Abstract
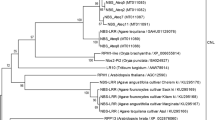
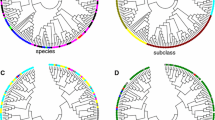
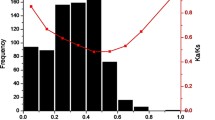
The Lesion Simulating Disease (LSD) genes encode a family of zinc finger proteins that play a role in programmed cell death (PCD) and other biological processes, such as plant growth and photosynthesis. In the present study, we report the reconstruction of the evolutionary history of the LSD gene family in Viridiplantae. Phylogenetic analysis revealed that the monocot and eudicot genes were distributed along the phylogeny, indicating that the expansion of the family occurred prior to the diversification between these clades. Sequences encoding proteins that present one, two, or three LSD domains formed separate groups. The secondary structure of these different LSD proteins presented a similar composition, with the β-sheets being their main component. The evolution by gene duplication was identified only to the genes that contain three LSD domains, which generated proteins with equal structure. Moreover, genes encoding proteins with one or two LSD domains evolved as single-copy genes and did not result from loss or gain in LSD domains. These results were corroborated by synteny analysis among regions containing paralogous/orthologous genes in Glycine max and Populus trichocarpa. The Ka/Ks ratio between paralogous/orthologous genes revealed that a subfunctionalization process possibly could be occurring with the LSD genes, explaining the involvement of LSD members in different biological processes, in addition to the negative regulation of PCD. This study presents important novelty in the evolutionary history of the LSD family and provides a basis for future research on individual LSD genes and their involvement in important pathway networks in plants.






Similar content being viewed by others
References
Atkinson NJ, Urwin PE (2012) The interaction of the plant biotic and abiotic stresses: from genes to the field. J Exp Bot 63:3523–3543
Bhatti KH, Xu C, Wu J, He C (2008) Overexpression of rice OsLOL2 gene confers disease resistance in tobacco to Pseudomonas syringae pv. tabaci. Prog Nat Sci 18:807–812
Buchan DW, Ward SM, Lobley AE, Nugent TC, Bryson K, Jones DT (2010) Protein annotation and modelling servers at University College London. Nucleic Acids Res 38:W563–W568
Cabreira C, Cagliari A, Bucker-Neto L, Wiebke-Strohm B, de Freitas LB, Marcelino-Guimaraes FC, Nepomuceno AL, Margis-Pinheiro MM, Bodanese-Zanettini MH (2013) The Lesion Simulating Disease (LSD) gene family as a variable in soybean response to Phakopsora pachyrhizi infection and dehydration. Funct Integr Genomics 13(3):323–338
Cagliari A, Turchetto-Zolet AC, Maraschin FS, Loss G, Margis R, Margis-Pinheiro M (2011) The evolutionary history of CBF transcription factors: gene duplication of CCAAT–binding factors NF-Y in plants In: Felix F (ed) Gene duplication, vol 1. InTech, Rijeka, pp 1–27
Cheng J, Baldi P (2005) Three-stage prediction of protein beta-sheets by neural networks, alignments and graph algorithms. Bioinformatics 21(Suppl 1):i75–i84
Coll NS, Vercammen D, Smidler A, Clover C, Van Breusegem F, Dangl JL, Epple P (2010) Arabidopsis type I metacaspases control cell death. Science 330(6009):1393–1397
Dietrich RA, Delaney TP, Uknes SJ, Ward ER, Ryals JA, Dangl JL (1994) Arabidopsis mutants simulating disease resistance response. Cell 77(4):565–577
Dietrich RA, Richberg MH, Schmidt R, Dean C, Dangl JL (1997) A novel zinc finger protein is encoded by the Arabidopsis LSD1 gene and functions as a negative regulator of plant cell death. Cell 88(5):685–694
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Epple P, Mack AA, Morris VR, Dangl JL (2003) Antagonistic control of oxidative stress-induced cell death in Arabidopsis by two related, plant-specific zinc finger proteins. Proc Natl Acad Sci USA 100(11):6831–6836
Fan C, Wang X, Hu R, Wang Y, Xiao C, Jiang Y, Zhang X, Zheng C, Fu YF (2013) The pattern of Phosphate transporter 1 genes evolutionary divergence in Glycine max L. BMC Plant Biol 13:48
