Abstract
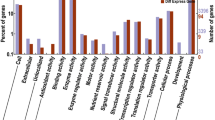
In order to unravel the molecular mechanism of maize ear development, a microarray containing ~56,000 probes was used to monitor the gene expression profiles of ears at four developmental stages. The results showed that 2,794 genes, accounting for 5.0% of the total probes, changed significantly during ear development. Among the 2,794 genes, 1,844 genes differentially expressed during the spikelet differentiation phase, 836 genes during the floret primordium differentiation phase and 645 genes during the floret organ differentiation phase. Hierarchical clustering revealed that the differentially expressed genes had 9 major expression patterns. Based on Mips Functional Catalogue, 684 differentially expressed genes were grouped into at least one functional category, including metabolism (30.4%), protein related function (29.2%), biogenesis of cellular components (15.4%) and transcription (13.7%). The analysis revealed that the auxin signaling pathway play an important role in ear development. Moreover, regulation of some transcription factors may play a key role during ear development. RT-PCR and in situ hybridization for some selected genes validated our microarray data and supplied additional information on ear developmental processes.





Similar content being viewed by others
References
Ambrose BA, Lerner DR, Ciceri P, Padilla CM, Yanofsky MF, Schmidt RJ (2000) Molecular and genetic analyses of the silky1 gene reveal conservation in floral organ specification between eudicots and monocots. Mol Cell 5:569–579. doi:10.1016/S1097-2765(00)80450-5
Becker A, Kaufmann K, Freialdenhoven A, Vincent C, Li MA, Saedler H, Theissen G (2002) A novel MADS-box gene subfamily with a sister-group relationship to class B floral homeotic genes. Mol Genet Genomics 266:942–950. doi:10.1007/s00438-001-0615-8
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300
Bommert P, Lunde C, Nardmann J, Vollbrecht E, Running M, Jackson D, Hake S, Werr W (2005) Thick tassel dwarf1 encodes a putative maize ortholog of the Arabidopsis CLAVATA1 leucine-rich repeat receptor-like kinase. Development 132:1235–1245. doi:10.1242/dev.01671
Cacharron J, Saedler H, Theissen G (1999) Expression of MADS box genes ZMM8 and ZMM14 during inflorescence development of Zea mays discriminates between the upper and the lower floret of each spikelet. Dev Genes Evol 209:411–420. doi:10.1007/s004270050271
Cheng P, Pareddy DR (1994) Morphology and development of the tassel and ear. In: Freeling M, Walbot V (eds) The maize handbook. Springer-Verlag, New York, USA, pp 37–43
Chuck G, Meeley RB, Hake S (1998) The control of maize spikelet meristem fate by the APETALA2-like gene indeterminate spikelet1. Genes Dev 12:1145–1154. doi:10.1101/gad.12.8.1145
Chuck G, Muszynski M, Kellogg E, Hake S, Schmidt RJ (2002) The control of spikelet meristem identity by the branched silkless1 gene in maize. Science 298:1238–1241. doi:10.1126/science.1076920
Dellaporta SL, Calderon-Urrea A (1994) The sex determination process in maize. Science 266:1501–1505. doi:10.1126/science.7985019
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868. doi:10.1073/pnas.95.25.14863
Firnhaber C, Puhler A, Kuster H (2005) EST sequencing and time course microarray hybridizations identify more than 700 Medicago truncatula genes with developmental expression regulation in flowers and pods. Planta 222:269–283. doi:10.1007/s00425-005-1543-3
Furutani I, Sukegawa S, Kyozuka J (2006) Genome-wide analysis of spatial and temporal gene expression in rice panicle development. Plant J 46:503–511. doi:10.1111/j.1365-313X.2006.02703.x
