:336–360
Zhang B, Pan X, Cannon CH, Cobb GP, Anderson TA (2006a) Conservation and divergence of plant microRNA genes. Plant J 46:243–259
Zhang BH, Pan XP, Anderson TA (2006b) Identification of 188 conserved maize microRNAs and their targets. FEBS Lett 580:3753–3762
Zhang BH, Wang QL, Wang KB, Pan XP, Liu F, Guo TL, Cobb GP, Anderson TA (2007) Identification of cotton microRNAs and their targets. Gene 397:26–37
Zhang BH, Pan XP, Stellwag EJ (2008a) Identification of soybean microRNAs and their targets. Planta 229:161–182
Zhang J, Zeng R, Chen J, Liu X, Liao Q (2008b) Identification of conserved microRNAs and their targets from Solanum lycopersicum Mill. Gene 423:1–7
Acknowledgements
This research was financially supported by the National Natural Science Foundation (No. 30921002), the Ministry of Agriculture of China (No. 200903044-3), the Fundamental Research Funds for the Central Universities (No. 2009PY022) and the Hubei provincial Natural Science Foundation (No. 2008CDA069). The authors thank Dr. Han-Hui Kuang (Huazhong Agric. Univ.) for his critical reading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. Gmitter
Rights and permissions
About this article
Cite this article
Wu, XM., Liu, MY., Xu, Q. et al. Identification and characterization of microRNAs from citrus expressed sequence tags. Tree Genetics & Genomes 7, 117–133 (2011). https://doi.org/10.1007/s11295-010-0319-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-010-0319-5