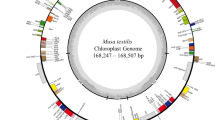
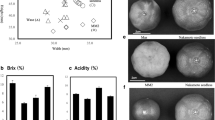
Abstract
Coffee is a popular beverage in the world and a valuable agricultural export commodity for several developing countries. Coffea arabica is one of the widely cultivated species and is believed to have originated from Ethiopia. Currently, a large number of C. arabica accessions are conserved ex situ in field gene banks in Ethiopia. However, there is no useful molecular barcoding to identify the conserved accessions, which presents a challenge to sustainable utilization of this economic species. To identify polymorphic chloroplast markers for the use in various studies, the complete chloroplast genome sequences of 24 C. arabica accessions were assembled and screened for variable regions in this study. The total length of the individual chloroplast genome sequence varied from 155,059 to 155,192 bp. Genome annotation revealed a total of 114 unique genes, consisting of 80 protein-coding, 30 tRNA, and four rRNA genes in all the 24 genomes. Twenty-four polymorphic regions were identified and developed. Most (71%) of the variable regions were found in non-coding sequence, and 54% of the variations detected are single base pairs. Three non-synonymous SNPs were found in accD, rpoA, and ycf1 genes. The markers developed were able to cluster the majority of the coffee accessions according to their geographical regions of collection. Five out of 24 newly developed markers were validated by assessing the diversity levels that were assessed in five original regions. The southwest region displayed the highest genetic diversity (HE = 0.224; I = 0.370) and might become a potential coffee genetic resource bank. Overall, the polymorphic markers developed and validated in this study could be a useful genomic resource for molecular breeding, identification, and biogeography studies of this commercially important crop.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.



Similar content being viewed by others
References
Aerts R, Berecha G, Gijbels P, Hundera K, Van Glabeke S, Vandepitte K, Muys B, Roldán-Ruiz I, Honnay O (2012) Genetic variation and risks of introgression in the wild Coffea arabica gene pool in south-western Ethiopian. Evol Appl:243–252. https://doi.org/10.1111/j.1752-4571.2012.00285.x
Aga E, Bekele E, Bryngelsson T (2005) Inter-simple sequence repeat (ISSR) variation in forest coffee trees (Coffea arabica L.) populations from Ethiopia. Genetica 124:213–214. https://doi.org/10.1007/s10709-005-1484-6
Alemayehu D (2017) Review on genetic diversity of coffee (Coffea arabica L.) in Ethiopia. Int J For Hortic 3(2):18–27. https://doi.org/10.20431/2454-9487.0302003
Alkimim ER, Caixeta ET, Sousa TV, da Silva FL, Sakiyama NS, Zambolim L (2018) High-throughput targeted genotyping using next-generation sequencing applied in Coffea canephora breeding. Euphytica 214:50–18. https://doi.org/10.1007/s10681-018-2126-2
Anthony F, Bertrand B, Quiros O, Wilches A, Lashermes P, Berthaud J, Charrier A (2001) Genetic diversity of wild coffee (Coffea arabica L.) using molecular markers. Euphytica 118(1):53–65. https://doi.org/10.1023/A:1004013815166
Benti T (2017) Progress in Arabica coffee breeding in Ethiopia : achievements, challenges and prospects. Int J Sci Basic Appl Res 33(2):15–25
Bi Y, Zhang M, Xue J, Dong R, Du YP, Zhang XH (2018) Chloroplast genomic resources for phylogeny and DNA barcoding : a case study on Fritillaria. Sci Rep 8:1–12. https://doi.org/10.1038/s41598-018-19591-9
CBOL Plant Working Group (2009) A DNA barcode for land plants. Proc Natl Acad Sci 106:12794–12797. https://doi.org/10.1073/pnas.0905845106
Chaney L, Mangelson R, Ramaraj T, Jellen EN, Maughan PJ (2016) The complete chloroplast genome sequences for four Amaranthus species. Appl Plant Sci 4(9):4–10. https://doi.org/10.3732/apps.1600063