Flagel LE, Wendel JF (2009) Gene duplication and evolutionary novelty in plants. New Phytol 183(3):557–564
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29(8):1023–1026
He S, Huang K, Zhang X, Yu X, Huang P, An C (2011a) The LSD1-type zinc finger motifs of Pisum sativa LSD1 are a novel nuclear localization signal and interact with importin alpha. PLoS One 6(7):e22131
He S, Tan G, Liu Q, Huang K, Ren J, Zhang X, Yu X, Huang P, An C (2011b) The LSD1-interacting protein GILP is a LITAF domain protein that negatively regulates hypersensitive cell death in Arabidopsis. PLoS One 6(4):e18750
Huang X, Li Y, Zhang X, Zuo J, Yang S (2010) The Arabidopsis LSD1 gene plays an important role in the regulation of low temperature-dependent cell death. New Phytol 187(2):301–312
Jabs T, Dietrich RA, Dangl JL (1996) Initiation of runaway cell death in an Arabidopsis mutant by extracellular superoxide. Science 273(5283):1853–1856
Jami SK, Clark GB, Ayele BT, Ashe P, Kirti PB (2012) Genome-wide comparative analysis of annexin superfamily in plants. PLoS One 7(11):e47801
Kaminaka H, Nake C, Epple P, Dittgen J, Schutze K, Chaban C, Holt BF III, Merkle T, Schafer E, Harter K, Dangl JL (2006) bZIP10-LSD1 antagonism modulates basal defense and cell death in Arabidopsis following infection. EMBO J 25(18):4400–4411
Kim KD, Shin JH, Van K, Kim DH, Lee SH (2009) Dynamic rearrangements determine genome organization and useful traits in soybean. Plant Physiol 151(3):1066–1076
Kliebenstein DJ, Dietrich RA, Martin AC, Last RL, Dangl JL (1999) LSD1 regulates salicylic acid induction of copper zinc superoxide dismutase in Arabidopsis thaliana. Mol Plant Microbe Interact 12(11):1022–1026
Kondrashov FA, Rogozin IB, Wolf YI, Koonin EV (2002) Selection in the evolution of gene duplications. Genome Biol 3(2):RESEARCH0008
Lee TH, Tang H, Wang X, Paterson AH (2013) PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res 41:D1152–D1158
Li W, Liu B, Yu L, Feng D, Wang H, Wang J (2009) Phylogenetic analysis, structural evolution and functional divergence of the 12-oxo-phytodienoate acid reductase gene family in plants. BMC Evol Biol 9:90
Li Y, Chen L, Mu J, Zuo J (2013) Lesion Simulating Disease1 interacts with catalases to regulate hypersensitive cell death in Arabidopsis. Plant Physiol 163(2):1059–1070
Lin L, Paterson AH (2011) Size variation in homologous segments across divergent plant genomes. Mob Genet Elements 1(2):92–96
Long M, de Souza SJ, Gilbert W (1995) Intron phase correlations and the evolution of intron/exon structure of genes. Proc Natl Acad Sci USA 92:1249512499
Lyons E, Pedersen B, Kane J, Alam M, Ming R, Tang H, Wang X, Bowers J, Paterson A, Lisch DMF (2008) Finding and comparing syntenic regions among Arabidopsis and the Outgroups Papaya, Poplar, and Grape: CoGe with Rosids. Bioinformatics 148:1772–1781
Mateo A, Muhlenbock P, Rusterucci C, Chang CC, Miszalski Z, Karpinska B, Parker JE, Mullineaux PM, Karpinski S (2004) Lesion Simulating Disease1 is required for acclimation to conditions that promote excess excitation energy. Plant Physiol 136(1):2818–2830
McClean PE, Mamidi S, McConnell M, Chikara S, Lee R (2010) Synteny mapping between common bean and soybean reveals extensive blocks of shared loci. BMC Genom 11:184
McGuffin LJ, Bryson K, Jones D (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405
Muhlenbock P, Plaszczyca M, Mellerowicz E, Karpinski S (2007) Lysigenous aerenchyma formation in Arabidopsis is controlled by Lesion Simulating Disease1. Plant Cell 19(11):3819–3830
Muhlenbock P, Szechynska-Hebda M, Plaszczyca M, Baudo M, Mateo A, Mullineaux PM, Parker JE, Karpinska B, Karpinski S (2008) Chloroplast signaling and Lesion Simulating Disease1 regulate crosstalk between light acclimation and immunity in Arabidopsis. Plant Cell 20(9):2339–2356
Mur LA, Kenton P, Lloyd AJ, Ougham H, Prats E (2008) The hypersensitive response; the centenary is upon us but how much do we know? J Exp Bot 59(3):501–520