Gallavotti A, Zhao Q, Kyozuka J, Meeley RB, Ritter MK, Doebley JF, Pe ME, Schmidt RJ (2004) The role of barren stalk1 in the architecture of maize. Nature 432:630–635. doi:10.1038/nature03148
Gomi K, Matsuoka M (2003) Gibberellin signalling pathway. Curr Opin Plant Biol 6:489–493. doi:10.1016/S1369-5266(03)00079-7
Guo QF, Wang QC, Wang LM (2004) China maize culture. Science and Technology Publisher, Shanghai
Hennig L, Gruissem W, Grossniklaus U, Kohler C (2004) Transcriptional programs of early reproductive stages in Arabidopsis. Plant Physiol 135:1765–1775. doi:10.1104/pp.104.043182
Hu W, Wang Y, Bowers C, Ma H (2003) Isolation, sequence analysis, and expression studies of florally expressed cDNAs in Arabidopsis. Plant Mol Biol 53:545–563. doi:10.1023/B:PLAN.0000019063.18097.62
Kerr MK, Churchill GA (2001) Experimental design for gene expression microarrays. Biostatistics 2:183–201. doi:10.1093/biostatistics/2.2.183
Laitinen RA, Immanen J, Auvinen P, Rudd S, Alatalo E, Paulin L, Ainasoja M, Kotilainen M, Koskela S, Teeri TH, Elomaa P (2005) Analysis of the floral transcriptome uncovers new regulators of organ determination and gene families related to flower organ differentiation in Gerbera hybrida (Asteraceae). Genome Res 15:475–486. doi:10.1101/gr.3043705
Lan L, Chen W, Lai Y, Suo J, Kong Z, Li C, Lu Y, Zhang Y, Zhao X, Zhang X, Zhang Y, Han B, Cheng J, Xue Y (2004) Monitoring of gene expression profiles and isolation of candidate genes involved in pollination and fertilization in rice (Oryza sativa L.) with a 10 K cDNA microarray. Plant Mol Biol 54:471–487. doi:10.1023/B:PLAN.0000038254.58491.c7
Liang X, Zhang Z (1995) Studies on the relation between panicle initiation and leaf age in maize. J S China Agric Univ 16:83–87
Lu XC, Gong HQ, Huang ML, Bai SL, He YB, Mao X, Geng Z, Li SG, Wei L, Yuwen JS, Xu ZH, Bai SN (2006) Molecular analysis of early rice stamen development using organ-specific gene expression profiling. Plant Mol Biol 61:845–861. doi:10.1007/s11103-006-0054-3
Mao X, Cai T, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793. doi:10.1093/bioinformatics/bti430
McSteen P, Hake S (2001) Barren inflorescence2 regulates axillary meristem development in the maize inflorescence. Development 128:2881–2891
Nemhauser JL, Feldman LJ, Zambryski PC (2000) Auxin and ETTIN in Arabidopsis gynoecium morphogenesis. Development 127:3877–3888
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247. doi:10.1093/dnares/10.6.239
Quint M, Gray WM (2006) Auxin signaling. Curr Opin Plant Biol 9:448–453. doi:10.1016/j.pbi.2006.07.006
Ruan Y, Gilmore J, Conner T (1998) Towards Arabidopsis genome analysis: monitoring expression profiles of 1400 genes using cDNA microarrays. Plant J 15:821–833. doi:10.1046/j.1365-313X.1998.00254.x
Sablowski RW, Meyerowitz EM (1998) A homolog of no apical meristem is an immediate target of the floral homeotic genes APETALA3/PISTILLATA. Cell 92:93–103. doi:10.1016/S0092-8674(00)80902-2
Schmidt RJ, Veit B, Mandel MA, Mena M, Hake S, Yanofsky MF (1993) Identification and molecular characterization of ZAG1, the maize homolog of the Arabidopsis floral homeotic gene AGAMOUS. Plant Cell 5:729–737
Schnable PS, Hochholdinger F, Nakazono M (2004) Global expression profiling applied to plant development. Curr Opin Plant Biol 7:50–56. doi:10.1016/j.pbi.2003.11.001
Shi YH, Zhu SW, Mao XZ, Feng JX, Qin YM, Zhang L, Cheng J, Wei LP, Wang ZY, Zhu YX (2006) Transcriptome profiling, molecular biological, and physiological studies reveal a major role for ethylene in cotton fiber cell elongation. Plant Cell 18:651–664. doi:10.1105/tpc.105.040303