Chen S, Chen S, Zhou Y, Chen Y, Gu J (2018) fastp : an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:884–890. https://doi.org/10.1093/bioinformatics/bty560
Chen S, Yao H, Han J, Liu C, Song J, Shi L, Zhu Y, Ma X, Gao T, Pang X, Luo K (2010) Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS One 5(1):1–8. https://doi.org/10.1371/journal.pone.0008613
Chung HY, Won SY, Kim YK, Kim JS (2019) Development of the chloroplast genome-based InDel markers in Niitaka (Pyrus pyrifolia) and its application. Plant Biotechnol Rep 13:51–61. https://doi.org/10.1007/s11816-018-00513-0
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2 : more models, new heuristics and parallel computing. Nat Methods 9(8):772. https://doi.org/10.1038/nmeth.2109
Davis AP, Govaerts R, Bridson DM, Stoffelen P (2006) An annotated taxonomic conspectus of the genus Coffea (Rubiaceae). Bot J Linn Soc 152:465–512
Davis AP, Tosh J, Ruch N, Fay MF (2011) Growing coffee : Psilanthus (Rubiaceae) subsumed on the basis of molecular and morphological data; implications for the size, morphology, distribution and evolutionary history of Coffea. Bot J Linn Soc 167:357–377
Dierckxsens N, Mardulyn P, Smits G (2017) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45(4):e18. https://doi.org/10.1093/nar/gkw955
Dong W, Liu J, Yu J, Wang L, Zhou S (2012) Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS One 7(4):1–9. https://doi.org/10.1371/journal.pone.0035071
Dong W, Xu C, Li C, Sun J, Zuo Y, Shi S, Cheng T, Guo J, Zhou S (2015) ycf1, the most promising plastid DNA barcode of land plants. Sci Rep 5:1–5. https://doi.org/10.1038/srep08348
Enan MR, Ahmed A (2014) DNA barcoding based on plastid mat K and RNA polymerase for assessing the genetic identity of date (Phoenix dactylifera L.) cultivars. Genet Mol Res 13(2):3527–3536
Fang Q, Wang L, Yu H, Huang Y, Jiang X, Deng X, Xu Q (2018) Development of species-specific InDel markers in citrus. Plant Mol Biol Rep 36(4):653–662
Fu C, Wu CS, Ye LJ, Mo ZQ, Liu J, Chang YW (2019) Prevalence of isomeric plastomes and effectiveness of plastome super-barcodes in yews (Taxus) worldwide. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-39161-x
Jain A, Roorkiwal M, Kale S, Garg V, Yadala R, Varshney RK (2019) InDel markers: an extended marker resource for molecular breeding in chickpea. PloS One 14(3):e0213999. https://doi.org/10.1371/journal.pone.0213999
Jarret RL (2008) DNA barcoding in a crop genebank : the Capsicum annuum species complex. Open Biol J 1:35–42
Jingade P, Huded AK, Kosaraju B, Mishra MK (2019) Diversity genotyping of Indian coffee (Coffea arabica L.) germplasm accessions by using SRAP markers. J Crop Improv 3(3):1–19. https://doi.org/10.1080/15427528.2019.1592050
Kaldate R, Rana M, Sharma V, Hirakawa H, Kumar R, Singh G, Chahota RK, Isobe SN, Sharma TR (2017) Development of genome-wide SSR markers in horsegram and their use for genetic diversity and cross-transferability analysis. Mol Breed 37(103):1–10. https://doi.org/10.1007/s11032-017-0701-1
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20(4):1160–1166. https://doi.org/10.1093/bib/bbx108
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kim TS, Lee JR, Raveendar S, Lee GA, Jeon YA, Lee HS, Ma KH, Lee SY, Chung JW (2016) Complete chloroplast genome sequence of Capsicum baccatum var. baccatum. Mol Breed 36(8):1–5. https://doi.org/10.1007/s11032-016-0532-5
Kode V, Mudd EA, Iamtham S, Day A (2005) The tobacco plastid accD gene is essential and is required for leaf development. Plant J 44(2):237–244. https://doi.org/10.1111/j.1365-313X.2005.02533.x
Krawczyk K, Nobis M, Myszczyński K, Klichowska E, Sawicki J (2018) Plastid super-barcodes as a tool for species discrimination in feather grasses (Poaceae: Stipa ). Sci Rep 8:1–10. https://doi.org/10.1038/s41598-018-20399-w