Roulin A, Auer PL, Libault M, Schlueter J, Farmer A, May G, Stacey G, Doerge RW, Jackson SA (2013) The fate of duplicated genes in a polyploid plant genome. Plant J 73:143–153
Rusterucci C, Aviv DH, Holt BF III, Dangl JL, Parker JE (2001) The disease resistance signaling components EDS1 and PAD4 are essential regulators of the cell death pathway controlled by LSD1 in Arabidopsis. Plant Cell 13(10):2211–2224
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463(7278):178–183
Sharp PA (1981) Speculations on RNA splicing. Cell 23:643–646
Simillion C, Vandepoele K, Van Montagu MC, Zabeau M, Van de Peer Y (2002) The hidden duplication past of Arabidopsis thaliana. Proc Natl Acad Sci USA 99(21):13627–13632
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49(2):369–381
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
van Doorn WG, Beers EP, Dangl JL, Franklin-Tong VE, Gallois P, Hara-Nishimura I, Jones AM, Kawai-Yamada M, Lam E, Mundy J, Mur LAJ, Petersen M, Smertenko A, Taliansky M, Van Breusegem F, Wolpert T, Woltering E, Zhivotovsky B, Bozhkov PV (2011) Morphological classification of plant cell deaths. Cell Death Differ 18:1241–1246
Wang L, Pei Z, Tian Y, He C (2005) OsLSD1, a rice zinc finger protein, regulates programmed cell death and callus differentiation. Mol Plant Microbe Interact 18(5):375–384
Wituszynska W, Slesak I, Vanderauwera S, Szechynska-Hebda M, Kornas A, Van Der Kelen K, Muhlenbock P, Karpinska B, Mackowski S, Van Breusegem F, Karpinski S (2013) Lesion Simulating Disease1, enhanced disease susceptibility1, and phytoalexin deficient 4 conditionally regulate cellular signaling homeostasis, photosynthesis, water use efficiency, and seed yield in Arabidopsis. Plant Physiol 161(4):1795–1805
Xu C, He C (2007) The rice OsLOL2 gene encodes a zinc finger protein involved in rice growth and disease resistance. Mol Genet Genomics 278(1):85–94
Xu G, Guo C, Shan H, Kong H (2011) Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci USA 109(4):187–1192
Yin G, Xu H, Xiao S, Qin Y, Li Y, Yan Y, Hu Y (2013) The large soybean (Glycine max) WRKY TF family expanded by segmental duplication events and subsequent divergent selection among subgroups. BMC Plant Biol 13(1):148
Yu J, Wang J, Lin W, Li S, Li H, Zhou J, Ni P, Dong W, Hu S, Zeng C, Zhang J, Zhang Y, Li R, Xu Z, Li X, Zheng H, Cong L, Lin L, Yin J, Geng J, Li G, Shi J, Liu J, Lv H, Li J, Deng Y, Ran L, Shi X, Wang X, Wu Q, Li C, Ren X, Li D, Liu D, Zhang X, Ji Z, Zhao W, Sun Y, Zhang Z, Bao J, Han Y, Dong L, Ji J, Chen P, Wu S, Xiao Y, Bu D, Tan J, Yang L, Ye C, Xu J, Zhou Y, Yu Y, Zhang B, Zhuang S, Wei H, Liu B, Lei M, Yu H, Li Y, Xu H, Wei S, He X, Fang L, Huang X, Su Z, Tong W, Tong Z, Ye J, Wang L, Lei T, Chen C, Chen H, Huang H, Zhang F, Li N, Zhao C, Huang Y, Li L, Xi Y, Qi Q, Li W, Hu W, Tian X, Jiao Y, Liang X, Jin J, Gao L, Zheng W, Hao B, Liu S, Wang W, Yuan L, Cao M, McDermott J, Samudrala R, Wong GK, Yang H (2005) The genomes of Oryza sativa: a history of duplications. PLoS Biol 3(2):e38
Zhang L, Vision TJ, Gaut BS (2002) Patterns of nucleotide substitution among simultaneously duplicated gene pairs in Arabidopsis thaliana. Mol Biol Evol 19(9):464–1473
Acknowledgment
This study was funded by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Programa de Pós-Graduação em Genética e Biologia Molecular da Universidade Federal do Rio Grande do Sul (PPGBM/UFRGS), GENOSOJA/CNPq, and BIOTECSUR II/MCT.
Conflict of interest
The authors declare that they have no conflicts of interest.
Compliance with ethical standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Yang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cabreira, C., Cagliari, A., Bücker-Neto, L. et al. The phylogeny and evolutionary history of the Lesion Simulating Disease (LSD) gene family in Viridiplantae. Mol Genet Genomics 290, 2107–2119 (2015). https://doi.org/10.1007/s00438-015-1060-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-015-1060-4