Abstract
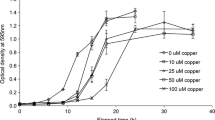
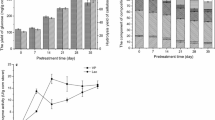
Degradation of most of the lignocellulose-rich agricultural residue in the cold regions of China is limited due to the cold climate. Lignin is the main component of lignocellulose, and the effective degradation of lignin is one of the most crucial processes in degrading lignocellulose. Psychrotrophic lignin-degrading bacteria and cold adapted ligninolytic enzymes have promising potential for the degradation and transformation of lignin, which are conducive to the resource utilization of lignocelluloses and energy-saving production under cold conditions. In this study, a newly psychrotrophic bacterial strain, Arthrobacter sp. C2, was isolated. The optimal enzyme activity conditions and lignin degradation pathways of C2 were investigated using sodium lignin sulfonate as substrate. The optimal conditions for enzyme activity included an initial pH of 6.74, a temperature of 14.9 °C, an incubation time of 6.87 days, and an inoculum size of 2.24%. Under the optimal conditions, the lignin peroxidase and manganese peroxidase activities and the degradation rate reached 29.8 U/L, 56.4 U/L and 40.1%, respectively. The biodegradation products including acids, phenols, aldehydes and alcohols were analyzed by gas chromatography–mass spectrometry and Fourier transform infrared spectroscopy. Further, the potential degradation pathways were proposed according to the results obtained in this study and those presented in the relevant literature. This study not only provides valuable psychrotrophic strain resources for the sustainable utilization of lignocellulose in cold regions, but also supplies potential application options for energy-saving production of useful chemicals using cold adapted enzymes.
Graphic abstract






Similar content being viewed by others
Abbreviations
- LiP:
-
Lignin peroxidase
- MnP:
-
Manganese peroxidase
- GC–MS:
-
Gas chromatography–mass spectrometry
- FTIR:
-
Fourier transform infrared spectroscopy
- RSM:
-
Response surface methodology
- BBD:
-
Box–Behnken design
- L-MSM:
-
Lignin mineral salt medium
- OD:
-
Optical density
- RT:
-
Retention time
- TCA:
-
Tricarboxylic acid
References
Ahmad M, Roberts JN, Hardiman EM, Singh R, Eltis LD, Bugg TD (2011) Identification of DypB from Rhodococcus jostii RHA1 as a lignin peroxidase. Biochemistry 50:5096–5107. https://doi.org/10.1021/bi101892z
Akila G, Chandra TS (2003) A novel cold-tolerant Clostridium strain PXYL1 isolated from a psychrophilic cattle manure digester that secretes thermolabile xylanase and cellulase. FEMS Microbiol Lett. 219:63–67. https://doi.org/10.1016/s0378-1097(02)01196-5
Asina F, Brzonova I, Voeller K, Kozliak E, Kubátová A, Yao B, Ji Y (2016) Biodegradation of lignin by fungi, bacteria and laccases. Bioresour Technol. 220:414–424. https://doi.org/10.1016/j.biortech.2016.08.016
Bai Y-l, Wang L, Lu Y-l, Yang L-p, Zhou L-p, Ni L, Cheng M-f (2015) Effects of long-term full straw return on yield and potassium response in wheat-maize rotation. J Integr Agric. 14:2467–2476. https://doi.org/10.1016/s2095-3119(15)61216-3
Blanco-Canqui H, Lal R (2009) Crop residue removal impacts on soil productivity and environmental quality. Crit Rev Plant Sci. 28:139–163. https://doi.org/10.1080/07352680902776507
