Abstract
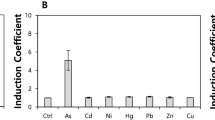
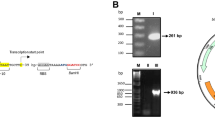
A number of human activities result in environmental contamination with copper compounds that can cause severe detrimental effects on the ecosystem as well as human health. The physico-chemical methods of metal detection have limitations such as inability to distinguish between total versus bio-available metals and differences in metal uptake in different organisms. The heavy metal resistance-encoding genetic systems of certain bacteria provide critical tools for development of biosensors for these purposes. This study reports a copper biosensor utilizing the cop operon of the heavy metal resistant bacterial isolate, Achromobacter sp. AO22, isolated from a contaminated site in Australia. A section located between the divergently transcribed putative response regulator gene copR and multicopper oxidase gene copA that included a palindromic cop box was identified as a copper-responsive promoter using a lacZ reporter construct, pCOPRP, in E. coli. The expression was found to be enhanced by inclusion of copR. Another engineered strain, AO22(pCOPRP), showed stronger induction, and the lacZ expression in both backgrounds was enhanced significantly (250–400 fold) by copper but minimally by other metals. The construct in Achromobacter sp. AO22 thus has a high potential as biosensor for detecting copper bioavailability (hence potential toxicity) in a soil bacterial background, while the construct in E. coli is ideal for laboratory-based testing.



Similar content being viewed by others
References
Ashley P, Lottermoser B (1999) Arsenic contamination at the Mole River mine, northern New South Wales. Austr J Earth Sci 46(6):861–874
Basim H, Minsavage GV, Stall RE, Wang J-F, Shanker S, Jones JB (2005) Characterisation of a unique chromosomal copper resistance gene cluster from Xanthomonas campestris pv. vesicatoria. Appl Environ Microbiol 71(12):8284–8291
Bondarenko O, RÃμlova T, Kahru A, Ivask A (2008) Bioavailability of Cd, Zn and Hg in soil to nine recombinant luminescent metal sensor bacteria. Sensors 8(11):6899–6923
Corbisier P, van der Lelie D, Borremans B, Provoost A, de Lorenzo V, Brown NL, Lloyd JR, Hobman JL, Csoregi E, Johansson G, Mattiasson B (1999) Whole cell- and protein-based biosensors for the detection of bioavailable heavy metals in environmental samples. Anal Chimi Acta 387(3):235–244
Diels L, Van Roy S, Taghavi S, Van Houdt R (2009) From industrial sites to environmental applications with Cupriavidus metallidurans. Antonie Van Leeuwenhoek 96(2):247–258
Dorsey A, Ingerman L (2004) Toxicology profile for copper. In US Department of Health and Human Services. Atlanta: Agency for toxic substances and disease registry
Gupta A, Matsui K, Lo JF, Silver S (1999) Molecular basis for resistance to silver cations in Salmonella. Nat Med 5(2):183–188
Hakkila K, Green T, Leskinen P, Ivask A, Marks R, Virta M (2004) Detection of bioavailable heavy metals in EILATox-Oregon samples using whole-cell luminescent bacterial sensors in suspension or immobilized onto fibre-optic tips. J Appl Toxicol 24(5):333–342
Harley CB, Reynolds RP (1987) Analysis of E. coli promoter sequences. Nucl Acids Res 15(5):2343–2361
Hettler J, Irion G, Lehmann B (1997) Environmental impact of mining waste disposal on a tropical lowland river system: a case study on the Ok Tedi Mine, Papua New Guinea. Miner Deposita 32(3):280–291
Leth S, Maltoni S, Simkus R, Mattiasson B, Corbisier P, Klimant I, Wolfbeis Otto S, Csöregi E (2002) Engineered bacteria based biosensors for monitoring bioavailable heavy metals. Electroanalysis 14(1):35–42
