Abstract
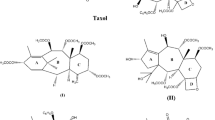
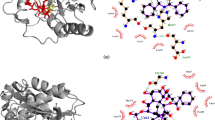
Taxanes (paclitaxel, docetaxel, cabazitaxel) are anticancer drugs as microtubule inhibitors. Following our previous studies on paclitaxel and docetaxel, in this work, we examine cabazitaxel and compare these three taxenes. The binding interaction of three taxanes with various β-tubulin isotypes is studied by homology modeling, molecular docking, and molecular dynamics simulations. The results show that the effects of docetaxel on βI-tubulin (− 29.5 kcal/mol) and of paclitaxel on βIIa-tubulin (− 25.5 kcal/mol) are much stronger than their effects on βIII-tubulin (− 17.8 kcal/mol and − 8.6 kcal/mol, respectively). However, the effect of cabazitaxel on βIII-tubulin (− 23.0 kcal/mol) is comparable with that on βI-tubulin (− 24.0 kcal/mol) and βIIa-tubulin (− 25.9 kcal/mol), consistent with the fact that overexpression of βIII-tubulin increases the drug resistance to paclitaxel and docetaxel, but has little influence for cabazitaxel. This theoretical research supports the use of cabazitaxel for patients who are resistant to the action of paclitaxel and docetaxel.







Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J. Clin. 68(1):7–30. https://doi.org/10.3322/caac.21442
Yan X, Han R, Zhou J, Yu H, Yang J, Wu M (2016) Incidence, mortality and survival of female breast cancer during 2003–2011 in Jiangsu province, China. Chin J Cancer Res 28(3):321–329. https://doi.org/10.21147/j.issn.1000-9604.2016.03.06
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016) Cancer statistics in China, 2015. CA Cancer J. Clin. 66(2):115–132. https://doi.org/10.3322/caac.21338
Zeng H, Chen W, Zheng R, Zhang S, Ji JS, Zou X, Xia C, Sun K, Yang Z, Li H, Wang N, Han R, Liu S, Li H, Mu H, He Y, Xu Y, Fu Z, Zhou Y, Jiang J, Yang Y, Chen J, Wei K, Fan D, Wang J, Fu F, Zhao D, Song G, Chen J, Jiang C, Zhou X, Gu X, Jin F, Li Q, Li Y, Wu T, Yan C, Dong J, Hua Z, Baade P, Bray F, Jemal A, Yu XQ, He J (2018) Changing cancer survival in China during 2003–15: a pooled analysis of 17 population-based cancer registries. Lancet Glob. Health 6(5):e555–e567. https://doi.org/10.1016/S2214-109X(18)30127-X
Chen JG, Chen HZ, Zhu J, Yang YL, Zhang YH, Huang PX, Chen YS, Zhu CY, Yang LP, Shen K, Qiang FL, Wang GR (2018) Cancer survival in patients from a hospital-based cancer registry, China. J. Cancer 9(5):851–860. https://doi.org/10.7150/jca.23039
Janke C, Bulinski JC (2011) Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat Rev Mol Cell Biol 12(12):773–786. https://doi.org/10.1038/nrm3227
Conde C, Caceres A (2009) Microtubule assembly, organization and dynamics in axons and dendrites. Nat. Rev. Neurosci. 10(5):319–332. https://doi.org/10.1038/nrn2631
Perez EA (2009) Microtubule inhibitors: differentiating tubulin-inhibiting agents based on mechanisms of action, clinical activity, and resistance. Mol. Cancer Ther. 8(8):2086–2095. https://doi.org/10.1158/1535-7163.MCT-09-0366
Giannakakou P, Gussio R, Nogales E, Downing KH, Zaharevitz D, Bollbuck B, Poy G, Sackett D, Nicolaou KC, Fojo T (2000) A common pharmacophore for epothilone and taxanes: molecular basis for drug resistance conferred by tubulin mutations in human cancer cells. Proc. Natl. Acad. Sci. 97(6):2904–2909. https://doi.org/10.1073/pnas.040546297
