Abstract
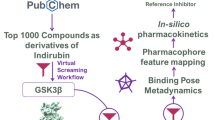
The striking structural resemblance between adenosine triphosphate (ATP) binding sites of glycogen synthase kinase-3 (GSK-3) and cyclin-dependent kinase-2 (CDK-2) raises numerous off-target selectivity problems in lead-identification processes that may jeopardize their progress into safe and effective drugs. The structural disparities between GSK-3 and CDK-2 in terms of inhibitors chemical space and binding site characteristics were investigated computationally by ligand-based (3D-similarity search) and structure-based (molecular docking) methods to reproduce the selectivity trend of indirubin derivatives. We attempted to assess distinctive key selectivity features of GSK-3 over CDK-2 with focus on indirubins and to provide a cascade virtual screening approach capable to identify suitable de novo GSK-3 selective scaffolds. Seven inhibitors with higher predicted interaction energies against GSK-3 compared to the highly active reference inhibitor were proposed. Concerted effects between 3D similarity search and docking afforded an exhaustive characterization of the binding site interactions. In spite of inherent challenges and limitations, the workflow developed hereby can be applied to other GSK-3 inhibitors, which display similar inhibitory profile against CDK-2, to rationally design potentially selective scaffolds.







Similar content being viewed by others
Abbreviations
- GSK-3:
-
Glycogen synthase kinase-3
- CDK-2:
-
Cyclin-dependent kinases-2
- MAPK:
-
Mitogen-activated protein kinase
- CLK:
-
CDC-like kinase
- ATP:
-
Adenosine triphosphate
- CDK-2 WII:
-
CDK-2 weak inhibitors and inactive dataset
- CDK-2 decoys:
-
DUD-E CDK-2 decoys dataset
- IXM:
-
Indirubin-3′-monoxime; (Z)-1H,1’H-[2,3′]bisindolylidene-3,2′-dione-3-oxime
- INR:
-
Indirubin-5-sulphonic acid; (2Z)-2′,3-dioxo-1,1′,2′,3-tetrahydro-2,3′-bisindole-5′-sulfonic acid
- TC:
-
Tanimoto combo
- ST:
-
Shape Tanimoto
- SC:
-
Scaled color
- CS:
-
Combo score
- CG4:
-
Chemgauss4
References
Woodgett JR (1990). EMBO J 9:2431–2438
Lee HC, Tsai JN, Liao PY, Tsai WY, Lin KY, Chuang CC, Sun CK, Chang WC, Tsai HJ (2007). BMC Dev Biol 7:93
Nikoulina SE, Ciaraldi TP, Mudaliar S, Carter L, Johnson K, Henry RR (2002). Diabetes 51:2190–2198
Cross DA, Alessi DR, Cohen P, Andjelkovich M, Hemmings BA (1995). Nature 378:785–789
Hart MJ, de los Santos R, Albert IN, Rubinfeld B, Polakis P (1998). Curr Biol 8:573–581
Ougolkov AV, Billadeau DD (2006). Future Oncol 2:91–100
Soos TJ, Meijer L, Nelson PJ (2006). Drug News Perspect 19:325–328
Ko HW, Kim EY, Chiu J, Vanselow JT, Kramer A, Edery I (2010). J Neurosci 30:12664–12675
Leclerc S, Garnier M, Hoessel R, Marko D, Bibb JA, Snyder GL, Greengard P, Biernat J, Wu YZ, Mandelkow EM, Eisenbrand G, Meijer L (2001). J Biol Chem 276:251–260
Bhat R, Xue Y, Berg S, Hellberg S, Ormö M, Nilsson Y, Radesäter A-C, Jerning E, Markgren P-O, Borgegård T, Nylöf M, Giménez-Cassina A, Hernández F, Lucas JJ, Díaz-Nido J, Avila J (2003). J Biol Chem 278:45937–45945
Rayasam GV, Tulasi VK, Sodhi R, Davis JA, Ray A (2009). J Pharmacol 156:885–898
Coghlan MP, Culbert AA, Cross DAE, Corcoran SL, Yates JW, Pearce NJ, Rausch OL, Murphy GJ, Carter PS, Roxbee Cox L, Mills D, Brown MJ, Haigh D, Ward RW, Smith DG, Murray KJ, Reith AD, Holder JC (2000). Chem Biol 7:793–803
