Abstract
Molecular marker-assisted backcrossing (MABC) is widely recommended for transferring favorable alleles from a donor to an elite variety. The question remains whether MABC is an effective approach to developing a competitive commercial variety. Here, we illustrate the transfer of a thermostable β-amylase allele Sd3 from wild barley into a commercial barley variety Gairdner. The elite lines were chosen for the Regional Crop Variety Test that followed a standard conventional breeding process. The results demonstrated that the Sd3 allele not only increased enzyme thermostability but dramatically enhanced diastatic power, an important malting quality trait. The BC1F1 individuals had a fundamental impact on the comprehensive agronomic and quality traits of the final progenies, demonstrating the importance of screening at the early stage of backcrossing in MABC. There was sufficient genetic variation in the BC3F3 families to select other malting quality and agronomic traits. Ten individual breeding lines with improved β-amylase thermostability also had improved yields and grain plumpness. Three elite lines with improved malting quality and agronomic traits were selected to provide a parental line to incorporate the wild barley allele for breeding a commercial variety. A new strategy should be considered that uses marker-assisted selection and backcrossing to transfer a favorable allele from a wild parent.




Similar content being viewed by others
References
Abeledo LG, Calderini DF, Slafer GA (2003) Genetic improvement of barley yield potential and its physiological determinants in Argentina (1944–1998). Euphytica 130(3):325–334. https://doi.org/10.1023/A:1023089421426
Ames N, Dreiseitl A, Steffenson B, Muehlbauer G (2015) Mining wild barley for powdery mildew resistance. Plant Pathol 64(6):1396–1406. https://doi.org/10.1111/ppa.12384
Analytica EBC (1998) European Brewery Convention Analytica-EBC, Method 4.5. Congress Method. Fachverlag Hans Carl, Nurnberg, Germany
Bankole F, Menkir A, Olaoye G, Crossa J, Hearne S, Unachukwu N, Gedil M (2017) Genetic gains in yield and yield related traits under drought stress and favorable environments in a maize population improved using marker assisted recurrent selection. Front Plant Sci 8(808). https://doi.org/10.3389/fpls.2017.00808
Chiapparino E, Donini P, Reeves J, Tuberosa R, O’Sullivan DM (2006) Distribution of β-amylase I haplotypes among European cultivated barleys. Mol Breeding 18(4):341–354. https://doi.org/10.1007/s11032-006-9035-0
Dai F, Nevo E, Wu D, Comadran J, Zhou M, Qiu L, Chen Z, Beiles A, Chen G, Zhang G (2012) Tibet is one of the centers of domestication of cultivated barley. Proc Natl Acad Sci U S A 109(42):16969–16973. https://doi.org/10.1073/pnas.1215265109
Das G, Rao G (2015) Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front Plant Sci 6(698). https://doi.org/10.3389/fpls.2015.00698
Dracatos PM, Singh D, Bansal U, Park RF (2015) Identification of new sources of adult plant resistance to Puccinia hordei in international barley (Hordeum vulgare L.) germplasm. Eur J Plant Pathol 141(3):463–476. https://doi.org/10.1007/s10658-014-0556-9
Eglinton J, Evans D, Brown A, Langridge P, McDonald G, Jefferies S, Barr A (1999) The use of wild barley (Hordeum vulgare ssp. spontaneum) in breeding for quality and adaptation. In: 9th Australian Barley Technical Symposium, Melbourne. http://www.regional.org.au/au/abts/1999/eglinton.htm#TopOfPage
Eglinton J, Langridge P, Evans D (1998) Thermostability variation in alleles of barley beta-amylase. J Cereal Sci 28(3):301–309. https://doi.org/10.1016/S0733-5210(98)90010-8
Ellis R, Forster B, Robinson D, Handley L, Gordon D, Russell J et al (2000) Wild barley: a source of genes for crop improvement in the 21st century? J Exp Bot 51(342):9–17. https://doi.org/10.1093/jexbot/51.342.9
Erkkilä MJ, Leah R, Ahokas H, Cameron-Mills V (1998) Allele-dependent barley grain β-amylase activity. Plant Physiol 117(2):679–685. https://doi.org/10.1104/pp.117.2.679