Lashermes P, Benoit B, Herv E (2009) Breeding coffee (Coffea arabica) for sustainable production. In: Jain SM, Priyadarshan PM (eds) Breeding plantation tree crops: tropical species. Springer Science+Business Media, LLC
Li B, Zheng Y (2018) Dynamic evolution and phylogenomic analysis of the chloroplast genome in Schisandraceae. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-27453-7
Li W, Liu Y, Yang Y, Xie X, Lu Y, Yang Z, Jin X, Dong W, Suo Z (2018) Interspecific chloroplast genome sequence diversity and genomic resources in Diospyros. BMC Plant Biol 18(210):1–11
Li W, Zhang C, Guo X, Liu Q, Wang K (2019) Complete chloroplast genome of Camellia japonica genome structures, comparative and phylogenetic analysis. PloS One 14(5). https://doi.org/10.1371/journal.pone.0216645
Li X, Yang Y, Henry RJ, Rossetto M, Wang Y, Chen S (2015) Plant DNA barcoding: from gene to genome. Biol Rev. 90:157–166. https://doi.org/10.1111/brv.12104
Lohse M, Drechsel O, Kahlau S, Bock R (2013) Organellar genome DRAW- a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:575–581. https://doi.org/10.1093/nar/gkt289
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res:1–4. https://doi.org/10.1093/nar/gkw413
Maurin O, Davis AP, Chester M, Mvung EF, Jaufeerally-Fakim Y, Fay MF (2007) Towards a phylogeny for Coffea (Rubiaceae): identifying well-supported lineages based on nuclear and plastid DNA Sequences. Ann Bot 100:1565–1583. https://doi.org/10.1093/aob/mcm257
Mehari B, Redi-Abshiro M, Chandravanshi BS, Combrinc S, McCrindle R, Atlabachew M (2019) GC-MS profiling of fatty acids in green coffee (Coffea arabica L.) beans and chemometric modelling for tracing geographical origins from Ethiopia. J Sci Food Agric. https://doi.org/10.1002/jsfa.9603
Mosa KA, Gairola S, Jamdade R, El-Keblawy A, Al Shaer KI, Al Harthi EK, Shabana HA, Mahmoud T (2019) The promise of molecular and genomic techniques for biodiversity research and DNA barcoding of the Arabian peninsula flora. Front Plant Sci 9:1–19. https://doi.org/10.3389/fpls.2018.01929
Ndjiondjop MN, Semagn K, Zhang J, Gouda AC, Kpeki SB, Goungoulou A, Wambugu P, Dramé KN, Bimpong IK, Zhao (2018) Development of species diagnostic SNP markers for quality control genotyping in four rice (Oryza L.) species. Mol Breed 38(131). https://doi.org/10.1007/s11032-018-0885-z
Nock CJ, Waters DL, Edwards MA, Bowen SG, Rice N, Cordeiro GM, Henry RJ (2011) Chloroplast genome sequences from total DNA for plant identification. Plant Biotechnol J 9:328–333. https://doi.org/10.1111/j.1467-7652.2010.00558.x
Onyia CO, Jideofor OP, Ojiego BO, Solomon BO, Ogundipe OO, Ogbadu LJ (2014) Current developments in the application of DNA barcoding to solving biodiversity conservation problems in developing countries. Glob J Biol Agric Health Sci 3(3):285–288
Park I, Yang S, Kim WJ, Noh P, Lee HO, Moon BC (2018) The complete chloroplast genomes of six Ipomoea species and indel marker development for the discrimination of authentic pharbitidis semen (seeds of I. nil or I. purpurea). Front Plant Sci 9:1–14. https://doi.org/10.3389/fpls.2018.00965
Peakall R, Smouse P (2012) GenAlEx 6.5:genetic analysis in excel. Population genetic software for teaching and research – an update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pečnikar ŽF, Buzan EV (2014) 20 years since the introduction of DNA barcoding: from theory to application. J Appl Genet 55:43–52. https://doi.org/10.1007/s13353-013-0180-y
Phumichai C, Phumichai T, Wongkaew A (2015) Novel chloroplast microsatellite (cpSSR) markers for genetic diversity assessment of cultivated and wild Hevea rubber. Plant Mol Biol Rep 33(5):1486–1498
Rambaut A (2017) Figtree-version 1.4. 3, a graphical viewer of phylogenetic trees.