Liao VH-C, Chien M-T, Tseng Y-Y, Ou K-L (2006) Assessment of heavy metal bioavailability in contaminated sediments and soils using green fluorescent protein-based bacterial biosensors. Environ Pollut 142(1):17–23
Miller J (1972) Experiments in molecular genetics. Cold Spring Harbor, New York
Mills SD, Lim CK, Cooksey DA (1994) Purification and characterization of CopR, a transcriptional activator protein that binds to a conserved domain (cop box) in copper-inducible promoters of Pseudomonas syringae. Mol Gen Genet 244(4):341–351
Munson GP, Lam DL, Outten FW, O'Halloran TV (2000) Identification of a copper-responsive two-component system on the chromosome of Escherichia coli K-12. J Bacteriol 182(20):5864–5871
Ng SP, Davis B, Palombo EA, Bhave M (2009) A Tn5051-like mer-containing transposon identified in a heavy metal tolerant strain Achromobacter sp. AO22. BMC Res Notes 2:38
Ng SP, Palombo EA, Bhave M (2012) The heavy metal tolerant soil bacterium Achromobacter sp. AO22 contains a unique copper homeostasis locus and two mer operons. J Microbiol Biotechnol (in press)
Nybroe O, Brandt KK, Ibrahim YM, Tom-Petersen A, Holm PE (2008) Differential bioavailability of copper complexes to bioluminescent Pseudomonas fluorescens reporter strains. Environ Toxicol Chem 27(11):2246–2252
Praszkier J, Wilson IW, Pittard AJ (1992) Mutations affecting translational coupling between the rep genes of an IncB miniplasmid. J Bacteriol 174(7):2376–2383
Riether K, Dollard MA, Billard P (2001) Assessment of heavy metal bioavailability using Escherichia coli zntAp:lux and copAp:lux-based biosensors. Appl Microbiol Biotechnol 57(5):712–716
Rouch DA, Brown NL (1997) Copper-inducible transcriptional regulation at two promoters in the Escherichia coli copper resistance determinant pco. Microbiology 143(Pt 4):1191–1202
Schneider K, Beck CF (1986) Promoter-probe vectors for the analysis of divergently arranged promoters. Gene 42(1):37–48
Shetty RS, Deo SK, Shah P, Sun Y, Rosen BP, Daunert S (2003) Luminescence-based whole-cell-sensing systems for cadmium and lead using genetically engineered bacteria. Anal Bioanal Chem 376(1):11–17
Tauriainen S, Karp M, Chang W, Virta M (1998) Luminescent bacterial sensor for cadmium and lead. Biosens Bioelec 13(9):931–938
Trajanovska S, Britz ML, Bhave M (1997) Detection of heavy metal ion resistance genes in Gram-positive and Gram-negative bacteria isolated from a lead-contaminated site. Biodegradation 8(2):113–124
van der Meer JR, Belkin S (2010) Where microbiology meets micro engineering: design and applications of reporter bacteria. Nat Rev Micro 8(7):511–522
Verma N, Singh M (2005) Biosensors for heavy metals. Biometals 18(2):121–129
Yamamoto K, Ishihama A (2005) Transcriptional response of Escherichia coli to external copper. Mol Microbiol 56(1):215–227
Zhuang P, McBride MB, Xia H, Li N, Li Z (2009) Health risk from heavy metals via consumption of food crops in the vicinity of Dabaoshan mine, South China. Sci Total Environ 407(5):1551–1561
Acknowledgments
We thank Dr Dena Lyras (Department of Microbiology, Monash University, Melbourne) and Prof Jim Camakaris (Department of Genetics, The University of Melbourne, Melbourne) for kindly providing E. coli CB454 and pMU2385, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ng, S.P., Palombo, E.A. & Bhave, M. Identification of a copper-responsive promoter and development of a copper biosensor in the soil bacterium Achromobacter sp. AO22. World J Microbiol Biotechnol 28, 2221–2228 (2012). https://doi.org/10.1007/s11274-012-1029-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-012-1029-y