Kruczynski A, Barret JM, Etievant C, Colpaert F, Fahy J, Hill BT (1998) Antimitotic and tubulin-interacting properties of vinflunine, a novel fluorinated vinca alkaloid. Biochem. Pharmacol. 55(5):635–648. https://doi.org/10.1016/s0006-2952(97)00505-4
Lu Y, Chen J, Xiao M, Li W, Miller DD (2012) An overview of tubulin inhibitors that interact with the colchicine binding site. Pharm. Res. 29(11):2943–2971. https://doi.org/10.1007/s11095-012-0828-z
Jordan MA, Wilson L (2004) Microtubules as a target for anticancer drugs. Nat. Rev. Cancer 4(4):253–265. https://doi.org/10.1038/nr1317
Ravelli RB, Gigant B, Curmi PA, Jourdain I, Lachkar S, Sobel A, Knossow M (2004) Insight into tubulin regulation from a complex with colchicine and a stathmin-like domain. Nature 428(6979):198–202. https://doi.org/10.1038/nature02393
Mukhtar E, Adhami VM, Mukhtar H (2014) Targeting microtubules by natural agents for cancer therapy. Mol. Cancer Ther. 13(2):275–284. https://doi.org/10.1158/1535-7163.mct-13-0791
Prota AE, Bargsten K, Zurwerra D, Field JJ, Diaz JF, Altmann KH, Steinmetz MO (2013) Molecular mechanism of action of microtubule-stabilizing anticancer agents. Science 339(6119):587–590. https://doi.org/10.1126/science.1230582
Field JJ, Diaz JF, Miller JH (2013) The binding sites of microtubule-stabilizing agents. Chem. Biol. 20(3):301–315. https://doi.org/10.1016/j.chembiol.2013.01.014
Blagosklonny MV, Fojo T (1999) Molecular effects of paclitaxel: myths and reality (a critical review). Int J Cancer 83(2):151–156. https://doi.org/10.1002/(sici)1097-0215(19991008)83:23.0.co;2-5
Lowe J, Li H, Downing KH, Nogales E (2001) Refined structure of αβ-tubulin at 3.5 Å resolution. J. Mol. Biol. 313(5):1045–1057. https://doi.org/10.1006/jmbi.2001.5077
Mitra A, Sept D (2008) Taxol allosterically alters the dynamics of the tubulin dimer and increases the flexibility of microtubules. Biophys. J. 95(7):3252–3258. https://doi.org/10.1529/biophysj.108.133884
Natarajan K, Senapati S (2012) Understanding the basis of drug resistance of the mutants of αβ-tubulin dimer via molecular dynamics simulations. PLoS One 7(8):e42351. https://doi.org/10.1371/journal.pone.0042351
Alushin GM, Lander GC, Kellogg EH, Zhang R, Baker D, Nogales E (2014) High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157(5):1117–1129. https://doi.org/10.1016/j.cell.2014.03.053
Kellogg EH, Hejab NMA, Howes S, Northcote P, Miller JH, Diaz JF, Downing KH, Nogales E (2017) Insights into the distinct mechanisms of action of taxane and non-taxane microtubule stabilizers from cryo-EM structures. J. Mol. Biol. 429(5):633–646. https://doi.org/10.1016/j.jmb.2017.01.001
Cao YN, Zheng LL, Wang D, Liang XX, Gao F, Zhou XL (2018) Recent advances in microtubule-stabilizing agents. Eur. J. Med. Chem. 143:806–828. https://doi.org/10.1016/j.ejmech.2017.11.062
Sun C, Zhu L, Zhang C, Song C, Wang C, Zhang M, Xie Y, Schaefer HF (2018) Conformers, properties, and docking mechanism of the anticancer drug docetaxel: DFT and molecular dynamics studies. J. Comput. Chem. 39(15):889–900. https://doi.org/10.1002/jcc.25165
Shah M, Dauffenbach L, Kerfoot CA, Cohen P, Culver KW, Sharma S, Wartmann M (2003) A comparative study of two novel, non-taxane microtubule stabilizing agents (XAA296A and EPO906A) utilizing the extreme drug resistance (EDR) assay in paclitaxel-sensitive or paclitaxel-resistant breast and ovarian cancers. Clin Cancer Res 9(16):6133s
Sgadari C, Toschi E, Palladino C, Barillari G, Carlei D, Cereseto A, Ciccolella C, Yarchoan R, Monini P, Sturzl M, Ensoli B (2000) Mechanism of paclitaxel activity in Kaposi’s sarcoma. J. Immunol. 165(1):509–517. https://doi.org/10.4049/jimmunol.165.1.509
