Abstract
Main conclusion
A 44-base-pair region in the Chlamydomonas reinhardtii LHCBM9 promoter is essential for sulphur responsiveness.
The photosynthetic light-harvesting complex (LHC) proteins play essential roles both in light capture, the first step of photosynthesis, and in photoprotective mechanisms. In contrast to the other LHC proteins and the majority of photosynthesis proteins, the Chlamydomonas reinhardtii photosystem II-associated LHC protein, LHCBM9, was recently reported to be up-regulated under sulphur deprivation conditions, which also induce hydrogen production. Here, we examined the sulphur responsiveness of the LHCBM9 gene at the transcriptional level, through promoter deletion analysis. The LHCBM9 promoter was found to be responsive to sulphur deprivation, with a 44-base-pair region between nucleotide positions −136 and −180 relative to the translation start site identified as essential for this response. Anaerobiosis was found to enhance promoter activity under sulphur deprivation conditions, however, alone was unable to induce promoter activity. The study of LHCBM9 is of biological and biotechnological importance, as its expression is linked to photobiological hydrogen production, theoretically the most efficient process for biofuel production, while the simplicity of using an S-deprivation trigger enables the development of a novel C. reinhardtii-inducible promoter system based on LHCBM9.









Similar content being viewed by others
References
Aksoy M, Pootakham W, Pollock SV, Moseley JL, Gonzalez-Ballester D, Grossman AR (2013) Tiered regulation of sulfur deprivation responses in Chlamydomonas reinhardtii and identification of an associated regulatory factor. Plant Physiol 162(1):195–211
Arguello-Astorga GR, Herrera-Estrella LR (1996) Ancestral multipartite units in light-responsive plant promoters have structural features correlating with specific phototransduction pathways. Plant Physiol 112(3):1151–1166
Boekema EJ, van Roon H, van Breemen JFL, Dekker JP (1999) Supramolecular organization of photosystem II and its light-harvesting antenna in partially solubilized photosystem II membranes. Eur J Biochem 266(2):444–452
Caffarri S, Frigerio S, Olivieri E, Righetti PG, Bassi R (2005) Differential accumulation of Lhcb gene products in thylakoid membranes of Zea mays plants grown under contrasting light and temperature conditions. Proteomics 5(3):758–768
Chen YB, Durnford DG, Koblizek M, Falkowski PG (2004) Plastid regulation of Lhcb1 transcription in the chlorophyte alga Dunaliella tertiolecta. Plant Physiol 136(3):3737–3750
Dall’Osto L, Holt NE, Kaligotla S, Fuciman M, Cazzaniga S, Carbonera D, Frank HA, Alric J, Bassi R (2012) Zeaxanthin protects plant photosynthesis by modulating chlorophyll triplet yield in specific light-harvesting antenna subunits. J Biol Chem 287(50):41820–41834
Davies JP, Yildiz F, Grossman AR (1994) Mutants of Chlamydomonas with aberrant responses to sulfur deprivation. Plant Cell 6(1):53–63
Davies JP, Yildiz FH, Grossman A (1996) Sac1, a putative regulator that is critical for survival of Chlamydomonas reinhardtii during sulfur deprivation. EMBO J 15(9):2150–2159
Davies JP, Yildiz FH, Grossman AR (1999) Sac3, an Snf1-like serine threonine kinase that positively and negatively regulates the responses of Chlamydomonas to sulfur limitation. Plant Cell 11(6):1179–1190
Dehostos EL, Schilling J, Grossman AR (1989) Structure and expression of the gene encoding the periplasmic arylsulfatase of Chlamydomonas reinhardtii. Mol Gen Genet 218(2):229–239
Dekker JP, Boekema EJ (2005) Supramolecular organization of thylakoid membrane proteins in green plants. Biochim Biophys Acta-Bioenergetics 1706(1–2):12–39
Ding J, Li XM, Hu HY (2012) Systematic prediction of cis-regulatory elements in the Chlamydomonas reinhardtii genome using comparative genomics. Plant Physiol 160(2):613–623
Doebbe A, Keck M, La Russa M, Mussgnug JH, Hankamer B, Tekce E, Niehaus K, Kruse O (2010) The interplay of proton, electron, and metabolite supply for photosynthetic H-2 production in Chlamydomonas reinhardtii. J Biol Chem 285(39):30247–30260
Durnford DG, Falkowski PG (1997) Chloroplast redox regulation of nuclear gene transcription during photoacclimation. Photosynth Res 53(2–3):229–241
Elrad D, Grossman AR (2004) A genome’s-eye view of the light-harvesting polypeptides of Chlamydomonas reinhardtii. Curr Genet 45(2):61–75
Elrad D, Niyogi KK, Grossman AR (2002) A major light-harvesting polypeptide of photosystem II functions in thermal dissipation. Plant Cell 14(8):1801–1816
Escoubas JM, Lomas M, Laroche J, Falkowski PG (1995) Light intensity regulation of cab gene transcription is signaled by the redox state of the plastoquinone pool. Proc Natl Acad Sci U S A 92(22):10237–10241
Ferrante P, Catalanotti C, Bonente G, Giuliano G (2008) An optimized, chemically regulated gene expression system for Chlamydomonas. PLoS One 3(9):e3200
Ferrante P, Ballottari M, Bonente G, Giuliano G, Bassi R (2012) LHCBM1 and LHCBM2/7 polypeptides, components of major LHCII complex, have distinct functional roles in photosynthetic antenna system of Chlamydomonas reinhardtii. J Biol Chem 287(20):16276–16288