Bloois EV, Pazmiño DET, Winter RT, Fraaije MW (2010) A robust and extracellular heme-containing peroxidase from Thermobifida fusca as prototype of a bacterial peroxidase superfamily. Appl Microbiol Biotechnol. 86:1419–1430. https://doi.org/10.1007/s00253-009-2369-x
Brodin M, Vallejos M, Opedal MT, Area MC, Chinga-Carrasco G (2017) Lignocellulosics as sustainable resources for production of bioplastics—a review. J Clean Prod. 162:646–664. https://doi.org/10.1016/j.jclepro.2017.05.209
Brown ME, Barros T, Chang MC (2012) Identification and characterization of a multifunctional dye peroxidase from a lignin-reactive bacterium. ACS Chem Biol. 7:2074–2081. https://doi.org/10.1021/cb300383y
Bugg TD, Ahmad M, Hardiman EM, Rahmanpour R (2011) Pathways for degradation of lignin in bacteria and fungi. Nat Prod Rep. 28:1883–1896. https://doi.org/10.1039/c1np00042j
Cao W, Xu H, Zhang H (2013) Architecture and functional groups of biofilms during composting with and without inoculation. Process Biochem. 48:1222–1226. https://doi.org/10.1016/j.procbio.2013.06.015
Chandra R, Bharagava RN (2013) Bacterial degradation of synthetic and kraft lignin by axenic and mixed culture and their metabolic products. J Environ Biol. 34:991–999. https://doi.org/10.1111/gbi.12061
Chen YH, Chai LY, Zhu YH, Yang ZH, Zheng Y, Zhang H (2012) Biodegradation of kraft lignin by a bacterial strain Comamonas sp. B-9 isolated from eroded bamboo slips. J Appl Microbiol. 112:900–906. https://doi.org/10.1111/j.1365-2672.2012.05275.x
Croce S, Wei Q, D'Imporzano G, Dong R, Adani F (2016) Anaerobic digestion of straw and corn stover: the effect of biological process optimization and pre-treatment on total bio-methane yield and energy performance. Biotechnol Adv. 34:1289–1304. https://doi.org/10.1016/j.biotechadv.2016.09.004
Datta R, Kelkar A, Baraniya D, Molaei A, Moulick A, Meena R, Formanek P (2017) Enzymatic degradation of lignin in soil: a review. Sustainability. 9:1163. https://doi.org/10.3390/su9071163
Diao Y, Song M, Zhang Y, Shi LY, Lv Y, Ran R (2017) Enzymic degradation of hydroxyethyl cellulose and analysis of the substitution pattern along the polysaccharide chain. Carbohydr Polym. 169:92–100. https://doi.org/10.1016/j.carbpol.2017.02.089
Dornez E, Verjans P, Arnaut F, Delcour JA, Courtin CM (2011) Use of psychrophilic xylanases provides insight into the xylanase functionality in bread making. J Agric Food Chem. 59:9553–9562. https://doi.org/10.1021/jf201752g
Duan J, Liang JD, Du WJ, Wang DQ (2014) Biodegradation of kraft lignin by a bacterial strain Sphingobacterium sp. HY-H. Adv Mat Res. 955–959:548–553
Fang S, An X, Liu H, Cheng Y, Hou N, Feng L, Huang X, Li C (2015) Enzymatic degradation of aliphatic nitriles by Rhodococcus rhodochrous BX2, a versatile nitrile-degrading bacterium. Bioresour Technol. 185:28–34. https://doi.org/10.1016/j.biortech.2015.02.078
Glenn JK, Akileswaran L, Gold MH (1986) Mn(II) oxidation is the principal function of the extracellular Mn-peroxidase from Phanerochaete chrysosporium. Arch Biochem Biophys. 251:688–696. https://doi.org/10.1016/0003-9861(86)90378-4
Guo X, Xie C, Wang L, Li Q, Wang Y (2019) Biodegradation of persistent environmental pollutants by Arthrobacter sp. Environ Sci Pollut Res Int. 26:8429–8443. https://doi.org/10.1007/s11356-019-04358-0
