Abstract
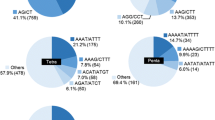
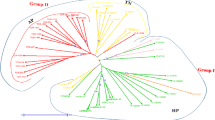
Bermudagrass lags in genomic and molecular breeding resources, particularly regarding a critical mass of robust, reproducible, and highly polymorphic molecular markers like simple sequence repeats (SSR). Here, 2017 sugarcane EST-SSR primer pairs (PPs) were screened for transferability and polymorphisms against commercial triploid hybrids and representatives of their parental species, tetraploid Cynodon dactylon and diploid C. transvaalensis. Fifty-four percent of PPs amplified target SSR in at least one of the two species, while 62% of these ‘transferable’ SSR were polymorphic. A subset of 228 polymorphic markers was utilized for genotyping 24 samples that included: 10 ‘Tif’ series cultivars (i.e., released from Tifton, GA), their parental species, 10 F1 hybrids between the two species, and two diverse Sorghum spp. A total of 90 and 144 PPs produced reproducible bands in the bermudagrass samples and the sorghum species, respectively. While 63 (70%) PPs were polymorphic among members of the ‘Tif’ diversity panel, 79 (54.8%) were polymorphic between the sorghum species. Further, 57 PPs polymorphic in the ‘Tif’ diversity panel were genotyped against 3 experimental lines and 7 commercial hybrids including ‘TifTuf’, a recent addition to the ‘Tif’ series. Joint analysis of 24 genotypes grouped bermudagrass accessions into two main clusters, one including C. dactylon and the other including C. transvaalensis. A small set of informative SSR readily differentiated the cultivar groups and identified potentially mischaracterized cultivars, but did not differentiate among mutants within cultivar groups. Heterologous EST-SSR from sugarcane are useful sources of polymorphic markers for cultivar identification in bermudagrass, and build on existing marker resources to facilitate genetic mapping, QTL analysis, and marker assisted selection.



Similar content being viewed by others
References
Aljanabi SM, Forget L, Dookun A (1999) An improved and rapid protocol for the isolation of polysaccharide- and polyphenol-free sugarcane DNA. Plant Mol Biol Rep 17:281–282
Bassam BJ, Caetano-Anollés G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Bethel CM, Sciara EB, Estill JC, Bowers JE, Hanna W, Paterson AH (2006) A framework linkage map of bermudagrass (Cynodon dactylon × transvaalensis) based on single-dose restriction fragments. Theor Appl Genet 112:727–737
Bowers JE, Abbey C, Anderson S, Chang C, Draye X, Hoppe AH, Jessup R, Lemke C, Lennington J, Li ZK, Lin YR, Liu SC, Luo LJ, Marler BS, Ming R, Mitchell SE, Qiang D, Reischmann K, Schulze SR, Skinner DN, Wang YW, Kresovich S, Schertz KF, Paterson AH (2003) A high-density genetic recombination map of sequence-tagged sites for Sorghum, as a framework for comparative structural and evolutionary genomics of tropical grains and grasses. Genetics 165:367–386
Burton GW (1960) Tifway Bermudagrass. US Golf Ass J 13:28–30
Burton GW (1966a) Tifway (Tifton 419) bermudagrass (Reg. No. 7). Crop Sci 6(1):93–94
Burton GW (1966b) Tifdwarf bermudagrass (Reg. No. 8). Crop Sci 6(1):94
Burton GW (1985) Registration of Tifway II bermudagrass. Crop Sci 25(2):364
Burton GW (1991) A history of turf research at Tifton. USGA Green Sect Rec 29:12–14
Caetano-Anollés G (1998) Genetic instability of bermudagrass (Cynodon) cultivars ‘Tifgreen’ and ‘Tifdwarf’ detected by DAF and ASAP analysis of accessions and off-types. Euphytica 101:165–173
Caetano-Anollés G, Callahan LM, Gresshoff M (1997) The origin of bermudagrass (Cynodon) types inferred by DNA amplification fingerprinting. Crop Sci 37(1):81–87
Chen Z, Raymer P, Waltz C, Wang ML (2009) Genetic diversity of warm-season turfgrass: seashore paspalum, bermudagrass, and zoysiagrass revealed by AFLPs. Floric Ornam Biotechnol 3:20–24
Clayton WD, Renvoize SA (1986) Genera graminum. Grasses of the World. Kew Bulletin Additional Series, pp 389–389
Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ (2001) Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Sci 160:1115–1123
De Bruijn J (2012) U.S. Patent No. PP22,963. Washington, DC: U.S. Patent and Trademark Office
De Wet JMJ, Harlan JR (1970) Biosystematics of Cynodon L. C. Rich. (Gramineae). International Bureau for Plant Taxonomy and Nomenclature, p 565