Samson N, Bausher MG, Lee SB, Jansen RK, Daniell H (2007) The complete nucleotide sequence of the coffee (Coffea arabica L.) chloroplast genome: organization and implications for biotechnology and phylogenetic relationships amongst angiosperms. Plant Biotechnol J 5:339–353. https://doi.org/10.1111/j.1467-7652.2007.00245.x
Silvestrini M, Junqueira MG, Favarin AC, Guerreiro-Filho O, Maluf MP, Silvarolla MB, Colombo CA (2007) Genetic diversity and structure of Ethiopian, Yemen and Brazilian Coffea arabica L. accessions using microsatellites markers. Genet Resour Crop Evol 54:1367–1379. https://doi.org/10.1007/s10722-006-9122-4
Steiger D, Nagai C, Moore P, Morden C, Osgood R, Ming R (2002) AFLP analysis of genetic diversity within and among Coffea arabica cultivars. Theor Appl Genet 105(2–3):209–215. https://doi.org/10.1007/s00122-002-0939-8
Sualeh A, Mekonnen N, Degefa M (2017) Hybrid coffee (Coffea arabica L.) genotypes quality evaluation under different environment of southern Ethiopia. Greener J Agric Sci 4(6):245–251. https://doi.org/10.15580/GJAS.2014.6.0523014244
Tesfaye K, Borsch T, Govers K, Bekele E (2007) Characterization of Coffea chloroplast microsatellites and evidence for the recent divergence of C. arabica and C. eugenioides chloroplast genomes. Genome 50:1112–1129. https://doi.org/10.1139/G07-088
Tesfaye K, Govers K, Bekele E, Borsch T (2014) ISSR fingerprinting of Coffea arabica throughout Ethiopia reveals high variability in wild populations and distinguishes them from landraces. Plant Syst Evol 300(5):881–897. https://doi.org/10.1007/s00606-013-0927-2
Tran HT, Lee LS, Furtado A, Smyth H, Henry RJ (2016) Advances in genomics for the improvement of quality in coffee. J Sci Food Agric 96:3300–3312. https://doi.org/10.1002/jsfa.7692
Trifinopoulos J, Nguyen L, Haeseler AV, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw256
Vaughan DA, Balazs E, Heslop-Harrison JS (2007) From crop domestication to super-domestication. Ann Bot 100:893–901. https://doi.org/10.1093/aob/mcm224
Wang M, Cui L, Feng K, Deng P, Du X, Wan F, Weining S, Nie X (2015) Comparative analysis of Asteraceae chloroplast genomes: structural organization, RNA editing and evolution. Plant Mol Biol Rep 33(5):1526–1538
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20(17):3252–3255. https://doi.org/10.1093/bioinformatics/bth352
Yan L, Ogutu C, Huang L, Wang X, Zhou H, Lv Y, Long Y, Dong Y, Han Y (2019) Genetic diversity and population structure of coffee germplasm collections in China revealed by ISSR markers. Plant Mol Biol Rep 37(3):204–213
Acknowledgments
The authors would like to thank the Ethiopian Biodiversity Institute for providing the coffee leaf samples used in this study. We gratefully acknowledge Jara Negash for his assistance during sample collection and Andrew W. Gichira for his help with the data analysis.
Funding
We thank Sino-Africa Joint Research Center, CAS (SAJC201614), for providing funding for the study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
• Comparative analysis of the newly sequenced 24 chloroplast genome sequences of ex situ conserved Coffea arabica accessions have been conducted.
• Three non-synonymous single-nucleotide polymorphisms were found in accD, rpoA, and ycf1 genes.
• The markers developed could be a useful genomic resource for molecular barcoding, molecular breeding, and biogeography studies.
Electronic Supplementary Material
Table S1
Informations of Coffea arabica accessions used for marker validation and polymorphism levels. (DOCX 17 kb)
Table S2
List of genes encoded by chloroplast genomes of Coffea arabica. (DOCX 14 kb)
Table S3
The twenty-four variable regions identified in the newly sequenced chloroplast genome sequences of Coffea arabica accessions. (DOCX 15 kb)
Table S4
Polymorphism detected across the 24 chloroplast genome sequences of Coffea arabica accessions. (DOCX 22 kb)
Rights and permissions
About this article
Cite this article
Mekbib, Y., Saina, J.K., Tesfaye, K. et al. Chloroplast Genome Sequence Variations and Development of Polymorphic Markers in Coffea arabica. Plant Mol Biol Rep 38, 491–502 (2020). https://doi.org/10.1007/s11105-020-01212-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-020-01212-3