He T, Ye Q, Sun Q, Cai X, Ni J, Li Z, Xie D (2018) Removal of nitrate in simulated water at low temperature by a novel psychrotrophic and aerobic bacterium, Pseudomonas taiwanensis. Strain J Biomed Res Int 2018:4984087. https://doi.org/10.1155/2018/4984087
Hendriks AT, Zeeman G (2009) Pretreatments to enhance the digestibility of lignocellulosic biomass. Bioresour Technol. 100:10–18. https://doi.org/10.1016/j.biortech.2008.05.027
Hernandez-Pérez G, Goma G, Rols JL (1997) Degradation of lignosulfonated compounds by Streptomyces viridosporus strain T7A. Biotechnol Lett. 19:285–290. https://doi.org/10.1023/a:1018374111610
Hildebrandt P, Wanarska M, Kur J (2009) A new cold-adapted beta-D-galactosidase from the Antarctic Arthrobacter sp. 32c—gene cloning, overexpression, purification and properties. BMC Microbiol. 9:151. Doi: 10.1186/1471-2180-9-151
Hong J, Ren L, Hong J, Xu C (2016) Environmental impact assessment of corn straw utilization in China. J Clean Prod. 112:1700–1708. https://doi.org/10.1016/j.jclepro.2015.02.081
Horton RN, Apel WA, Thompson VS, Sheridan PP (2006) Low temperature reduction of hexavalent chromium by a microbial enrichment consortium and a novel strain of Arthrobacter aurescens. BMC Microbiol 6:1–8. https://doi.org/10.1186/1471-2180-6-5
Hou N, Feng F, Shi Y, Cao H, Li C, Cao Z, Cheng Y (2014) Characterization of the extracellular biodemulsifiers secreted by Bacillus cereus LH-6 and the enhancement of demulsifying efficiency by optimizing the cultivation conditions. Environ Sci Pollut Res Int. 21:10386–10398. https://doi.org/10.1007/s11356-014-2931-7
Hou N, Wen L, Cao H, Liu K, An X, Li D, Wang H, Du X, Li C (2017) Role of psychrotrophic bacteria in organic domestic waste composting in cold regions of China. Bioresour Technol. 236:20–28. https://doi.org/10.1016/j.biortech.2017.03.166
Karimi M, Esfandiar R, Biria D (2017) Simultaneous delignification and saccharification of rice straw as a lignocellulosic biomass by immobilized Thrichoderma viride sp. to enhance enzymatic sugar production. Renew Energy. 104:88–95. https://doi.org/10.1016/j.renene.2016.12.012
Kim SM, Park H, Choi JI (2016) Cloning and characterization of cold-adapted α-amylase from Antarctic Arthrobacter agilis. Appl Biochem Biotechnol. 181:1–12. https://doi.org/10.1007/s12010-016-2267-5
Kricka W, Fitzpatrick J, Bond U (2014) Metabolic engineering of yeasts by heterologous enzyme production for degradation of cellulose and hemicellulose from biomass: a perspective. Front Microbiol. 5:1–11. https://doi.org/10.3389/fmicb.2014.00174
Kumar L, Rathore V, Srivastava H (2001) 14C-[lignin]-lignocellulose biodegradation by bacteria isolated from polluted soil. Indian J Exp Biol. 39:584–589
Li C, Zang H, Yu Q, Lv T, Cheng Y, Cheng X, Liu K, Liu W, Xu P, Lan C (2016a) Biodegradation of chlorimuron-ethyl and the associated degradation pathway by Rhodococcus sp. D310–1. Environ Sci Pollut Res Int. 23:8794–8805. https://doi.org/10.1007/s11356-015-5976-3
Li D, Feng L, Liu K, Cheng Y, Hou N, Li C (2016b) Optimization of cold-active CMCase production by psychrotrophic Sphingomonas sp. FLX-7 from the cold region of China. Cellulose 23:1335–1347. https://doi.org/10.1007/s10570-016-0859-4
