Abstract
White bulb colors of onion (Allium cepa L.) are determined by the C and I loci which control the so-called recessive and dominant white bulb colors, respectively. To identify the causal gene responsible for the I locus, a combined approach of bulked segregant analysis and RNA-Seq was used in the present study. A total of 68 contigs containing homozygous single nucleotide polymorphisms (SNPs) between red and dominant white bulked RNAs were identified. The position of I locus was found to be located at chromosome 3 by performing comparative analysis of these contigs and using a previously constructed linkage map. After verification of homozygous SNPs by sequencing of PCR products, 12 high resolution melting, one cleaved amplified polymorphic sequence, and one InDel markers were developed. A linkage map flanking the I locus was constructed using these markers. Two tightly linked markers (DW51596 and DW35019) flanking the I locus were identified by analyzing 1457 F4 individuals. A total of 104 and 39 contigs showing more than tenfold increase of expression in red and dominant white bulks, respectively, were identified. Transcriptions of all structural genes encoding enzymes in flavonoid biosynthesis pathway were significantly reduced in the dominant white bulk. Transcription levels of most contigs showing more than tenfold reduced expression in dominant white were also significantly reduced in the recessive white bulbs controlled by the C locus. Genomic DNA sequences of 12 genes encoding transcription factors assumed to regulate flavonoid biosynthesis were analyzed. However, the causal gene for the I locus could not be identified.




Similar content being viewed by others
References
Amawi H, Ashby CR Jr, Tiwari AK (2017) Cancer chemoprevention through dietary flavonoids: what’s limiting? Chin J Cancer 36:50
Andersen JR, Lübberstedt T (2003) Functional markers in plants. Trends Plant Sci 11:554–560
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9:208–218
Baek G, Kim C, Kim S (2017) Development of a molecular marker tightly linked to the C locus conferring a white bulb color in onion (Allium cepa L.) using bulked segregant analysis and RNA-seq. Mol Breed 37:94
Cao J, Chen W, Zhang Y, Zhang Y, Zhao X (2010) Content of selected flavonoids in 100 edible vegetables and fruits. Food Sci Technol Res 16:395–402
Chin S, Behm CA, Mathesius U (2018) Functions of flavonoids in plant-nematode interactions. Plants 7:85
Clarke AE, Jones HA, Little TM (1944) Inheritance of bulb color in the onion. Genetics 29:569–575
Davis EW (1954) Rapid identification of recessive onion bulbs by use of ammonia fumes. J Hered 45:122
Dixon RA, Pasinetti GM (2010) Flavonoids and isoflavonoids: from plant biology to agriculture and neuroscience. Plant Physiol 154:453–457
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Duangjit J, Bohanec B, Chan AP, Town CD, Havey MJ (2013) Transcriptome sequencing to produce SNP-based genetic maps of onion. Theor Appl Genet 126:2093–2101
El-Shafie MW, Davis GN (1967) Inheritance of bulb color in the onion (Allium cepa L.). Hilgardia 38:607–622
Farhadi F, Khameneh B, Iranshahi M, Iranshahy M (2018) Antibacterial activity of flavonoids and their structure–activity relationship: an update review. Phytother Res 33:13–40
Fernández-Rojas B, Gutiérrez-Venegas G (2018) Flavonoids exert multiple periodontic benefits including anti-inflammatory, periodontal ligament-supporting, and alveolar bone-preserving effects. Life Sci 209:435–454
Fini A, Brunetti C, Di Ferdinando M, Ferrini F, Tattini M (2011) Stress-induced flavonoid biosynthesis and the antioxidant machinery of plants. Plant Signal Behav 6:709–711
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, Macmanes MD, Ott M, Orvis J, Pochet N, Strozzi F, Weeks N, Westerman R, William T, Dewey CN, Henschel R, Leduc RD, Friedman N, Regev A (2013) De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8:1494–1512
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Window 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hichri I, Barrieu F, Bogs J, Kappel C, Delrot S, Lauvergeat V (2011) Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J Exp Bot 62:2465–2483
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1070–1083
Jaakola L (2013) New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci 18:477–483
Jo C, Kim S (2020) Transposition of a non-autonomous DNA transposon in the gene coding for a bHLH transcription factor results in a white bulb color of onions (Allium cepa L.). Theor Appl Genet 133:317–328
Kim B, Kim S (2019) Identification of a variant of CMS-T cytoplasm and development of high resolution melting markers for distinguishing cytoplasm types and genotyping a restorer-of-fertility locus in onion (Allium cepa L.). Euphytica 215:164
Kim S, Binzel ML, Yoo K, Park S, Pike LM (2004a) Pink (P), a new locus responsible for pink trait in onions (Allium cepa) resulting from natural mutations of anthocyanidin synthase. Mol Gen Genom 272:18–27