Serenó L, Coma M, Rodríguez M, Sánchez-Ferrer P, Sánchez MB, Gich I, Agulló JM, Pérez M, Avila J, Guardia-Laguarta C, Clarimón J, Lleó A, Gómez-Isla T (2009). Neurobiol Dis 35:359–367
Takashima A (2009). J Pharmacol Sci 109:174–178
Mazanetz MP, Fischer PM (2007). Nat Rev Drug Discov 6:464–479
Spittaels K, van den Haute C, van Dorpe J, Geerts H, Mercken M, Bruynseels K, Lasrado R, Vandezande K, Laenen I, Boon T, van Lint J, Vandenheede J, Moechars D, Loos R, van Leuven F (2000). J Biol Chem 275:41340–41349
Caballero J, Zilocchi S, Tiznado W, Collina S, Rossi D (2011). Chem Biol Drug Des 78:631–641
Bertrand JA, Thieffine S, Vulpetti A, Cristiani C, Valsasina B, Knapp S, Kalisz HM, Flocco M (2003). J Mol Biol 333:393–407
Berg S, Bergh M, Hellberg S, Hogdin K, Lo-Alfredsson Y, Soderman P, Von Berg S, Weigelt T, Ormo M, Xue Y, Tucker J, Neelissen J, Jerning E, Nilsson Y, Bhat R (2012). J Med Chem 55:9107–9119
Georgievska B, Sandin J, Doherty J, Mörtberg A, Neelissen J, Andersson A, Gruber S, Nilsson Y, Schött P, Arvidsson PI, Hellberg S, Osswald G, Berg S, Fälting J, Bhat RV (2013). J Neurochem 125:446–456
Pradeep H, Rajanikant GK (2012). Mol Divers 16:553–562
Quesada-Romero L, Mena-Ulecia K, Tiznado W, Caballero J (2014). PLoS One 9:e102212
Meijer L, Greengard P, Knockaert M, Skaltsounis A (2007) Patent US 2007/0276025 A1
Polychronopoulos P, Magiatis P, Skaltsounis AL, Myrianthopoulos V, Mikros E, Tarricone A, Musacchio A, Roe SM, Pearl L, Leost M, Greengard P, Meijer L (2004). J Med Chem 47:935–946
Vougogiannopoulou K, Skaltsounis AL (2012). Planta Med 78:1515–1528
Vougogiannopoulou K, Ferandin Y, Bettayeb K, Myrianthopoulos V, Lozach O, Fan Y, Johnson CH, Magiatis P, Skaltsounis A-L, Mikros E, Meijer L (2008). J Med Chem 51:6421–6431
Choi S-J, Lee J-E, Jeong S-Y, Im I, Lee S-D, Lee E-L, Lee SK, Kwon SM, Ahn S-G, Yoon J-H, Han S-Y, Kim J-I, Kim Y-C (2010). J Med Chem 53:3696–3706
Suzuki K, Adachi R, Hirayama A, Watanabe H, Otani S, Watanabe Y, Kasahara T (2005). Br J Haematol 130:681–690
Ferandin Y, Bettayeb K, Kritsanida M, Lozach O, Polychronopoulos P, Magiatis P, Skaltsounis AL, Meijer L (2006). J Med Chem 49:4638–4649
Hoessel R, Leclerc S, Endicott JA, Nobel ME, Lawrie A, Tunnah P, Leost M, Damiens E, Marie D, Marko D, Niederberger E, Tang W, Eisenbrand G, Meijer L (1999) Nature. Cell Biol 1:60–67
Davies TG, Tunnah P, Meijer L, Marko D, Eisenbrand G, Endicott JA, Noble ME (2001). Structure 9:389–397
RCSB Protein Data Bank, RCSB PDB, https://www.rcsb.org/structure/1Q41 (accessed on January 2018)
RCSB Protein Data Bank, RCSB PDB, https://www.rcsb.org/structure/1E9H (accessed on January 2018)
Crisan L, Avram S, Pacureanu L (2017). Mol Divers 21:385–405
Aouidate A, Ghaleb A, Ghamali M, Chtita S, Ousaa A, M’b C, Sbai A, Bouachrine M, Lakhlifi T (2018). Struct Chem. https://doi.org/10.1007/s11224-018-1134-0
Crisan L, Pacureanu L, Bora A, Avram S, Kurunczi L, Simon Z (2013). Cent Eur J Chem 1:63–77
Katritzky AR, Pacureanu LM, Dobchev DA, Fara DC, Duchowicz PR, Karelson M (2006). Bioorg Med Chem 14:4987–5002
Pacureanu L, Crisan L, Bora A, Avram S, Kurunczi L (2012). Monatsh Chem 143:1559–1573
Crisan L, Pacureanu L, Bora A, Avram S, Kurunczi L (2013). Cent Eur J Chem 11:1644–1656