Liang YL, Zhang Z, Wu M, Wu Y, Feng JX (2014) Isolation, screening, and identification of cellulolytic bacteria from natural reserves in the subtropical region of China and optimization of cellulase production by Paenibacillus terrae ME27-1. Biomed Res Int. 2014:1–13. https://doi.org/10.1155/2014/512497
Liu Y, Hu T, Wu Z, Zeng G, Huang D, Shen Y, He X, Lai M, He Y (2014) Study on biodegradation process of lignin by FTIR and DSC. Environ Sci Pollut Res Int. 21:14004–14013. https://doi.org/10.1007/s11356-014-3342-5
Liu J, Sidhu SS, Ming LW, Tonnis B, Habteselassie M, Mao J, Huang Q (2015) Evaluation of various fungal pretreatment of switchgrass for enhanced saccharification and simultaneous enzyme production. J Clean Prod. 104:480–488. https://doi.org/10.1016/j.jclepro.2015.04.094
Margesin R, Moertelmaier C, Mair J (2013) Low-temperature biodegradation of petroleum hydrocarbons (n-alkanes, phenol, anthracene, pyrene) by four actinobacterial strains. Int Biodeterior Biodegrad. 84:185–191. https://doi.org/10.1016/j.ibiod.2012.05.004
Masai E, Katayama Y, Fukuda M (2007a) Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. J Agric Chem Soc Jpn. 71:1–15. https://doi.org/10.1271/bbb.60437
Masai E, Yamamoto Y, Inoue T, Takamura K, Hara H, Kasai D, Katayama Y, Fukuda M (2007b) Characterization of ligV essential for catabolism of vanillin by Sphingomonas paucimobilis SYK-6. Biosci Biotechnol Biochem. 71:2487. https://doi.org/10.1271/bbb.70267
Miranda-Ríos JA, Ramírez-Trujillo JA, Nova-Franco B, Lozano-Aguirre Beltrán LF, Iturriaga G, Suárez-Rodríguez R (2015) Draft genome sequence of Arthrobacter chlorophenolicus strain Mor30.16, isolated from the bean rhizosphere. Genome Announc. 3:1–2. https://doi.org/10.1128/genomeA.00360-15
Mongodin EF, Shapir N, Daugherty SC, DeBoy RT, Emerson JB, Shvartzbeyn A, Radune D, Vamathevan J, Riggs F, Grinberg V, Khouri H, Wackett LP, Nelson KE, Sadowsky MJ (2006) Secrets of soil survival revealed by the genome sequence of Arthrobacter aurescens TC1. PLoS Genet. 2:e214. https://doi.org/10.1371/journal.pgen.0020214
Niewerth H, Schuldes J, Parschat K, Kiefer P, Vorholt JA, Daniel R, Fetzner S (2012) Complete genome sequence and metabolic potential of the quinaldine-degrading bacterium Arthrobacter sp. Rue61a. Bmc Genom. 13:534–534. https://doi.org/10.1186/1471-2164-13-534
Panagiotopoulos IA, Lignos GD, Bakker RR, Koukios EG (2012) Effect of low severity dilute-acid pretreatment of barley straw and decreased enzyme loading hydrolysis on the production of fermentable substrates and the release of inhibitory compounds. J Clean Prod. 32:45–51. https://doi.org/10.1016/j.jclepro.2012.03.019
Perestelo F, Carnicero A, De lFG (1996) Short communication: isolation of a bacterium capable of limited degradation of industrial and labelled, natural and synthetic lignins. World J Microbiol Biotechnol. 12:111–112. https://doi.org/10.1007/bf00327817
Priefert H, Rabenhorst J, Steinbüchel A (1997) Molecular characterization of genes of Pseudomonas sp. strain HR199 involved in bioconversion of vanillin to protocatechuate. J Bacteriol. 179:2595–2607. https://doi.org/10.1128/jb.179.8.2595-2607.1997
Raj A, Chandra R, Reddy MMK, Purohit HJ, Kapley A (2006) Biodegradation of kraft lignin by a newly isolated bacterial strain, Aneurinibacillus aneurinilyticus from the sludge of a pulp paper mill. World J Microbiol Biotechnol. 23:793–799. https://doi.org/10.1007/s11274-006-9299-x