Kim S, Jones R, Yoo K, Pike LM (2004b) Gold color in onions (Allium cepa): a natural mutation of the chalcone isomerase gene resulting in a premature stop codon. Mol Gen Genom 272:411–419
Kim S, Jones R, Yoo K, Pike LM (2005a) The L locus, one of complementary genes required for anthocyanin production in onions (Allium cepa), encodes anthocyanidin synthase. Theor Appl Genet 111:120–127
Kim S, Yoo K, Pike LM (2005b) Development of a PCR-based marker utilizing a deletion mutation in the DFR (dihydroflavonol 4-reductase) gene responsible for the lack of anthocyanin production in yellow onions (Allium cepa). Theor Appl Genet 110:588–595
Kim S, Yoo K, Pike LM (2005c) The basic color factor, the C locus, encodes a regulatory gene controlling transcription of chalcone synthase genes in onions (Allium cepa). Euphytica 142:273–282
Kim S, Kim M, Kim Y, Yeom S, Cheong K, Kim K, Jeon J, Kim S, Kim D, Sohn S, Lee Y, Choi D (2015a) Integrative structural annotation of de novo RNA-Seq provides an accurate reference gene set of the enormous genome of the onion (Allium cepa L.). DNA Res 22:19–27
Kim S, Park JY, Yang T (2015b) Characterization of three active transposable elements recently inserted in three independent DFR-A alleles and one high-copy DNA transposon isolated from the Pink allele of the ANS gene in onion (Allium cepa L.). Mol Genet Genom 290:1027–1037
Kim E, Kim C, Kim S (2016) Identification of two novel mutant ANS alleles responsible for inactivation of anthocyanidin synthase and failure of anthocyanin production in onion (Allium cepa L.). Euphytica 212:427–437
Lee YG, Cho J, Kim Y, Moon J (2016) Change in flavonoid composition and antioxidative activity during fermentation of onion (Allium cepa L.) by Leuconostoc mesenteroides with different salt concentrations. J Food Sci 81:C1385–C1393
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12:323
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup (2009) The Sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25:2078–2079
Mitra J, Shrivastava SL, Rao PS (2012) Onion dehydration: a review. J Food Sci Technol 49:267–277
Morita Y, Saitoh M, Hoshino A, Nitasaka E, Iida S (2006) Isolation of cDNAs for R2R3-MYB, bHLH and WDR transcriptional regulators and identification of c and ca mutations conferring white flowers in the Japanese morning glory. Plant Cell Physiol 47:457–470
Nakatsuka A, Yamagishi M, Nakano M, Tasaki K, Kobayashi N (2009) Light-induced expression of basic helix-loop-helix genes involved in anthocyanin biosynthesis in flowers and leaves of Asiatic hybrid lily. Sci Hortic 121:84–91
Passeri V, Koes R, Quattrocchio FM (2016) New challenges for the design of high value plant products: stabilization of anthocyanins in plant vacuoles. Front Plant Sci 7:153
Petroni K, Tonelli C (2011) Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci 181:219–229
Ramsay NA, Glover BJ (2005) MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10:63–70
Scarano A, Chieppa M, Santino A (2018) Looking at flavonoid biodiversity in horticultural crops: a colored mine with nutritional benefits. Plants 7:98
Slimestad R, Fossen T, Vågen IM (2007) Onions: a source of unique dietary flavonoids. J Agric Food Chem 55:10067–10080
Song S, Kim C, Moon JS, Kim S (2014) At least nine independent natural mutations of the DFR-A gene are responsible for appearance of yellow onions (Allium cepa L.) from red progenitors. Mol Breed 33:173–186
Spelt C, Quattrocchio F, Mol JN, Koes RE (2000) anthocyanin1 of Petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12:1619–1631
Van Ooijen JW, Voorrips RE (2001) JoinMap® 3.0, Software for the calculation of genetic linkage maps. Plant Res Int, Wageningen
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Xu Z, Feng K, Que F, Wang F, Xiong A (2017) A MYB transcription factor, DcMYB6, is involved in regulating anthocyanin biosynthesis in purple carrot taproots. Sci Rep 7:45324
Yamazaki M, Makita Y, Springob K, Saito K (2003) Regulatory mechanisms for anthocyanin biosynthesis in chemotypes of Perilla frutescens var. crispa. Biochem Eng J 14:191–197
Zaynab M, Fatima M, Abbas S, Sharif Y, Umair M, Zafar MH, Bahadar K (2018) Role of secondary metabolites in plant defense against pathogens. Microb Pathog 124:198–202
Acknowledgements
This research was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through Agriculture, Food and Rural Affairs Convergence Technologies Program for Educating Creative Global Leader funded by the Ministry of Agriculture, Food and Rural Affairs (710011-03), Golden Seed Project (Center for Horticultural Seed Development, No. 213007-05-4-SBB10), and a grant from the Next-Generation BioGreen 21 Program (Plant Molecular Breeding Center No. PJ013400). The authors thank Ji-wha Hur, Jeong-Ahn Yoo, and Su-jung Kim for their dedicated technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig.