Evans D, Li C, Eglinton J (2009) The properties and genetics of barley malt starch degrading enzymes. In: Genetics and improvement of barley malt quality. Springer, berlin, Heidelberg, pp 143–189. https://doi.org/10.1007/978-3-642-01279-2_6
Evans DE, Collins H, Eglinton J, Wilhelmson A (2005) Assessing the impact of the level of diastatic power enzymes and their thermostability on the hydrolysis of starch during wort production to predict malt fermentability. J Am Soc Brew Chem 63(4):185–198. https://doi.org/10.1094/asbcj-63-0185
Genger R, Williams K, Raman H, Read B, Wallwork H, Burdon J, Brown A (2003) Leaf scald resistance genes in Hordeum vulgare and Hordeum vulgare ssp. spontaneum: parallels between cultivated and wild barley. Crop Pasture Sci 54(12):1335–1342. https://doi.org/10.1071/AR02230
Gong X, Westcott S, Zhang X-Q, Yan G, Lance R, Zhang G, Sun D, Li C (2013) Discovery of novel Bmy1 alleles increasing β-amylase activity in Chinese landraces and Tibetan wild barley for improvement of malting quality via MAS. PLoS One 8(9):e72875. https://doi.org/10.1371/journal.pone.0072875
He J, Zhao X, Laroche A, Lu Z-X, Liu H, Li Z (2014) Genotyping-by-sequencing (GBS), an ultimate marker-assisted selection (MAS) tool to accelerate plant breeding. Front Plant Sci 5:484. https://doi.org/10.3389/fpls.2017.01809
Howell P, Leigh F, Bates R, Gosman N, Trafford K, Powell, Smith AM, Greenland A (2014) Rapid marker-assisted development of advanced recombinant lines from barley starch mutants. Mol Breeding 33 (1):243–248. https://doi.org/10.1007/s11032-013-9930-0
Jakob SS, Rödder D, Engler JO, Shaaf S, Özkan H, Blattner FR, Kilian B (2014) Evolutionary history of wild barley (Hordeum vulgare subsp. spontaneum) analyzed using multilocus sequence data and paleodistribution modeling. Genome Biol Evol 6 (3):685–702. https://doi.org/10.1093/gbe/evu047
Karsai I, Szűcs P, Mészáros K, Filichkina T, Hayes P, Skinner J, Láng L, Bedő Z (2005) The Vrn-H2 locus is a major determinant of flowering time in a facultative× winter growth habit barley (Hordeum vulgare L.) mapping population. Theor Appl Genet 110(8):1458–1466. https://doi.org/10.1007/s00122-005-1979-7
Li C, Cakir M, Lance R (2009) Genetic improvement of malting quality through conventional breeding and marker-assisted selection. In: Genetics and improvement of barley malt quality. Springer, berlin, Heidelberg, pp 260–292. https://doi.org/10.1007/978-3-642-01279-2_9
Liu X-Q, Rong J-Y, Liu X-Y (2008) Best linear unbiased prediction for linear combinations in general mixed linear models. J Multivariate Anal 99(8):1503–1517. https://doi.org/10.1016/j.jmva.2008.01.004
Malysheva-Otto L, Röder M (2006) Haplotype diversity in the endosperm specific β-amylase gene Bmy1 of cultivated barley (Hordeum vulgare L.). Mol Breeding 18(2):143–156. https://doi.org/10.1007/s11032-006-9023-4
Manes Y, Gomez H, Puhl L, Reynolds M, Braun H, Trethowan R (2012) Genetic yield gains of the CIMMYT international semi-arid wheat yield trials from 1994 to 2010. Crop Sci 52(4):1543–1552. https://doi.org/10.2135/cropsci2011.10.0574
Mondal S, Rutkoski JE, Velu G, Singh PK, Crespo-Herrera LA, Guzmán C, Bhavani S, Lan C, He X, Singh RP (2016) Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front Plant Sci 7(991). https://doi.org/10.3389/fpls.2016.00991
Nevo E (2015) Evolution of wild barley at “evolution canyon”: adaptation, speciation, pre-agricultural collection, and barley improvement. Isr J Plant Sci 62(1–2):22–32. https://doi.org/10.1080/07929978.2014.940783
Nevo E, Chen G (2010) Drought and salt tolerances in wild relatives for wheat and barley improvement. Plant Cell Environ 33(4):670–685. https://doi.org/10.1111/j.1365-3040.2009.02107.x
Paris M, Jones MG, Eglinton JK (2002) Genotyping single nucleotide polymorphisms for selection of barley β-amylase alleles. Plant Mol Biol Report 20(2):149–159. https://doi.org/10.1007/BF02799430