Ellis JR, Burke JM (2007) EST-SSRs as a resource for population genetic analyses. Heredity 99:125–132
Felsenstein J (1985) Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Society for the Study of Evolution, p 783
Felsenstein J (1988) The PHYLIP phylogeny inference package. Abstracts with Programs. Geol Soc Am 20:185
Gulsen O, Sever-Mutlu S, Mutlu N, Tuna M, Karaguzel O, Shearman RC, Riordan TP, Heng-Moss TM (2009) Polyploidy creates higher diversity among Cynodon accessions as assessed by molecular markers. Theor Appl Genet 118:1309–1319
Guo Y, Wu Y, Anderson JA, Moss JQ, Zhu L (2015) Disomic Inheritance and Segregation Distortion of SSR Markers in Two Populations of Cynodon dactylon (L.) Pers. var. dactylon. PLoS ONE 10:1–10
Hanna WW, Elsner JE (1999) Registration of ‘TifEagle’ bermudagrass. Crop Sci 39:1258
Hanna WW, Schwartz BM (2016) U.S. Patent No. PP27392 P2. Washington, DC: U.S. Patent and Trademark Office
Hanna WW, Burton GW, Johnson AW (1990) Registration of ‘Tifton 10’ turf bermudagrass. Crop Sci 30(6):1355–1356
Hanna WW, Carrow RN, Powell AJ (1997) Registration of ‘Tift 94’ bermudagrass. Crop Sci 37:1012
Hanna WW, Braman SK, Schwartz BM (2010) ‘ST-5’, a shade-tolerant turf bermudagrass. HortScience 45(1):132–134
Harlan JR (1970) Cynodon species and their value for grazing and hay. Herbage Abstracts 40:233–238
Harris-Shultz KR, Schwartz BM, Hanna WW, Brady JA (2010a) Development, linkage mapping, and use of microsatellites in bermudagrass. J Am Soc Hortic Sci 135:511–520
Harris-Shultz KR, Schwartz BM, Paterson AH, Brady JA (2010b) Identification and mapping of nucleotide binding site-leucine-rich repeat resistance gene analogs in bermudagrass. J Am Soc Hortic Sci 135:74–82
Harris-Shultz KR, Schwartz BM, Brady JA (2011) Identification of simple sequence repeat markers that differentiate bermudagrass cultivars derived from ‘Tifgreen’. J Am Soc Hortic Sci 136:211–218
Hein MA (1953) Registration of varieties and strains of bermuda grass, II (Cynodon dactylon (L.) Pers.). Agron J 45(11):572–573
Hein MA (1961) Registration of varieties and strains of bermudagrass, III. (Cynodon dactylon (L.) Pers.). Agron J 53:276
Jewell MC, Frere CH, Prentis PJ, Lambrides CJ, Godwin ID (2010) Characterization and multiplexing of EST-SSR primers in Cynodon (Poaceae) species. Am J Bot 97:e99–e101
Kamps TL, Williams NR, Ortega VM, Chamusco KC, Harris-Shultz K, Scully BT, Chase CD (2011) DNA polymorphisms at bermudagrass microsatellite loci and their use in genotype fingerprinting. Crop Sci 51:1122–1131
Kantety RV, Ml Rota, Matthews DE, Sorrells ME (2002) Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant Mol Biol 48:501–510
Kim C, Jang CS, Kamps TL, Robertson JS, Feltus FA, Paterson AH (2008) Transcriptome analysis of leaf tissue from Bermudagrass (Cynodon dactylon) using a normalised cDNA library. Funct Plant Biol 35:585–594
Kim C, Tang H, Paterson AH (2009) Duplication and divergence of grass genomes: integrating the Chloridoids. Trop Plant Biol 2:51–62
Kim C, Zhang D, Auckland SA, Rainville LK, Jakob K, Kronmiller B, Sacks EJ, Deuter M, Paterson AH (2012) SSR-based genetic maps of Miscanthus sinensis and M. sacchariflorus, and their comparison to sorghum. Theor Appl Genet 124(7):1325–1338
Kong W, Jin H, Franks CD, Kim C, Bandopadhyay R, Rana MK, Auckland SA, Goff VH, Rainville LK, Burow GB, Woodfin C, Burke JJ, Paterson AH (2013) Genetic analysis of recombinant inbred lines for Sorghum bicolor × Sorghum propinquum. G3-Genes Genom Genet 3:101–108
Ling Y, Zhang XQ, Ma X, Chen SY, Chen TT, Liu W (2012) Analysis of genetic diversity among wild bermudagrass germplasm from southwest China using SSR markers. Genet Mol Res 11:4598–4608
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Lu Y, Jiang J, Yi Z (2012) Study on the transferability of maize SSR and sugarcane EST-SSR markers to Miscanthus (Poaceae). Acta Prataculturae Sinica 21:86–95
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Parsons DLV, Lehman VG (2007) U.S. Patent No. PP18,247. Washington, DC: U.S. Patent and Trademark Office
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang H, Wang X, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Paterson AH, Wang X, Tang H, Kim C (2013) Comparative genomics of grasses: a saccharinae-centric view. In: Paterson AH (ed) Genomics of the Saccharinae. Springer, New York, pp 429–445
