Abstract
As the enzyme nucleoside hydrolase (NH) is widely found in nature but has not yet been detected in mammals, it is considered an ideal target in the development of chemotherapy against parasitic diseases and bacterial infections like anthrax. Considering the risk that this biological warfare agent represents nowadays, the search for new drugs and new molecular targets in the development of chemotherapy against anthrax is imperative. On this basis, we performed docking studies of six known NH inhibitors at the active site of NH from Bacillus anthracis (BaNH). Subsequently, molecular dynamics (MD) simulations of these compounds inside BaNH were carried out in order to complement the docking studies and select the most promising compounds as leads for the design of potential BaNH inhibitors. Most of the docking and MD results obtained agreed well with each other and showed good correlation with experimental data.

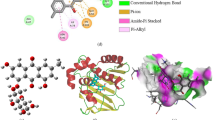
Interactions of compound 6 in the active sites of TvNH and BaNH


















Similar content being viewed by others
References
Goldman DL, Arturo C (2008) Anthrax-associated shock. Front Biosc 13:4009–4014
Sterne M (1967) Distribution and economic importance of anthrax. Fed Pro 26:1493–1495
Dixon TC, Meselson M, Guillemin J, Hanna PC (1999) Anthrax. N Eng J Med 341:815–826
Turnbull PC (1996) B. Anthrax is alive and well PHLS. Microbiol Digest 9:103–106
Lindler LE, Lebeda FJ, Korch GW (2005) Biological weapons defense: infectious diseases and counter bioterrorism. Humana, Totowa
Parkin DW, Schramm VL (1995) Binding modes for substrate and a proposed transition—state analogue of protozoan nucleoside hydrolase. Biochemistry 34:13961–13966
Gopaul DN, Meyer SL, Sacchettini JC, Schramm VL (1996) Inosine-uridine nucleoside hydrolase from crithidia fasciculate: genetic characterization, crystallization, and identification of histidine 241 as a catalytic site residue. Biochemistry 35:5963–5970
Degano M, Almo SC, Sacchettini JC, Schramm VL (1998) Trypanossomal nucleoside hydrolase. A novel mechanism from the structure with a transition-state inhibitor. Biochemistry 37:6277–6285
Shi W, Schramm VL, Almo SC (1999) Nucleoside hydrolase from Leishmania major: cloning, expression, catalytic properties, transition state inhibitors, and the 2.5-Å crystal structure. Biol Chem 274:21114–21120
Cui L, Rajasekariah GR, Martin SK (2001) A nonspecific nucleoside hydrolase from Leishmania donovani: implications for purine salvage by the parasite. Gene 280:153–162
Santana DM, Borja-cabrera GP, de Souza EP, Sturm NR, Palatnik CB, Campbell DA (2002) Nucleoside hydrolase from Leishmania donovani is an antigen diagnostic for visceral leishmaniasis. Mol Biochem Parasitol 120:315–319
Versées W, Steyaert J (2003) Catalysis by nucleoside hydrolases. Curr Opin Struct Biol 13:731–738
Ogawa J, Takeda S, Xie SX, Hatanaka H, Ashikari T, Amachi T, Shimizu S (2001) Purification, characterisation, and gene cloning of purine nucleosidase from Ochrobactrum anthropi. Appl Environ Microbio 67:1783–1787
Petersen C, Moller LB (2001) The RihA, RihB, and RihC ribonucleoside hydrolases of Escherichia coli. Substrate specificity, gene expression, and regulation. J Biol Chem 276:884–894
Kurtz JE, Exinger F, Erbs P, Jund R (2002) The URH1 uridine ribohydrolase of Saccharomyces cerevisae. Curr Genet 41:132–141
Pellé R, Schramm VL, Parkin DW (2001) Molecular cloning and expression of a purine-specific N-ribohydrolase from Trypanosoma brucei brucei: sequence, expression, and molecular analysis. J Biol Chem 273:2118–2126
Ribeiro JM, Valenzuela JG (2003) The salivary purine nucleosidase of the mosquito, Aedes aegypti Insect. Biochem Mol Biol 33:13–22
França TCC, Rocha MRM, Reboredo BM, Rennó MN, Tinoco LW, Villar JDF (2008) Design of inhibitors for nucleoside hydrolase from Leishmania donovani using molecular dynamics studies. J Braz Chem Soc 19:64–73
França TCC, Pascutti PG, Ramalho TC, Villar JDF (2005) A three-dimensional structure of Plasmodium falciparum serine hydroxymethyltransferase in complex with glycine and 5-formyl-tetrahydrofolate. Homology modeling and molecular dynamics. Biophys Chem 115:1–10
França TCC, Wilter A, Pascutti PG, Ramalho TC, Villar JDF (2006) Molecular dynamics of the interaction of Plasmodium falciparum and human serine hydroxymethyltransferase with 5-formyl-6-hydrofolic acid analogues: design of new potential antimalarials. J Braz Chem Soc 17:1383–1392
Gonçalves AS, França TCC, Wilter A, Villar JDF (2006) Molecular dynamics of the interaction of pralidoxime and deazapralidoxime with acetylcholinesterase inhibited by the neurotoxic agent tabun. J Braz Chem Soc 17:968–975