Raj A, Krishna Reddy MM, Chandra R (2007) Identification of low molecular weight aromatic compounds by gas chromatography–mass spectrometry (GC–MS) from kraft lignin degradation by three Bacillus sp. Int Biodeter Biodegr. 59:292–296. https://doi.org/10.1016/j.ibiod.2006.09.006
Ramachandra M, Crawford DL, Hertel G (1988) Characterization of an extracellular lignin peroxidase of the lignocellulolytic actinomycete Streptomyces viridosporus. Appl Environ Microbiol. 54:3057
Rouches E, Herpoëlgimbert I, Steyer JP, Carrere H (2016) Improvement of anaerobic degradation by white-rot fungi pretreatment of lignocellulosic biomass: a review. Renew Sustain Energy Rev. 59:179–198. https://doi.org/10.1016/j.rser.2015.12.317
Ruan Z, Zhou S, Jiang S, Sun L, Zhai Y, Wang Y, Chen C, Zhao B (2013) Isolation and characterization of a novel cinosulfuron degrading Kurthia sp. from a methanogenic microbial consortium. Bioresour Technol. 147:477–483. https://doi.org/10.1016/j.biortech.2013.08.017
Sainsbury PD, Hardiman EM, Ahmad M, Otani H, Seghezzi N, Eltis LD, Bugg TD (2013) Breaking down lignin to high-value chemicals: the conversion of lignocellulose to vanillin in a gene deletion mutant of Rhodococcus jostii RHA1. ACS Chem Biol. 8:2151–2156. https://doi.org/10.1021/cb400505a
Sathish L, Pavithra N, Ananda K (2012) Antimicrobial activity and biodegrading enzymes of endophytic fungi from eucalyptus. Int J Pharm Sci Res. 3:2574–2583
See-Too WS, Ee R, Lim YL, Convey P, Pearce DA, Mohidin TBM, Yin WF, Chan KG (2017) Complete genome of Arthrobacter alpinus strain R3.8, bioremediation potential unraveled with genomic analysis. Stand Genomic Sci. 12:52. doi:10.1186/s40793–017–0264–0
Shi Y, Chai L, Tang C, Yang Z, Zhang H, Chen R, Chen Y, Zheng Y (2013) Characterization and genomic analysis of kraft lignin biodegradation by the beta-proteobacterium Cupriavidus basilensis B-8. Biotechnol Biofuels. 6:1–1. https://doi.org/10.1186/1754-6834-6-1
Song J, Gu J, Yi Z, Wei W, Wang H, Ruan Z, Shi Y, Yan Y (2013) Biodegradation of nicosulfuron by a Talaromyces flavus LZM1. Bioresour Technol. 140:243–248. https://doi.org/10.1016/j.biortech.2013.02.086
Struvay C, Feller G (2012) Optimization to Low Temperature Activity in Psychrophilic Enzymes. Int J Mol Sci. 13:11643–11665. https://doi.org/10.3390/ijms130911643
Sun Y, Qiu X, Liu Y (2013) Chemical reactivity of alkali lignin modified with laccase. Biomass Bioenergy. 55:198–204. https://doi.org/10.1016/j.biombioe.2013.02.006
Tang M, Zhang F, Yao S, Liu Y, Chen J (2015) Application of Pseudomonas flava WD-3 for sewage treatment in constructed wetland in winter. Environ Technol. 36:1205–1211. https://doi.org/10.1080/21622515.2014.983183
Tanvi CS, Dhanker R, Devi S, Goyal S (2018) Optimization of Physical and Nutritional Factors for Enhanced Production of Lignocellulolytic Enzymes by Aspergillus terreus FJAT-31011 under Submerged Conditions. Int J Curr Microbiol Appl Sci. 7:150–162. https://doi.org/10.20546/ijcmas.2018.707.019
Tien M, Kirk TK (1988) Lignin peroxidase of Phanerochaete chrysosporium. Methods Enzymol. 161:238–249. https://doi.org/10.1016/0076-6879(88)61025-1