Crisan L, Pacureanu L, Avram S, Bora A, Avram S, Kurunczi L (2014) J Enz Inhib. Med Chem 29:599–610
Quesada-Romero L, Caballero J (2014). Mol Divers 18:149–159
Li X, Wang X, Tian Z, Zhao H, Liang D, Li W, Qiu Y, Lu S (2014). J Mol Model 20:2407
Kirchmair J, Distinto S, Schuster D, Spitzer G, Langer T, Wolber G (2008). Curr Med Chem 15:2040–2053
Kim S, Bolton EE, Bryant SH (2011). J Cheminform 3:26
Grant JA, Gallardo MA, Pickup B (1996). J Comp Chem 17:1653–1666
Gaulton A, Hersey A, Nowotka M, Bento AP, Chambers J, Mendez D, Mutowo P, Atkinson F, Bellis LJ, Cibrián-Uhalte E, Davies M, Dedman N, Karlsson A, Magariños MP, Overington JP, Papadatos G, Smit I, Leach AR (2017) The ChEMBL database in 2017. Nucleic Acids Res. 45(D1):D945–D954
Mysinger MM, Carchia M, Irwin JJ, Shoichet BK (2012). J Med Chem 55:6582–6594
Dean PM (1990) In: Maggiora GM, Johnson MA (eds) Concepts and applications of molecular similarity. Wiley&Sons, New York
Akabli T, Toufik H, Yasri A, Bih H, Lamchouri F (2018). Struct Chem. https://doi.org/10.1007/s11224-018-1141-1
OMEGA v.2.5.1.4 OpenEye Scientific Software Inc. Santa Fe NM, USA www.eyesopen.com
Hawkins PCD, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010). J Chem Inf Model 50:572–584
Hawkins PCD, Nicholls A (2012). J Chem Inf Model 52:2919–2936
Boström J, Greenwood JR, Gottfries J (2003). J Mol Graph Model 21:449–462
ROCS v.3.2.1.4 OpenEye Scientific Software Inc. Santa Fe NM, USA www.eyesopen.com
Hawkins PCD, Skillman AG, Nicholls A (2007). J Med Chem 50:74–82
Venhorst J, Nunez S, Terpstra JW, Kruse CG (2008). J Med Chem 51:3222–3229
Sheridan RP, McGaughey GB, Cornell WD (2008). J Comput Aided Mol Des 22:257–265
Rush TS, Grant JA, Mosyak L, Nicholls A (2005). J Med Chem 48:1489–1495
FRED v.3.2.0.2 OpenEye Scientific Software Inc. Santa Fe NM, USA www.eyesopen.com
McGann M (2011). J Chem Inf Model 51:578–596
Sotriffer C, Stahl M (2003) In: Abraham DJ (ed) Docking and scoring functions/virtual screening. Wiley & Sons, New York, p 1
Henrich S, Feierberg I, Wang T, Blomberg N, Wade RC (2010). Proteins 78:135–153
Meijer L, Skaltsounis AL, Magiatis P, Polychronopoulos P, Knockaert M, Leost M, Ryan XP, Vonica CA, Brivanlou A, Dajani R, Crovace C, Tarricone C, Musacchio A, Roe SM, Pearl L, Greengard P (2003). Chem Biol 10:1255–1266
Ribas J, Bettayeb K, Ferandin Y, Knockaert M, Garrofé-Ochoa X, Totzke F, Schächtele C, Mester J, Polychronopoulos P, Magiatis P, Skaltsounis AL, Boix J, Meijer L (2006). Oncogene 25:6304–6318
Olesen PH, Sørensen AR, Ursø B, Kurtzhals P, Bowler AN, Ehrbar U, Hansen BF (2003). J Med Chem 46:3333–3341
Kaidanovich-Beilin O, Woodgett JR (2011). Front Mol Neurosci 4:40
Meijer L, Flajolet M, Greengard P (2004). Trends Pharmacol Sci 25:471–480
Bain J, McLauchlan H, Elliot M, Cohen P (2003). Biochem J 371:199–204
FILTER v.2.5.1.4 OpenEye Scientific Software Inc. Santa Fe, NM USA www.eyesopen.com
Egan WJ, Merz KM, Baldwin JJ (2000). J Med Chem 43:3867–3877
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kipple KD (2002). J Med Chem 45:2615–2623
Martin YC (2005). J Med Chem 48:3164–3170
Schrödinger Release 2016–1: LigPrep v.3.1 (2016) Schrödinger, LLC, New York, NY
Schrödinger Release 2016–1: Maestro v.10.5 (2016) Schrödinger, LLC, New York, NY
Make Receptor v.3.2.0.2 OpenEye Scientific Software Inc., Santa Fe NM, USA www.eyesopen.com