Seoud MAF, Shamseddine A, Khalil AM, Charafeddine M, Geara FB (2006) Randomized trial comparing concurrent radiotherapy with cisplatin compared with paclitaxel in cervical cancer. Obstet Gynecol 107(4):43S. https://doi.org/10.1097/00006250-200604001-00101
Von Hoff DD, Goldstein D, Renschler MF (2014) Albumin-bound paclitaxel plus gemcitabine in pancreatic cancer REPLY. N. Engl. J. Med. 370(5):479–480
McGrogan BT, Gilmartin B, Carney DN, McCann A (2008) Taxanes, microtubules and chemoresistant breast cancer. Biochim. Biophys. Acta 1785(2):96–132. https://doi.org/10.1016/j.bbcan.2007.10.004
Joensuu H, Bono P, Kataja V, Alanko T, Kokko R, Asola R, Utriainen T, Turpeenniemi-Hujanen T, Jyrkkio S, Moykkynen K, Helle L, Ingalsuo S, Pajunen M, Huusko M, Salminen T, Auvinen P, Leinonen H, Leinonen M, Isola J, Kellokumpu-Lehtinen PL (2009) Fluorouracil, epirubicin, and cyclophosphamide with either docetaxel or vinorelbine, with or without trastuzumab, as adjuvant treatments of breast cancer: final results of the FinHer trial. J. Clin. Oncol. 27(34):5685–5692. https://doi.org/10.1200/JCO.2008.21.4577
Gernone A, Pagliarulo A, Calderoni G, Pagliarulo V (2011) Elderly patients with metastatic castrate-resistant prostate cancer (mCRPC): safety and efficacy of docetaxel retreatment. J Clin Oncol 29(7):161. https://doi.org/10.1200/jco.2011.29.7_suppl.161
Urano N, Fujiwara Y, Doki Y, Kim SJ, Miyoshi Y, Noguchi S, Miyata H, Takiguchi S, Yasuda T, Yano M, Monden M (2006) Clinical significance of class III β-tubulin expression and its predictive value for resistance to docetaxel-based chemotherapy in gastric cancer. Int. J. Oncol. 28(2):375–381
Galsky MD, Dritselis A, Kirkpatrick P, Oh WK (2010) Cabazitaxel. Nat. Rev. Drug Discov. 9(9):677–678. https://doi.org/10.1038/nrd3254
Duran GE, Wang YC, Francisco EB, Rose JC, Martinez FJ, Coller J, Brassard D, Vrignaud P, Sikic BI (2015) Mechanisms of resistance to cabazitaxel. Mol. Cancer Ther. 14(1):193–201. https://doi.org/10.1158/1535-7163.MCT-14-0155
Paller CJ, Antonarakis ES (2011) Cabazitaxel: a novel second-line treatment for metastatic castration-resistant prostate cancer. Drug Des Devel Ther 5:117–124. https://doi.org/10.2147/DDDT.S13029
Pean E, Demolis P, Moreau A, Hemmings RJ, O'Connor D, Brown D, Shepard T, Abadie E, Pignatti F (2012) The European Medicines Agency Review of Cabazitaxel (Jevtana®) for the treatment of hormone-refractory metastatic prostate cancer: summary of the Scientific Assessment of the Committee for Medicinal Products for Human Use. Oncologist 17(4):543–549. https://doi.org/10.1634/theoncologist.2011-0364
Luduena RF (1998) Multiple forms of tubulin: different gene products and covalent modifications. Int. Rev. Cytol. 178:207–275
Santoshi S, Naik PK (2014) Molecular insight of isotypes specific β-tubulin interaction of tubulin heterodimer with noscapinoids. J. Comput. Aided Mol. Des. 28(7):751–763. https://doi.org/10.1007/s10822-014-9756-9
Derry WB, Wilson L, Khan IA, Luduena RF, Jordan MA (1997) Taxol differentially modulates the dynamics of microtubules assembled from unfractionated and purified β-tubulin isotypes. Biochemistry 36(12):3554–3562. https://doi.org/10.1021/bi962724m
Yang CH, Yap EH, Xiao H, Fiser A, Horwitz SB (2016) 2-(m-Azidobenzoyl)taxol binds differentially to distinct β-tubulin isotypes. Proc. Natl. Acad. Sci. 113(40):11294–11299. https://doi.org/10.1073/pnas.1613286113