Silva ML, Gonçalves AS, Batista PR, Villar JDF, Pascutti PG, França TCC (2010) Design, docking studies and molecular dynamics of new potential selective inhibitors of Plasmodium falciparum serine hydroxymethyltransferase. Mol Simul 36:5–14
Ramalho TC, França TCC, Rennó MN, Guimarães AP, Cunha EFF, Kuča K (2010) Development of new acetylcholinesterase reactivators: molecular modeling versus in vitro data. Chem Biol Interact 187:436–440
Gonçalves AS, França TCC, Villar JDF, Pascutti PG (2010) Conformational analysis of toxogonine, TMB-4 and HI-6 using PM6 and RM1 methods. J Braz Chem Soc 21:179–184
Gonçalves AS, França TCC, Villar JDF, Pascutti PG (2011) Molecular dynamics simulations and QM/MM studies of the reactivation by 2-PAM of tabun inhibited human acethylcholinesterase. J Braz Chem Soc 22:155–165
Guimarães AP, Oliveira AA, Cunha EFF, Ramalho TC, França TCC (2011) Design of new chemotherapeutics against the deadly anthrax disease. Docking and molecular dynamics studies of inhibitors containing pyrrolidine and riboamidrazone rings on nucleoside hydrolase from Bacillus anthracis. J Biol Struct Dyn 28:455–469
Goeminne A, McNaughton M, Bal G, Surpateanu G, van der Veken P, de Prol S, Versées W, Steyaert J, Haemers A, Augustyns K (2008) Synthesis and biochemical evaluation of guanidino-alkyl-ribitol derivatives as nucleoside hydrolase inhibitors. Eur J Med Chem 43:315–326
Goeminne A, McNaughton M, Bal G, Surpateanu G, van der Veken P, de Prol S, Versées W, Steyaert J, Haemers A, Augustyns K (2008) N-Arylmethyl substituted iminoribitol derivatives as inhibitors of a purine specific nucleoside hydrolase. Bioorg Med Chem 16:6752–6763
Versées W, Decanniere K, Pellé R, van Holsbeve E, Steyaert J (2002) Enzyme-substrate interactions in the purine-specific nucleoside hydrolase from Trypanosoma vivax. J Biol Chem 277:15938–15946
Versées W, Barlow J, Steyaert J (2006) Transition-state complex of the purine-specific nucleoside hydrolase of T.vivax: enzyme conformational changes and implications for catalysis. J Mol Biol 359:331–346
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Guex N, Peitsch MC (1997) SWISS-MODEL and Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Thomsen R, Christensen MH (2006) MolDock: a new technique for high-accuracy molecular docking. J Med Chem 49:3315–3321
Hehre WJ, Deppmeier BJ, Klunzinger PE (1999) PC SPARTAN Pro. Wavefunction, Inc., Irvine
Van der Spoel D, van Buuren AR, Apol E, Meulenhoff PJ, Tieleman DP, Sijbers ALT M, Hess B, Feentra KA, Lindahl E, van Drunen R, Berendsen HJC (2001) GROMACS user manual, version 3.0. University of Groningen, Groningen
Schuettelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein±ligand complexes. Acta Crystallogr D60:1355–1363
Frisch MJ, Trucks GW, Schlegel HB, Scuseria ES, Pople JA et al (2001) Gaussian 98, revision A.11. Gaussian, Inc., Pittsburgh
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Edwards PM (2002) Origin 7.0: scientific graphing and data analysis software. J Chem Inf Comput Sci 42:1270–1271
Koradi R, Billeter M, Wüthrich K (1996) MOLMOL: a program for display and analysis of macromolecular structures. J Mol Graph 14:51–55
Warren D (2002) The PyMOL molecular graphics system. DeLano Scientific, San Carlos
Warren GL, Webster AC, Capelli AM, Clarke B, LaLonde J, Lambert MH, Lindvall M, Nevins N, Semus SF, Senger S, Tedesco G, Wall ID, Woolven JM, Peishoff CE, Head MS (2006) A critical assessment of docking programs and scoring functions. J Med Chem 49:5912–5931
Leach AR, Shoichet BK, Peishoff CE (2006) Prediction of protein–ligand interactions. Docking and scoring: successes and gaps. J Med Chem 49:5851–5855
Kontoyanni M, McClellan LM, Sokol GS (2004) Evaluation of docking performance: comparative data on docking algorithms. J Med Chem 47:558–565
Acknowledgments
The authors wish to thank the Brazilian financial agencies Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo ao Ensino e Pesquisa do Estado do Rio de Janeiro (FAPERJ), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (PAPEMIG) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior/Ministério da Defesa (CAPES/MD) (Edital PRODEFESA 2008) for financial support, and the Military Institute of Engineering for providing the physical infrastructure and working space.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Guimarães, A.P., Oliveira, A.A., da Cunha, E.F.F. et al. Analysis of Bacillus anthracis nucleoside hydrolase via in silico docking with inhibitors and molecular dynamics simulation. J Mol Model 17, 2939–2951 (2011). https://doi.org/10.1007/s00894-011-0968-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-0968-9