Ueda M, Goto T, Nakazawa M, Miyatake K, Sakaguchi M, Inouye K (2010) A novel cold-adapted cellulase complex from Eisenia foetida: characterization of a multienzyme complex with carboxymethylcellulase, beta-glucosidase, beta-1,3 glucanase, and beta-xylosidase. Comp Biochem Physiol B Biochem Mol Biol. 157:26–32. https://doi.org/10.1016/j.cbpb.2010.04.014
Wang D, Lin Y, Du W, Liang J, Ning Y (2013) Optimization and characterization of lignosulfonate biodegradation process by a bacterial strain Sphingobacterium sp. HY-H. Int Biodeterior Biodegrad. 85:365–371. https://doi.org/10.1016/j.ibiod.2013.06.032
Wang J, Wang X, Xu M, Feng G, Zhang W, Ca Lu (2015a) Crop yield and soil organic matter after long-term straw return to soil in China. Nutr Cycl Agroecosys. 102:371–381. https://doi.org/10.1007/s10705-015-9710-9
Wang P, Chang J, Yin Q, Wang E, Zhu Q, Song A, Lu F (2015b) Effects of thermo-chemical pretreatment plus microbial fermentation and enzymatic hydrolysis on saccharification and lignocellulose degradation of corn straw. Bioresour Technol. 194:165–171. https://doi.org/10.1016/j.biortech.2015.07.012
Xiong M, Li C, Pan J, Cheng X, Xi C (2011) Isolation and characterization of Rhodococcus sp.BX2 capable of degrading bensulfuron-methyl. Afr J Microbiol Res. 5:4296–4302. https://doi.org/10.5897/ajmr11.051
Yadav M, Paritosh K, Pareek N, Vivekanand V (2019) Coupled treatment of lignocellulosic agricultural residues for augmented biomethanation. J Clean Prod. 213:75–88. https://doi.org/10.1016/j.jclepro.2018.12.142
Yoon JH, Oh HM, Yoon BD, Kang KH, Park YH (2003) Paenibacillus kribbensis sp. nov. and Paenibacillus terrae sp. nov., bioflocculants for efficient harvesting of algal cells. Int J Syst Evol Microbiol. 53:295–301. https://doi.org/10.1099/ijs.0.02108-0
Zang H, Yu Q, Lv T, Cheng Y, Feng L, Cheng X, Li C (2016) Insights into the degradation of chlorimuron-ethyl by Stenotrophomonas maltophilia D310–3. Chemosphere 144:176–184. https://doi.org/10.1016/j.chemosphere.2015.08.073
Zhang R, Zhou J, Gao Y, Guan Y, Li J, Tang X, Xu B, Ding J, Huang Z (2015) Molecular and biochemical characterizations of a new low-temperature active mannanase. Folia Microbiol. 60:483–492. https://doi.org/10.1007/s12223-015-0391-1
Zheng Y, Chai L-y, Yang Z-h, Zhang H, Chen Y-h (2013) Characterization of a newly isolated Bacterium Pandoraea sp. B-6 capable of degrading kraft lignin. J Cent South Univ. 20:757–763. https://doi.org/10.1007/s11771-013-1545-4
Acknowledgments
We would like to acknowledge “Northeast Agricultural University/Key Laboratory of Swine Facilities Engineering, Ministry of Agriculture, People’s Republic of China” for excellent technical assistance. This research was supported by the National Natural Science Foundation of China (Grant Numbers 41771559).
Author information
Authors and Affiliations
Contributions
CYL and HLZ designed the whole scheme of the study and conducted the experiments. CJ, YC, and XC performed experiments and XHS and JMW analyzed data. CJ and CYL wrote the manuscript, and YW and YTZ helped to revise. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, C., Cheng, Y., Zang, H. et al. Biodegradation of lignin and the associated degradation pathway by psychrotrophic Arthrobacter sp. C2 from the cold region of China. Cellulose 27, 1423–1440 (2020). https://doi.org/10.1007/s10570-019-02858-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-019-02858-3