Abstract
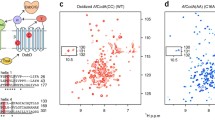
The purified native mercuric reductase (MerA) from Ralstonia metallidurans CH34 contains an N-terminal sequence of 68 amino acids predicted to be homologous to MerP, the periplasmic mercury-binding protein. This MerP-like protein has now been expressed independently. The protein was named MerAa by homology with Ccc2a, the first soluble domain of the copper-transporting ATPase from yeast. Δa has been characterized using a set of biophysical techniques. The binding of mercury was followed using circular dichroism spectroscopy and electrospray mass spectrometry. The two cysteine residues contained in the consensus sequence GMTCXXC are involved in the binding of one mercury atom, with an apparent affinity comparable to that of MerP for the same metal. The metal-binding site is confirmed by NMR chemical shift changes observed between apo- and metal-bound MerAa in solution. NMR shift and NOE data also indicate that only minor structural changes occur upon metal binding. Further NMR investigation of the fold of MerAa using long-range methyl–methyl NOE and backbone residual dipolar coupling data confirm the expected close structural homology with MerP. 15N relaxation data show that MerAa is a globally rigid molecule. An increased backbone mobility was observed for the loop region connecting the first β-strand and the first α-helix and comprising the metal-binding domain. Although significantly reduced, this loop region keeps some conformational flexibility upon metal binding. Altogether, our data suggest a role of MerAa in mercury trafficking.







Similar content being viewed by others

Abbreviations
- CCA:
-
α-cyano-4-hydroxy-trans-cinnamic acid
- CSI:
-
chemical shift index
- HSQC:
-
1H-detected heteronuclear single-quantum coherence
- MerAa:
-
the 68 amino acid N-terminal extension of the mercuric reductase
- NOE:
-
nuclear Overhauser effect
- RDCs:
-
residual dipolar couplings
- TCEP-HCl:
-
tris(2-carboxyethyl)phosphine hydrochloride
References
Silver S, Phung LT (1996) Annu Rev Microbiol 50:753–789
Hobman JL, Brown NL (1997) Metal Ions Biol Syst 34:527–568
Liebert CA, Hall RM, Summers AO (1999) Microbiol Mol Biol Rev 63:507–522
Barkay T, Miller SM, Summers AO (2003) FEMS Microbiol Rev 27:355–384
O’Halloran TV, Walsh CT (1987) Science 235:211–214
Brown NL, Stoyanov JV, Kidd SP, Hobman JL (2003) FEMS Microbiol Rev 27:145–163
Sahlman L, Jonsson B-H (1992) Eur J Biochem 205:375–381
Brown NL, Camakaris J, Lee BT, Williams T, Morby AP, Parkhill J, Rouch DA (1991) J Cell Biochem 46:106–114
Fox B, Walsh CT (1982) J Biol Chem 257:2498–2503
Moore MJ, Walsh CT (1989) Biochemistry 28:1183–1194
Brown NL, Shih Y-C, Leang C, Glendinning KJ, Hobman JL, Wilson JR (2002) Biochem Soc Trans 30:715–718
Sedlmeier R, Altenbuchner J (1992) Mol Gen Genet 236:76–85
Shiering N, Kabsch W, Moore MJ, Distefano MD, Walsh CT, Pai EF (1991) Nature 352:168–172
Steele RA, Opella SJ (1997) Biochemistry 36:6885–6895
Qian H, Sahlman L, Eriksson P-O, Hambraeus C, Edlund U, Sethson I (1998) Biochemistry 37:9316–9322
Opella SJ, Desilva TM, Veglia G (2002) Curr Opin Chem Biol 6:217–223