Hari M, Yang H, Zeng C, Canizales M, Cabral F (2003) Expression of class IIIβ-tubulin reduces microtubule assembly and confers resistance to paclitaxel. Cell Motil. Cytoskeleton 56(1):45–56. https://doi.org/10.1002/cm.10132
Seve P, Isaac S, Tredan O, Souquet PJ, Pacheco Y, Perol M, Lafanechere L, Penet A, Peiller EL, Dumontet C (2005) Expression of class III β-tubulin is predictive of patient outcome in patients with non-small cell lung cancer receiving vinorelbine-based chemotherapy. Clin. Cancer Res. 11(15):5481–5486. https://doi.org/10.1158/1078-0432.CCR-05-0285
Ferrandina G, Martinelli E, Zannoni GF, Distefano M, Paglia A, Ferlini C, Scambia G (2007) Expression of class III β-tubulin in cervical cancer patients administered preoperative radiochemotherapy: correlation with response to treatment and clinical outcome. Gynecol. Oncol. 104(2):326–330. https://doi.org/10.1016/j.ygyno.2006.08.046
Sève P, Dumontet C (2008) Is class III β-tubulin a predictive factor in patients receiving tubulin-binding agents? Lancet Oncol 9(2):168–175. https://doi.org/10.1016/s1470-2045(08)70029-9
Katsetos CD, Herman MM, Mork SJ (2003) Class III β-tubulin in human development and cancer. Cell Motil. Cytoskeleton 55(2):77–96. https://doi.org/10.1002/cm.10116
Ferrandina G, Zannoni GF, Martinelli E, Paglia A, Gallotta V, Mozzetti S, Scambia G, Ferlini C (2006) Class III β-tubulin overexpression is a marker of poor clinical outcome in advanced ovarian cancer patients. Clin. Cancer Res. 12(9):2774–2779. https://doi.org/10.1158/1078-0432.ccr-05-2715
Magnani M, Ortuso F, Soro S, Alcaro S, Tramontano A, Botta M (2006) The βI/βIII-tubulin isoforms and their complexes with antimitotic agents. FEBS J. 273(14):3301–3310. https://doi.org/10.1111/j.1742-4658.2006.05340.x
Akasaka K, Maesawa C, Shibazaki M, Maeda F, Takahashi K, Akasaka T, Masuda T (2009) Loss of class III β-tubulin induced by histone deacetylation is associated with chemosensitivity to paclitaxel in malignant melanoma cells. J. Investig. Dermatol. 129(6):1516–1526. https://doi.org/10.1038/jid.2008.406
Smiyun G, Azarenko O, Miller H, Rifkind A, LaPointe NE, Wilson L, Jordan MA (2017) βIII-tubulin enhances efficacy of cabazitaxel as compared with docetaxel. Cancer Chemother. Pharmacol. 80(1):151–164. https://doi.org/10.1007/s00280-017-3345-2
Yusuf RZ, Duan Z, Lamendola DE, Penson RT, Seiden MV (2003) Paclitaxel resistance: molecular mechanisms and pharmacologic manipulation. Curr. Cancer Drug Targets 3(1):1–19. https://doi.org/10.2174/1568009033333754
Ferlini C, Raspaglio G, Cicchillitti L, Mozzetti S, Prislei S, Bartollino S, Scambia G (2007) Looking at drug resistance mechanisms for microtubule interacting drugs: does TUBB3 work? Curr. Cancer Drug Targets 7(8):704–712. https://doi.org/10.2174/156800907783220453
Sissung TM, Baum CE, Deeken J, Price DK, Aragon-Ching J, Steinberg SM, Dahut W, Sparreboom A, Figg WD (2008) ABCB1 genetic variation influences the toxicity and clinical outcome of patients with androgen-independent prostate cancer treated with docetaxel. Clin. Cancer Res. 14(14):4543–4549. https://doi.org/10.1158/1078-0432.ccr-07-4230
Kavallaris M (2010) Microtubules and resistance to tubulin-binding agents. Nat. Rev. Cancer 10(3):194–204. https://doi.org/10.1038/nrc2803
