Abstract
Self-incompatibility (SI) is an important trait of Citrus plants that is exploited by farmers to produce seedless fruit. However, the molecular mechanism of SI in Citrus is not well understood. Wuzishatangju (Citrus reticulata Blanco) (SI) is an excellent seedless cultivar selected from a seedy Shatangju cultivar (self-compatible, SC) through spontaneous bud mutation. The two cultivars are therefore excellent materials for studying the mechanisms of SI and/or SC in Citrus. In this study, an integrative strategy combining eight suppression subtractive hybridization libraries with cDNA microarray was used to study the molecular mechanisms that differ between Wuzishatangju and Shatangju (control) mandarins. A custom microarray screen resulted in a total of 1,830 up- or down-regulated clones (false discovery rate <0.05 and a fold change \({ \geqq }\)2) obtained from 9,810 positive clones. The expression of genes involved in embryonic development, ubiquitination pathway, Ca2+-signaling pathway, gibberellins, and auxin was significantly up-regulated in SI Wuzishatangju compared with SC Shatangju mandarin. The microarray analysis suggested that the ubiquitin-mediated proteolysis pathway might be involved in the SI reaction of Wuzishatangju. Additionally, our research highlighted some main genes (mitogen-activated protein kinase, SI S1 family protein, ubiquitin-conjugating factor E4-like, auxin transporter protein 1, and gibberellin receptor) that participate in the SI reaction of Wuzishatangju and could be beneficial for understanding the evolution of SI systems and for breeding seedless citrus fruits in the future.





Similar content being viewed by others
References
Brock AK, Willmann R, Kolb D, Grefen L, Lajunen HM, Bethke G, Lee J, Nürnberger T, Gust AA (2010) The Arabidopsis mitogen-activated protein kinase phosphatase PP2C5 affects seed germination, stomatal aperture, and abscisic acid-inducible gene expression. Plant Physiol 153:1098–1111
Busch JW, Schoen DJ (2008) The evolution of self-incompatibility when mates are limiting. Trends Plant Sci 13:128–136
Caruso M, Merelo P, Distefano G, La Malfa S, Lo Piero AR, Tadeo FR, Talon M, Gentile A (2012) Comparative transcriptome analysis of stylar canal cells identifies novel candidate genes implicated in the self-incompatibility response of Citrus clementina. BMC Plant Biol 12:20
Chai L, Biswas MK, Ge X, Deng X (2010) Isolation, characterization, and expression analysis of an Skp 1 -like gene from ‘Shatian’ Pummelo (Citrus grandis Osbeck). Plant Mol Biol Rep 28:569–577
Chai L, Ge X, Biswas MK, Deng X (2011) Molecular analysis and expression of a floral organ-relative F-box gene isolated from ‘Zigui Shatian’ pummelo (Citrus grandis Osbeck). Mol Biol Rep 38:4429–4436
de Nettancourt D (1997) Incompatibility in angiosperms. Sex Plant Reprod 10:185–199
Distefano G, Caruso M, La Malfa S, Gentile A, Tribulato E (2009) Histological and molecular analysis of pollen-pistil interaction in clementine. Plant Cell Rep 28:1439–1451
Foote HC, Ride JP, Franklin-Tong VE, Walker EA, Lawrence MJ, Franklin FC (1994) Cloning and expression of a distinctive class of self-incompatibility (S) gene from Papaver rhoeas L. Proc Natl Acad Sci USA 91:2265–2269
Gao ZH, Wang PP, Zhuang WB, Zhang Z (2013) Sequence analysis of new S-RNase and SFB alleles in Japanese apricot (Prunus mume). Plant Mol Biol Rep 31:751–762
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435
Gu C, Wu J, Du YH, Yang YN, Zhang SL (2013) Two different Prunus SFB alleles have the same function in the self-incompatibility reaction. Plant Mol Biol Rep 31:425–434