Nicholls A, McGaughey GB, Sheridan RP, Good AC, Warren G, Mathieu M, Muchmore SW, Brown SP, Grant JA, Haigh JA, Nevins N, Jain AN, Kelley B (2010). J Med Chem 53:3862–3886
Yan X, Li J, Liu Z, Zheng M, Ge H, Xu J (2013). J Chem Inf Model 53:1967–1978
Fontaine F, Bolton E, Borodina Y, Bryant SH (2007). Chem Cent J 6:1–12
Muchmore SW, Debe DA, Metz JT, Brown SP, Martin YC, Hajduk PJ (2008). J Chem Inf Model 48:941–948
Bortolato A, Perruccio F, Moro S (2011) In: Miteva MA (ed) Successful applications of in silico Approaches for lead/drug discovery, Bentham Science Publishers
Sutherland JJ, Nandigam RK, Erickson JA, Vieth M (2007). J Chem Inf Model 47:2293–2302
Kelley BP, Brown SP, Warren GL, Muchmore SW (2015). J Chem Inf Model 55:1771–1780
Bemis GW, Murcko MA (1996). J Med Chem 39:2887–2893
Dassault Systèmes BIOVIA (2015) Discovery Studio Visualizer. v4.5.0, vol 15071. Dassault Systèmes, San Diego. www.3dsbiovia.com
Lu SY, Jiang YJ, Lv J, Zou JW, Wu TX (2011). J Comput Chem 32:1907–1918
Zhang B, Tan VBC, Lim KM, Tay TE (2007). J Chem Inf Model 47:1877–1885
Tirado-Rives J, Jorgensen WL (2006). J Med Chem 49:5880–5884
Chang CA, Chen W, Gilson MK (2007). Proc Natl Acad Sci USA 104:1534–1539
Duca JS, Madison VS, Voigt JH (2008). J Chem Inf Model 48:659–668
Sadowski J, Gasteiger J, Klebe G (1994). J Chem Inf Comput Sci 34:1000–1008
Boström J, Hogner A, Schmitt S (2006). J Med Chem 49:6716–6725
Kramer T, Schmidt B, Lo Monte F (2012). Int J Alzheimers Dis 2012:381029
Chohan TA, Qian H-Y, Pan Y-L, Chen J-Z (2015). Mol BioSyst 12:145–161
ChemSpider http://www.chemspider.com/ (accessed on July 2018)
SureChem http://www.surechem.com/ (accessed on July 2018)
Segraves NL, Robinson SJ, Garcia D, Said SA, Fu X, Schmitz FJ, Pietraszkiewicz H, Valeriote FA, Crews P (2004). J Nat Prod 67:783–792
Kim HM, Kim C-S, Lee J-H, Jang SJ, Hwang JJ, Ro S, Choi J (2013). PLoS ONE 8:e60383
Gustin JP, Karakas B, Weiss MB, Abukhdeir AM, Lauring J, Garay JP, Cosgrove D, Tamaki A, Konishi H, Konishi Y, Mohseni M, Wang G, Rosena DM, Denmeade SR, Higgins MJ, Vitolo MI, Bachman KE, Park BH (2009). Proc Natl Acad Sci U S A 106:2835–2840
Kunnimalaiyaan S, Schwartz VK, Alao Jackson I, Gamblin TC, Kunnimalaiyaan M (2018). BMC Cancer 18:560–567
Acknowledgments
We thank OpenEye Scientific Software, Chemaxon, for providing us an academic license, to Dr. Ramona Curpan, Institute of Chemistry Timisoara, for providing access to Schrodinger software acquired through the project PN–II–RU PD_502 funded by UEFISCDI–CNCSIS Romania, to BIOVIA Discovery Studio for the free license and SureChem for the free trial license.
Funding
This project was financially supported by the Institute of Chemistry Timisoara of the Romanian Academy, Project 1.2.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest. The authors alone are responsible for the content and writing of the paper.
Electronic supplementary material
ESM 1
(PDF 891 kb)
Rights and permissions
About this article
Cite this article
Pacureanu, L., Avram, S., Bora, A. et al. Portraying the selectivity of GSK-3 inhibitors towards CDK-2 by 3D similarity and molecular docking. Struct Chem 30, 911–923 (2019). https://doi.org/10.1007/s11224-018-1224-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1224-z