Kulkarni SA, Feng SS (2011) Effects of surface modification on delivery efficiency of biodegradable nanoparticles across the blood-brain barrier. Nanomedicine 6(2):377–394. https://doi.org/10.2217/nnm.10.131
Culine S (2018) Refining the use of cabazitaxel in metastatic castrate-resistant prostate cancer. Eur. J. Cancer 97:30–32. https://doi.org/10.1016/j.ejca.2018.04.002
Zhang ML, Song C, Yao Z, Ji Q (2012) Theoretical studies of the structure and properties of anticancer drug taxol. Curr Org Chem 16(19):2321–2331. https://doi.org/10.2174/138527212803520281
Durrant JD, McCammon JA (2011) Molecular dynamics simulations and drug discovery. BMC Biol. 9:71. https://doi.org/10.1186/1741-7007-9-71
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Petersson GA, Nakatsuji H, Li X, Caricato M, Marenich AV, Bloino J, Janesko BG, Gomperts R, Mennucci B, Hratchian HP, Ortiz JV, Izmaylov AF, Sonnenberg JL, Williams, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski VG, Gao J, Rega N, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery Jr. JA, Peralta JE, Ogliaro F, Bearpark MJ, Heyd JJ, Brothers EN, Kudin KN, Staroverov VN, Keith TA, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Adamo C, Cammi R, Ochterski JW, Martin RL, Morokuma K, Farkas O, Foresman JB, Fox DJ (2016) Gaussian 16 Rev. B.01. Wallingford, CT
Cavasotto CN, Phatak SS (2009) Homology modeling in drug discovery: current trends and applications. Drug Discov. Today 14(13–14):676–683. https://doi.org/10.1016/j.drudis.2009.04.006
The UniProt C (2017) UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45(D1):D158–D169. https://doi.org/10.1093/nar/gkw1099
McWilliam H, Li W, Uludag M, Squizzato S, Park YM, Buso N, Cowley AP, Lopez R (2013) Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 41:W597–W600. https://doi.org/10.1093/nar/gkt376
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30:S162–S173. https://doi.org/10.1002/elps.200900140
Benkert P, Biasini M, Schwede T (2011) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics (Oxford, England) 27(3):343–350. https://doi.org/10.1093/bioinformatics/btq662
Bienert S, Waterhouse A, de Beer TA, Tauriello G, Studer G, Bordoli L, Schwede T (2017) The SWISS-MODEL repository-new features and functionality. Nucleic Acids Res. 45(D1):D313–D319. https://doi.org/10.1093/nar/gkw1132
Bertoni M, Kiefer F, Biasini M, Bordoli L, Schwede T (2017) Modeling protein quaternary structure of homo- and hetero-oligomers beyond binary interactions by homology. Sci. Rep. 7(1):10480. https://doi.org/10.1038/s41598-017-09654-8
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46(W1):W296–W303. https://doi.org/10.1093/nar/gky427
Gilson MK, Zhou HX (2007) Calculation of protein-ligand binding affinities. Annu. Rev. Biophys. 36:21–42. https://doi.org/10.1146/annurev.biophys.36.040306.132550
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J. Comput. Chem. 26(16):1701–1718. https://doi.org/10.1002/jcc.20291
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J. Comput. Chem. 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Leontyev IV, Stuchebrukhov AA (2010) Electronic polarizability and the effective pair potentials of water. J. Chem. Theory Comput. 6(10):3153–3161. https://doi.org/10.1021/ct1002048
Hess B (2008) P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4(1):116–122. https://doi.org/10.1021/ct700200b
Mozzetti S, Ferlini C, Concolino P, Filippetti F, Raspaglio G, Prislei S, Gallo D, Martinelli E, Ranelletti FO, Ferrandina G, Scambia G (2005) Class III β-tubulin overexpression is a prominent mechanism of paclitaxel resistance in ovarian cancer patients. Clin. Cancer Res. 11(1):298–305