Guo Y, Guo H, Zhang L, Xie H, Zhao X, Wang F, Li Z, Wang Y, Ma S, Tao J, Wang W, Zhou Y, Yang W, Cheng J (2005) Genomic analysis of anti-hepatitis B virus (HBV) activity by small interfering RNA and lamivudine in stable HBV-producing cells. J Virol 79:14392–14403
Hatakeyama S, Yada M, Matsumoto M, Ishida N, Nakayama KI (2001) U box proteins as a new family of ubiquitin-protein ligases. J Biol Chem 276:33111–33120
Honsho C, Kotsubo M, Fukuda Y, Hamabata Y (2009) Reproductive characteristics for self-compatibility and seedlessness in ‘Nishiuchi Konatsu’, a bud mutation of Hyuganatsu (Citrus tamurana hort. ex Tanaka). HortScience 44:1547–1551
Iwano M, Takayama S (2012) Self/non-self discrimination in angiosperm self-incompatibility. Curr Opin Plant Biol 15:78–83
Koegl M, Hoppe T, Schlenker S, Ulrich HD, Mayer TU, Jentsch S (1999) A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96:635–644
Kubo K, Entani T, Takara A, Wang N, Fields AM, Hua Z, Toyoda M, Kawashima S, Ando T, Isogai A, Kao TH, Takayama S (2010) Collaborative non-self recognition system in S-RNase-Based self-incompatibility. Science 330:796–799
Li S, Samaj J, Franklin-Tong VE (2007) A mitogen-activated protein kinase signals to programmed cell death induced by self-incompatibility in Papaver pollen. Plant Physiol 145:236–245
Liu Q, Zhu A, Chai L, Zhou W, Yu K, Ding J, Xu J, Deng X (2009) Transcriptome analysis of a spontaneous mutant in sweet orange [Citrus sinensis (L.) Osbeck] during fruit development. J Exp Bot 60:801–813
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the \(2^{{{-}{\Delta \Delta }C_{\text{T}} }}\) method. Methods 25:402–408
Luo M, Xiao Y, Hou L, Luo X, Li D, Pei Y (2003) Cloning and expression analysis of LIM-domain protein gene from cotton (Gossypium hirsutum L.). Acta Genet Sin 30:175–182
McGinnis KM, Thomas SG, Soule JD, Strader LC, Zale JM, Sun TP, Steber CM (2003) The Arabidopsis SLEEPY1 gene encodes a putative F-box subunit of an SCF E3 ubiquitin ligase. Plant Cell 15:1120–1130
Mesejo C, Yuste R, Martínez-Fuentes A, Reig C, Iglesias DJ, Primo-Millo E, Agustí M (2013) Self-pollination and parthenocarpic ability in developing ovaries of self-incompatible Clementine mandarins (Citrus clementina). Physiol Plant 148:87–96
Miao H, Qin Y, Teixeira da Silva JA, Ye Z, Hu G (2011) Cloning and expression analysis of S-RNase homologous gene in Citrus reticulate Blanco cv. Wuzishatangju. Plant Sci 180:358–367
Miao H, Qin Y, Teixeira da Silva JA, Ye Z, Hu G (2013a) Identification of differentially expressed genes in pistils from self-incompatible Citrus reticulata by suppression subtractive hybridization. Mol Biol Rep 40:159–169
Miao H, Qin Y, Ye Z, Hu G (2013b) Molecular characterization and expression analysis of ubiquitin-activating enzyme E1 gene in Citrus reticulata. Gene 513:249–259
Miao H, Ye Z, Teixeira da Silva JA, Qin Y, Hu G (2013c) Identifying differentially expressed genes in pollen from self-incompatible “Wuzishatangju” and self-compatible “Shatangju” mandarins. Int J Mol Sci 14:8538–8555
Miao H, Ye Z, Qin Y, Hu G (2013d) Molecular characterization and expression analysis of S1 self-incompatibility locus-linked pollen 3.15 gene in Citrus reticulata. J Integr Plant Biol 55:443–452
Moon J, Parry G, Estelle M (2004) The ubiquitin-proteasome pathway and plant development. Plant Cell 16:3181–3195
Ngo BX, Wakana A, Kim JH, Mori T, Sakai K (2010) Estimation of self-incompatibility S genotypes of Citrus cultivars and plants based on controlled pollination with restricted number of pollen grains. J Fac Agric Kyushu Univ 55:67–72
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11:3194–3205
Poulter NS, Staiger CJ, Rappoport JZ, Franklin-Tong VE (2010) Actin-binding proteins implicated in the formation of the punctuate actin foci stimulated by the self-incompatibility response in Papaver. Plant Physiol 152:1274–1283