Peakall R, Smouse PE (2006) Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Reasor EH, Brosnan JT, Trigiano RN, Elsner JE, Henry GM, Schwartz BM (2016) The genetic and phenotypic variability of interspecific hybrid bermudagrasses (Cynodon dactylon (L.) Pers. × C. transvaalensis Burtt-Davy) used on golf course putting greens. Planta doi:10.1007/s00425-016-2573-8
Riley RJ (2000) U.S. Patent No. PP11,181. Washington, DC: U.S. Patent and Trademark Office
Saha MC, Mian MAR, Eujayl I, Zwonitzer JC, Wang L, May GD (2004) Tall fescue EST-SSR markers with transferability across several grass species. Theor Appl Genet 109:783–791
Sharma RK, Gupta P, Sharma V, Sood A, Mohapatra T, Ahuja PS (2008) Evaluation of rice and sugarcane SSR markers for phylogenetic and genetic diversity analyses in bamboo. Genome 51:91–103
Taliaferro CM, Martin DL, Anderson JA, Anderson MP (2006) U.S. Patent No. PP16,801. Washington, DC: U.S. Patent and Trademark Office
Tan C, Wu Y, Taliaferro CM, Anderson M, Tauer C, Samuels T (2012) Development of simple sequence repeat markers for bermudagrass from its expressed sequence tag sequences and preexisting sorghum SSR markers. Mol Breed 29:23–30
Tan C, Wu Y, Taliaferro CM, Bell GE, Martin DL, Smith MW (2014) Development and characterization of genomic SSR markers in Cynodon transvaalensis Burtt-Davy. Mol Genet Genomics 289:523–531
Varshney RK, Thiel T, Stein N, Langridge P, Graner A (2002) In silico analysis on frequency and distribution of microsatellites in ESTs of some cereal species. Cell Mol Biol Lett 7:537–546
Wang ML, Barkley NA, Yu JK, Dean RE, Newman ML, Sorrells ME, Pederson GA (2005) Transfer of simple sequence repeat (SSR) markers from major cereal crops to minor grass species for germplasm characterization and evaluation. Plant Genet Resour C 3:45–57
Wang Z, Samuels T, Tan C, Gao H, Wu Y, Dl Martin (2010) Identification of vegetatively propagated turf bermudagrass cultivars using simple sequence repeat markers. Crop Sci 50:2103–2111
Wu YQ, Taliaferro CM, Bai GH, Anderson MP (2004) AFLP analysis of Cynodon dactylon (L.) Pers. var. dactylon genetic variation. Genome 47:689–696
Wu YQ, Taliaferro CM, Bai GH, Anderson MP (2005) Genetic diversity of Cynodon transvaalensis Burtt-Davy and its relatedness to hexaploid C. dactylon (L.) Pers. as indicated by AFLP markers. Crop Sci 45:848–853
Wu YQ, Martin DL, Taliaferro CM, Anderson JA, Moss JQ (2013) U.S. Patent No. PP24,116. Washington, DC: U.S. Patent and Trademark Office
Wu YQ, Martin DL, Taliaferro CM, Anderson JA, Moss JQ (2014) U.S. Patent No. PP24,271. Washington, DC: U.S. Patent and Trademark Office
Yerramsetty PN, Anderson MP, Taliaferro CM, Martin DL (2008) Genetic variations in clonally propagated bermudagrass cultivars identified by DNA fingerprinting. Plant Omics 1:1–8
Zhang LH, Ozias-Akins P, Kochert G, Kresovich S, Dean R, Hanna W (1999) Differentiation of bermudagrass (Cynodon spp.) genotypes by AFLP analyses. Theor Appl Genet 98:895–902
Zhiyong W, Li L, Xuejun Y, Hailin G, Aigui G, Jianxiu L (2013) Genetic diversity analysis of Cynodon dactylon (bermudagrass) accessions and cultivars from different countries based on ISSR and SSR markers. Biochem Syst Ecol 46:108–115
Acknowledgements
We thank Wayne Hanna for his feedback on the manuscript. This work was financially supported by the United States Golf Association (USGA), Georgia Seed Development Commission (GSDC), Georgia Crop Improvement Association (GCIA), and Georgia Agricultural Experiment Station.
Author contribution
SK was responsible for designing and performing the experiments, data analysis and manuscript writing as a part of his PhD research. BS served as SK’s PhD committee member, provided germplasm included in the study, maintained the mapping population, and provided feedback on the proposed work and manuscript before submission.CK helped in marker screening and reviewed data analysis. SAA, LKR and JA performed experiments. AHP conceived the project, served as SK’s major advisor and revised the manuscript before submission.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Khanal, S., Schwartz, B.M., Kim, C. et al. Cross-taxon application of sugarcane EST-SSR to genetic diversity analysis of bermudagrass (Cynodon spp.). Genet Resour Crop Evol 64, 2059–2070 (2017). https://doi.org/10.1007/s10722-017-0496-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-017-0496-2