Polakova K, Laurie D, Vaculova K, Ovesna J (2003) Characterization of β-amylase alleles in 79 barley varieties with pyrosequencing. Plant Mol Biol Report 21(4):439–447. https://doi.org/10.1007/BF02772593
Russell J, Mascher M, Dawson IK, Kyriakidis S, Calixto C, Freund F, Bayer M, Milne I, Marshall-Griffiths T, Heinen S (2016) Exome sequencing of geographically diverse barley landraces and wild relatives gives insights into environmental adaptation. Nat Genet 48(9):1024–1030. https://doi.org/10.1038/ng.3612
Schmalenbach I, Pillen K (2009) Detection and verification of malting quality QTLs using wild barley introgression lines. Theor Appl Genet 118(8):1411–1427. https://doi.org/10.1007/s00122-009-0991-8
Shavrukov Y, Gupta NK, Miyazaki J, Baho MN, Chalmers KJ, Tester M, Langridge P, Collins NC (2010) HvNax3—a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct Integr Genomics 10(2):277–291. https://doi.org/10.1007/s10142-009-0153-8
Thomas W, Baird E, Fuller J, Lawrence P, Young G, Russell J, Ramsay L, Waugh R, Powell W (1998) Identification of a QTL decreasing yield in barley linked to Mlo powdery mildew resistance. Mol Breeding 4(5):381–393. https://doi.org/10.1023/A:1009646115967
Vishwakarma MK, Arun B, Mishra V, Yadav P, Kumar H, Joshi AK (2016) Marker-assisted improvement of grain protein content and grain weight in Indian bread wheat. Euphytica 208(2):313–321. https://doi.org/10.1007/s10681-015-1598-6
von Korff M, Wang H, Léon J, Pillen K (2008) AB-QTL analysis in spring barley: III. Identification of exotic alleles for the improvement of malting quality in spring barley (H. vulgare ssp. spontaneum). Mol Breeding 21(1):81–93. https://doi.org/10.1007/s11032-007-9110-1
Wang J, Yang J, Zhang Q, Zhu J, Jia Q, Hua W, Shang Y, Li C, Zhou M (2015a) Mapping a major QTL for malt extract of barley from a cross between TX9425 × Naso Nijo. Theor Appl Genet 128(5):943–952. https://doi.org/10.1007/s00122-015-2481-5
Wang X, Zhang X, Cai S, Ye L, Zhou M, Chen Z, Zhang G, Dai F (2015b) Genetic diversity and QTL mapping of thermostability of limit dextrinase in barley. J Agric Food Chem 63(14):3778–3783. https://doi.org/10.1021/acs.jafc.5b00190
Yu Y, Huang Y, Zhang W (2012) Changes in rice yields in China since 1980 associated with cultivar improvement, climate and crop management. Field Crops Res 136:65–75. https://doi.org/10.1016/j.fcr.2012.07.021
Zhang WS, Li X, Liu JB (2007) Genetic variation of Bmy1 alleles in barley (Hordeum vulgare L.) investigated by CAPS analysis. Theor Appl Genet 114(6):1039–1050. https://doi.org/10.1007/s00122-006-0497-6
Zhou G, Panozzo J, Zhang X-Q, Cakir M, Harasymow S, Li C (2016) QTL mapping reveals genetic architectures of malting quality between Australian and Canadian malting barley (Hordeum vulgare L.). Mol Breeding 36(6):1–12. https://doi.org/10.1007/s11032-016-0492-9
Acknowledgements
The project funds were provided by the Australian Grain Research and Development Corporation (DAW00233) and National Natural Science Foundation of China (31471496, 31501309).
Author information
Authors and Affiliations
Contributions
CL and WZ designed the experiments; XY, XQZ, SH, and SW conducted the research; XY, WZ, and CL wrote and finalized the manuscript.
Corresponding authors
Ethics declarations
This research does not involve human participants or animals.
Conflict of interests
The authors declare that they have no competing interests.
Electronic supplementary material
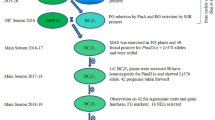
Figure. S1
Scheme of marker-assisted selection on thermostable β-amylase through backcrossing the introgressing Sd3 allele from wild barley to a commercial variety. (JPG 29 kb)
Rights and permissions
About this article
Cite this article
Xu, Y., Zhang, XQ., Harasymow, S. et al. Molecular marker-assisted backcrossing breeding: an example to transfer a thermostable β-amylase gene from wild barley. Mol Breeding 38, 63 (2018). https://doi.org/10.1007/s11032-018-0828-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-018-0828-8