Qiao H, Wang F, Zhao L, Zhou J, Lai Z, Zhang Y, Robbins TP, Xue Y (2004) The F-box protein AhSLF-S2 controls the pollen function of S-RNase-based self-incompatibility. Plant Cell 16:2307–2322
Qiu W, Zhu A, Wang Y, Chai L, Ge X, Deng X, Guo W (2012) Comparative transcript profiling of gene expression between seedless Ponkan mandarin and its seedy wild type during floral organ development by suppression subtractive hybridization and cDNA microarray. BMC Genomics 13:397
Roiz L, Goren R, Shoseyov O (1995) Stigmatic RNase in calamondin (Citrus reticulata var. austera × Fortunella sp.). Physiol Plant 94:585–590
Rudd JJ, Franklin-Tong VE (2003) Signals and targets of the self-incompatibility response in pollen of Papaver rhoeas. J Exp Bot 54:141–148
Sasaki A, Itoh H, Gomi K, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Jeong DH, An G, Kitano H, Ashikari M, Matsuoka M (2003) Accumulation of phosphorylated repressor for gibberellin signaling in an F-box mutant. Science 299:1896–1898
Sassa H, Kakui H, Minamikawa M (2010) Pollen-expressed F-box gene family and mechanism of S-RNase-based gametophytic self-incompatibility (GSI) in Rosaceae. Sex Plant Reprod 23:39–43
Schomburg FM, Bizzell CM, Lee DJ, Zeevaart JA, Amasino RM (2003) Overexpression of a novel class of gibberellin 2-oxidases decreases gibberellin levels and creates dwarf plants. Plant Cell 15:151–163
Sullivan JA, Shirasu K, Deng XW (2003) The diverse roles of ubiquitin and the 26S proteasome in the life of plants. Nat Rev Genet 4:948–958
Sun P, Kao TH (2013) Self-incompatibility in Petunia inflata: the relationship between a self-incompatibility locus F-box protein and its non-self S-RNases. Plant Cell 25:470–485
Thomas SG, Franklin-Tong VE (2004) Self-incompatibility triggers programmed cell death in Papaver pollen. Nature 429:305–309
Tsuchimatsu T, Suwabe K, Shimizu-lnatsugi R, Isokawa S, Pavlidis P, Städler T, Suzuki G, Takayama S, Watanabe M, Shimizu KK (2010) Evolution of self-incompatibility in Arabidopsis by a mutation in the male specificity gene. Nature 464:1342–1346
Wakana A, Ngo BX, Fukudome I, Kajiwara K (2004) Estimation of the degree of self-incompatibility reaction during flower bud development and production of self-fertilized seeds by bud pollination in self-incompatible Citrus cultivars. J Fac Agric Kyushu Univ 49:307–320
Wang P, Lü L (2009) Self-incompatible reaction parts in Citrus grandis ‘Guanximiyou’ and ‘Duweimiyou’—observation of pollination on different pistil parts in vitro. Chin J Trop Crops 30:1105–1108
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J, Speed TP (2002) Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 30:15
Ye Z, Zeng T, Xu J, Luo Z, Hu G, Zhang Z, Ji Z, Chen Y, Chen G, Chen L, Lin S (2006) Wuzishatangju, a new mandarin cultivar. J Fruit Sci 23:149–150
Ye W, Qin Y, Ye Z, Teixeira da Silva JA, Zhang L, Wu X, Lin S, Hu G (2009) Seedless mechanism of a new mandarin cultivar Wuzishatangju (Citrus reticulata Blanco). Plant Sci 177:19–27
Zhang SW, Ding F, He XH, Luo C, Huang GX, Hu Y (2014) Characterization of the ‘Xiangshui’ lemon transcriptome by de novo assembly to discover genes associated with self-incompatibility. Mol Genet Genomics. doi:10.1007/s00438-014-0920-7
Acknowledgments