1. Development of an HRM marker (DW12796) based on a single SNP between dominant white and red bulked DNAs. A. Alignment of nucleotide sequences flanking the SNP. Vertical arrow indicates the position of SNP. Horizontal arrows indicate primer-binding sites. B. Normalized melting peaks (upper) and curves (bottom) of the DW12796 marker. (TIFF 262 kb)
Supplementary Fig.
2. Flowchart showing the process determining linkage relationship of screened contigs relative to the position of I locus. Recombinants (R1 and R2) and Markers (M1, M2, M3, and M4) are examples used to describe the principle. (TIFF 178 kb)
Supplementary Fig.
3. Correlation for expression levels of all contigs between dominant white and red bulked RNAs. A. Reference transcriptome. B. De novo-assembled contigs. (TIFF 69 kb)
Supplementary Fig.
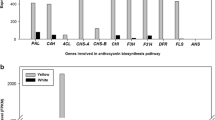
4. Comparison of expression levels of structural genes involved in flavonoid biosynthesis pathway between dominant white and red bulked RNAs. Detailed information of structural genes is shown in Baek et al. (2017). A. Reference transcriptome, B. De novo-assembled contigs. (TIFF 126 kb)
Supplementary Fig.
5. Genotyping of five types of onion bulb colors using molecular markers. Two CAPS markers, DW51596 and ANS-PS, were used to genotype the I locus and the L locus, respectively. DW: dominant white; YW: heterozygous white; RW: reddish white; R: red; Y: yellow; A: homozygous dominant control; H: heterozygous control; B: homozygous recessive control. (TIFF 154 kb)
Supplementary Fig.
6. Comparison of expression levels of onion bHLH-coding genes between dominant white and red bulked RNAs. Detail information of bHLH-coding genes is described in Baek et al. (2017). A. Reference transcriptome, B. De novo-assembled contigs. (TIFF 70 kb)
Supplementary Fig.
7. Comparison of expression levels of onion WD40-coding genes between dominant white and red bulked RNAs. Detail information of onion WD40-coding genes is described in Baek et al. (2017). A. Reference transcriptome, B. De novo-assembled contigs. (TIFF 91 kb)
Supplementary Fig.
8. Comparison of expression levels of onion MYB-coding genes between dominant white and red bulked RNAs. Detail information of onion MYB-coding genes is described in Baek et al. (2017). A. Reference transcriptome, B. De novo-assembled contigs. (TIFF 132 kb)
Supplementary Fig.
9. Comparison of bulb colors of reddish white and dominant white bulbs of the F3 segregating population (WR15). A. Reddish white onion bulbs, B. Dominant white onion bulbs. (TIFF 943 kb)
Rights and permissions
About this article
Cite this article
Seo, I., Kim, JG., Moon, JH. et al. Construction of a linkage map flanking the I locus controlling dominant white bulb color and analysis of differentially expressed genes between dominant white and red bulbs in onion (Allium cepa L.). Euphytica 216, 97 (2020). https://doi.org/10.1007/s10681-020-02638-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-020-02638-2