Seve P, Mackey J, Isaac S, Tredan O, Souquet PJ, Perol M, Lai R, Voloch A, Dumontet C (2005) Class III β-tubulin expression in tumor cells predicts response and outcome in patients with non-small cell lung cancer receiving paclitaxel. Mol. Cancer Ther. 4(12):2001–2007. https://doi.org/10.1158/1535-7163.MCT-05-0244
Gan PP, Pasquier E, Kavallaris M (2007) Class III β-tubulin mediates sensitivity to chemotherapeutic drugs in non-small cell lung cancer. Cancer Res. 67(19):9356–9363. https://doi.org/10.1158/0008-5472.CAN-07-0509
Guo J, Wang X, Sun H, Liu H, Yao X (2012) The molecular basis of IGF-II/IGF2R recognition: a combined molecular dynamics simulation, free-energy calculation and computational alanine scanning study. J. Mol. Model. 18(4):1421–1430. https://doi.org/10.1007/s00894-011-1159-4
Jing X, Xiaoqiang H, Yushan Z (2019) Using molecular dynamics simulations to evaluate active designs of cephradine hydrolase by molecular mechanics/Poisson – Boltzmann surface area and molecular mechanics/generalized born surface area methods. RSC Adv. 9:13868–13877. https://doi.org/10.1039/c9ra02406a
Kumari R, Kumar R, Open Source Drug Discovery C, Lynn A (2014) g_mmpbsa--a GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. Model. 54(7):1951–1962. https://doi.org/10.1021/ci500020m
Stepanian SG, Reva ID, Radchenko ED, Adamowicz L (1998) Conformational behavior of α-alanine. Matrix-isolation infrared and theoretical DFT and ab initio study. J Phys Chem A 102(24):4623–4629. https://doi.org/10.1021/jp973479z
Yu D, Kang W, Hao F, Li Z (2017) Spectroscopic studies and structural elucidation of cabazitaxel. Chinese J Magn Reson 34(2):191–199
Baker D, Sali A (2001) Protein structure prediction and structural genomics. Science 294(5540):93–96. https://doi.org/10.1126/science.1065659
Bowie JU, Luthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253(5016):164–170
Luthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356(6364):83–85. https://doi.org/10.1038/356083a0
Kumbhar BV, Borogaon A, Panda D, Kunwar A (2016) Exploring the origin of differential binding affinities of human tubulin isotypes αβII, αβIII and αβIV for DAMA-colchicine using homology modelling, molecular docking and molecular dynamics simulations. PLoS One 11(5):e0156048. https://doi.org/10.1371/journal.pone.0156048
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grants No. 10904111, 11604238, and 21772146), the Tianjin Natural Science Foundation (11JCYBJC14500), the Science & Technology Development Fund of Tianjin Education Commission for Higher Education (2019KJ175), and the China Postdoctoral Science Foundation (20100470792). We appreciate the supply of the computing resources by USTC. The research at the University of Georgia was supported by the National Science Foundation (Grant CHE-1661604). The visit of MZ to the Center for Computational Quantum Chemistry, the University of Georgia was very helpful.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 11727 kb)
Rights and permissions
About this article
Cite this article
Zhu, L., Zhang, C., Lü, X. et al. Binding modes of cabazitaxel with the different human β-tubulin isotypes: DFT and MD studies. J Mol Model 26, 162 (2020). https://doi.org/10.1007/s00894-020-04400-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-020-04400-w