This work was supported by National Natural Science Foundation of China (No. 31000899 and 31471858), Research Fund for the Doctoral Program of Higher Education of China (No. 20104404120015 and 20114404110018), Guangdong Province Science Foundation of China (No. S2013020013084, S2013010011950 and 06025843), Foundation for Higher Education Discipline and Specialty Construction of Guangdong Provincial Department of Education (No. 2013KJCX0031), Science and Technology Planning Project of Guangzhou (2010r1-C771), the Open Foundation of State Key Laboratory for Conservation and Utilization of Subtropical Agro-bioresources, South China Agricultural University (No. KSL-CUSAb-2012-09), Key Laboratory of Innovation and Utilization for Germplasm Resources in Horticultural Crops in Southern China of Guangdong Higher Education Institutes, South China Agricultural University (No. KBL11008) and “211” Construction Fund for Key Subjects of College of Horticulture, South China Agricultural University.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2015_204_MOESM3_ESM.tif
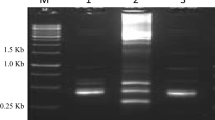
Supplement S3 Agarose gel electrophoresis of total RNA. 1: Pistils of Wuzishatangju; 2: Pistils of Shatangju; 3: Pollen of Wuzishatangju; 4: Pollen of Shatangju; 5: 72 h styles after self-pollination of Wuzishatangju; 6: 72 h styles after cross-pollination of Wuzishatangju × Shatangju; 7, 9, 11, 13, 15, 17: Pistils in differential stages (0, 4, 24, 48, 72, and 96 h) after self-pollination of Wuzishatangju; 8, 10, 12, 14, 16, 18: Pistils in differential stages (0, 4, 24, 48, 72, and 96 h) after cross-pollination of Wuzishatangju. (TIFF 326 kb)
11032_2015_204_MOESM4_ESM.tif
Supplement S4 TreeView representation of ESTs from eight SSH libraries. WY: Forward SSH library of Wuzishatangju pistils; YW: Reverse SSH library of Wuzishatangju pistils; H: Forward SSH library of Wuzishatangju pollen; F: Reverse SSH library of Wuzishatangju pollen; T1: Forward SSH library of 72 h styles after self-pollination of Wuzishatangju; T2: Reverse SSH library of 72 h styles after cross-pollination of Wuzishatangju × Shatangju; 0 h1, 24 h5, 48 h7, 72 h9, 96 h11: Forward SSH library of pistils in differential stages (0, 24, 48, 72, and 96 h) after self-pollination of Wuzishatangju; 0 h2, 24 h6, 48 h8, 72 h10, 96 h12: Reverse SSH library of pistils in differential stages (0, 4, 24, 48, 72, and 96 h) after cross-pollination of Wuzishatangju. (TIFF 1730 kb)
11032_2015_204_MOESM5_ESM.tif
Supplement S5 Scatter plots for the eight SSH libraries. WY: Forward SSH library of Wuzishatangju pistils; YW: Reverse SSH library of Wuzishatangju pistils; H: Forward SSH library of Wuzishatangju pollen; F: Reverse SSH library of Wuzishatangju pollen; T1: Forward SSH library of 72 h styles after self-pollination of Wuzishatangju; T2: Reverse SSH library of 72 h styles after cross-pollination of Wuzishatangju × Shatangju; 72 h9: Forward SSH library of pistils in 72 h after self-pollination of Wuzishatangju; 72 h10: Reverse SSH library of pistils in 72 h after cross-pollination of Wuzishatangju × Shatangju. (TIFF 1780 kb)
11032_2015_204_MOESM7_ESM.tif
Supplement S7 Up- and down- regulated clones of eight SSH libraries. A (0, 24, 48, 72, and 96 h): Pistils in different stages (0, 24, 48, 72, and 96 h) after self-pollination of Wuzishatangju mandarin. B (0, 24, 48, 72, and 96 h): Pistils in differential stages (0, 24, 48, 72, and 96 h) after cross-pollination of Wuzishatangju × Shatangju mandarin. (TIFF 3079 kb)
11032_2015_204_MOESM9_ESM.jpg
Supplement S9 A tentative model regarding the main genes and/or pathways involved in the SI reaction of Wuzishatangju mandarin ubiquitin-mediated proteolysis signaling pathway (JPEG 165 kb)
Rights and permissions
About this article
Cite this article
Miao, H., Ye, Z., Hu, G. et al. Comparative transcript profiling of gene expression between self-incompatible and self-compatible mandarins by suppression subtractive hybridization and cDNA microarray. Mol Breeding 35, 47 (2015). https://doi.org/10.1007/s11032-015-0204-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0204-x