Smith DL, Fedoroff NV (1995) LRP1, a gene expressed in lateral and adventitious root primordia of Arabidopsis. Plant Cell 7:735–745
Smyth GK (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170. doi:10.1016/S0092-8674(00)81093-4
Taguchi-Shiobara F, Yuan Z, Hake S, Jackson D (2001) The fasciated ear2 gene encodes a leucine-rich repeat receptor-like protein that regulates shoot meristem proliferation in maize. Genes Dev 15:2755–2766. doi:10.1101/gad.208501
Tan QK, Irish VF (2006) The Arabidopsis zinc finger-homeodomain genes encode proteins with unique biochemical properties that are coordinately expressed during floral development. Plant Physiol 140:1095–1108. doi:10.1104/pp.105.070565
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signalling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7:847–859. doi:10.1038/nrm2020
Theissen G, Strater T, Fischer A, Saedler H (1995) Structural characterization, chromosomal localization and phylogenetic evaluation of two pairs of AGAMOUS-like MADS-box genes from maize. Gene 156:155–166. doi:10.1016/0378-1119(95)00020-7
Ullah H, Chen JG, Temple B, Boyes DC, Alonso JM, Davis KR, Ecker JR, Jones AM (2003) The beta-subunit of the Arabidopsis G protein negatively regulates auxin-induced cell division and affects multiple developmental processes. Plant Cell 15:393–409. doi:10.1105/tpc.006148
Vollbrecht E, Springer PS, Goh L, Buckler ES, Martienssen R (2005) Architecture of floral branch systems in maize and related grasses. Nature 436:1119–1126. doi:10.1038/nature03892
Wang Z, Liang Y, Li C, Xu Y, Lan L, Zhao D, Chen C, Xu Z, Xue Y, Chong K (2005) Microarray analysis of gene expression involved in anther development in rice (Oryza sativa L.). Plant Mol Biol 58:721–737. doi:10.1007/s11103-005-8267-4
Wellmer F, Riechmann JL, ves-Ferreira M, Meyerowitz EM (2004) Genome-wide analysis of spatial gene expression in Arabidopsis flowers. Plant Cell 16:1314–1326. doi:10.1105/tpc.021741
Whipple CJ, Ciceri P, Padilla CM, Ambrose BA, Bandong SL, Schmidt RJ (2004) Conservation of B-class floral homeotic gene function between maize and Arabidopsis. Development 131:6083–6091. doi:10.1242/dev.01523
Xu Y, Chong K, Xu Z, Tan K (2001) Expression patterns of a vernalization-related genes responding to jasmonate. Acta Bot Sin 43:871–873
Zhang X, Feng B, Zhang Q, Zhang D, Altman N, Ma H (2005) Genome-wide expression profiling and identification of gene activities during early flower development in Arabidopsis. Plant Mol Biol 58:401–419. doi:10.1007/s11103-005-5434-6
Zhu Y, Wang M, Jia Z, Lian Y, Jin Y, Wang G (2007) An effective method for extracting total RNA from young ear of maize. Chin Bull Bot 24:624–628
Acknowledgments
The authors thank Dr. Jingrui Dai and Qifeng Xu (China Agricultural University) for offering maize seeds, Dr. Zhen Su (China Agricultural University) and Wenying Xu (Institute of Genetics and Development) for helpful advice in microarray design and hybridization. We also thank Dr. Patrick S Schnable (Iowa State University, USA) and Dr. Xinmin Li (University of Chicago, USA) for their pre-review and helpful suggestions. This work was supported by the National Basic Research Program of China (2006CB101700).
Author information
Authors and Affiliations
Corresponding author
Additional information
Yun Zhu and Junjie Fu authors contributed equally to this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhu, Y., Fu, J., Zhang, J. et al. Genome-wide analysis of gene expression profiles during ear development of maize. Plant Mol Biol 70, 63–77 (2009). https://doi.org/10.1007/s11103-009-9457-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-009-9457-2