Arnesano F, Banci L, Bertini I, Ciofi-Baffoni S, Molteni E, Huffman DL, O’Halloran TV (2002) Genome Res 12:255–271
Mergeay M, Monchy S, Vallaeys T, Auquier V, Benotmane A, Bertin P, Taghavi S, Dunn J, van der Lelie D, Wattiez R (2003) FEMS Microbiol Rev 27:385–410
Banci L, Bertini I, Ciofi-Baffoni S, Huffman DL, O’Halloran TV (2001) J Biol Chem 276:8415–8426
Bersch B, Rossy E, Covès J, Brutscher B (2003) J Biomol NMR 27:57–67
Laemmli UK (1970) Nature 227:680–685
Zhu G, Bax A (1990) J Magn Reson 101:114–119
Grzesiek S, Anglister J, Bax A (1993) J Magn Reson B101:114–119
Bax A, Mehlkopf AF, Smidt J (1979) J Magn Reson 35:167–169
Rückert M, Otting G (2000) J Am Chem Soc 122:7793–7797
Farrow NA, Muhandiram R, Singer AU, Pascal SM, Kay CM, Gish, G, Shoelson SE, Pawson T, Forman-Kay JD, Kay LE (1994) Biochemistry 34:868–878
Tsan P, Hus J-C, Caffrey M, Marion D, Blackledge M (2000) J Am Chem Soc 121:2311–2312
Blackledge M, Medvedeva S, Poncin M, Guerlesquin F, Bruschi M, Marion D (1995) J Mol Biol 245:661–681
Hus J-C, Marion D, Blackledge M (2000) J Mol Biol 298:927–936
Sibille N, Pardi A, Simorre J-P, Blackledge M (2001) J Am Chem Soc 49:12135–12146
Gurd F (1972) Methods Enzymol 25:424–438
Metzler WJ, Wittekind M, Goldfarb V, Mueller L, Farmer BT (1996) J Am Chem Soc 119:6800–6801
Smith B, Ito Y, Raine A, Teichmann S, Ben-Tovim L, Nietlispach D, Broadhurst RW, Terada T, Kelly M, Oschkinat H, Shibata T, Yokoyama S, Laue E (1996) J Biomol NMR 8:360–369
Gardner K, Rosen MK, Kay LE (1997) Biochemistry 36:1389–1401
Gardner K, Kay LE (1998) Annu Rev Biophys Biomol Struct 27:357–406
Banci L, Bertini I, Ciofi-Baffoni S, Finney LA, Outten CE, O’Halloran TV (2002) J Mol Biol 323:883–897
Lipari G, Szabo A (1982a) J Am Chem Soc 104:4546–4559
Lipari G, Szabo A (1982a) J Am Chem Soc 104:4559–4570
Dosset P, Hus J-C, Blackledge M, Marion D (2000) J Biomol NMR 16:23–28
Lu W, Zelazowski AJ, Stillman MJ (1993) Inorg Chem 32:919–926
Lu W, Stillman MJ (1993) J Am Chem Soc 115:3291–3299
Kihlken MA, Leech AP, Le Brun NE (2002) Biochem J 368:729–739
Morby AP, Hobman JL, Brown NL (1995) Mol Microbiol 17:25–35
DeSilva T, Veglia G, Porcelli F, Pranter AM, Opella SJ (2002) Biopolymers 64:189–197
Guex N, Peitsch MC (1997) Electrophoresis 1:2714–2723
Acknowledgements
We are very grateful to David Lemaire and David Lascoux for their help in mass spectrometry experiments, to Jean-Pierre Andrieu for N-terminal sequencing, to Elisabeth Mintz and Vincent Forge for the access to the CD apparatus, the use of the SPECFIT software, and for fruitful discussions. Finally, Tatiana Valleys and Max Mergeay are acknowledged for constant encouragements. E.R. and L.C. were supported by PhD grants from the Région Rhône-Alpes and the Commissariat à l’Energie Atomique, respectively.
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Rossy, E., Champier, L., Bersch, B. et al. Biophysical characterization of the MerP-like amino-terminal extension of the mercuric reductase from Ralstonia metallidurans CH34. J Biol Inorg Chem 9, 49–58 (2004). https://doi.org/10.1007/s00775-003-0495-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-003-0495-y