Fuhrmann M, Hausherr A, Ferbitz L, Schodl T, Heitzer M, Hegemann P (2004) Monitoring dynamic expression of nuclear genes in Chlamydomonas reinhardtii by using a synthetic luciferase reporter gene. Plant Mol Biol 55(6):869–881
Ghirardi ML, Zhang JP, Lee JW, Flynn T, Seibert M, Greenbaum E, Melis A (2000) Microalgae: a green source of renewable H-2. Trends Biotechnol 18(12):506–511
Giuliano G, Pichersky E, Malik VS, Timko MP, Scolnik PA, Cashmore AR (1988) An evolutionarily conserved protein binding sequence upstream of a plant light-regulated gene. Proc Natl Acad Sci U S A 85(19):7089–7093
Gonzalez-Ballester D, Pollock SV, Pootakham W, Grossman AR (2008) The central role of a SNRK2 kinase in sulfur deprivation responses. Plant Physiol 147(1):216–227
Gonzalez-Ballester D, Casero D, Cokus S, Pellegrini M, Merchant SS, Grossman AR (2010) RNA-Seq analysis of sulfur-deprived chlamydomonas cells reveals aspects of acclimation critical for cell survival. Plant Cell 22(6):2058–2084
Green PJ, Kay SA, Chua NH (1987) Sequence-specific interactions of a pea nuclear factor with light-responsive elements upstream of the rbcS-3A gene. EMBO J 6(9):2543–2549
Grewe S, Ballottari M, Alcocer M, D’Andrea C, Blifernez-Klassen O, Hankamer B, Mussgnug JH, Bassi R, Kruse O (2014) Light-harvesting complex protein LHCBM9 is critical for photosystem II activity and hydrogen production in Chlamydomonas reinhardtii. Plant Cell 26:1598–1611
Grossman AR, Gonzalez-Ballester D, Shibagaki N, Pootakham W, Moseley J (2010) Responses to macronutrient deprivation. In: Pareek A, Sopory SK, Bohnert HJ, Govindjee X (eds) Abiotic stress adaptation in plants: physiological, molecular and genomic foundation. Springer, New York, pp 307–348
Hahn D, Kuck U (1999) Identification of DNA sequences controlling light- and chloroplast-dependent expression of the lhcb1 gene from Chlamydomonas reinhardtii. Curr Genet 34(6):459–466
Haldrup A, Jensen PE, Lunde C, Scheller HV (2001) Balance of power: a view of the mechanism of photosynthetic state transitions. Trends Plant Sci 6(7):301–305
Harris EH (1989) The Chlamydomonas Sourcebook: A Comprehensive Guide to Biology and Laboratory Use. Academic Press, San Diego
Hell R (1997) Molecular physiology of plant sulfur metabolism. Planta 202(2):138–148
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27(1):297–300
Hirakawa Y, Kofuji R, Ishida K (2008) Transient transformation of a chlorarachniophyte alga, Lotharella amoebiformis (chlorarachniophyceae), with uidA and egfp reporter genes. J Phycol 44(3):814–820
Humby PL, Cunningham ML, Saunders HL, Price JA, Durnford DG (2009) Compartmental cross-talk in the regulation of light harvesting complex transcription under short-term light and temperature stress in Chlamydomonas reinhardtii. Botany 87(4):375–386
Huner NPA, Oquist G, Sarhan F (1998) Energy balance and acclimation to light and cold. Trends Plant Sci 3(6):224–230
Hwang SB, Herrin DL (1994) Control of the lhc gene transcription by the circadian clock in Chlamydomonas reinhardtii. Plant Mol Biol 26(2):557–569
Irihimovitch V, Stern DB (2006) The sulfur acclimation SAC3 kinase is required for chloroplast transcriptional repression under sulfur limitation in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 103(20):7911–7916
Jacobshagen S, Kindle KL, Johnson CH (1996) Transcription of CABII is regulated by the biological clock in Chlamydomonas reinhardtii. Plant Mol Biol 31(6):1173–1184
Jahns P, Latowski D, Strzalka K (2009) Mechanism and regulation of the violaxanthin cycle: the role of antenna proteins and membrane lipids. Biochim Biophys Acta-Bioenergetics 1787(1):3–14
Jansson S (1994) The light-harvesting chlorophyll a/b-binding proteins. Biochim Biophys Acta-Bioenergetics 1184(1):1–19
Jansson S (1999) A guide to the Lhc genes and their relatives in Arabidopsis. Trends Plant Sci 4(6):236–240
Jarvis EE, Brown LM (1991) Transient expression of firefly luciferase in protoplasts of the green alga Chlorella ellipsoidea. Curr Genet 19(4):317–321
Jensen PE, Bassi R, Boekema EJ, Dekker JP, Jansson S, Leister D, Robinson C, Scheller HV (2007) Structure, function and regulation of plant photosystem I. Biochim Biophys Acta-Bioenergetics 1767(5):335–352
Johnson MP, Ruban AV (2011) Restoration of Rapidly Reversible Photoprotective Energy Dissipation in the Absence of PsbS Protein by Enhanced Delta pH. J Biol Chem 286(22):19973–19981
Kakinuma M, Ikeda M, Coury DA, Tominaga H, Kobayashi I, Amano H (2009) Isolation and characterization of the rbcS genes from a sterile mutant of Ulva pertusa (Ulvales, Chlorophyta) and transient gene expression using the rbcS gene promoter. Fisheries Sci 75(4):1015–1028
Kindle KL (1987) Expression of a gene for a light-harvesting chlorophyll a/b-binding protein in Chlamydomonas reinhardtii: effect of light and acetate. Plant Mol Biol 9(6):547–563
Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 87(3):1228–1232
Koblenz B, Lechtreck KF (2005) The NIT1 promoter allows inducible and reversible silencing of centrin in Chlamydomonas reinhardtii. Eukaryot Cell 4(11):1959–1962
Kopriva S, Rennenberg H (2004) Control of sulphate assimilation and glutathione synthesis: interaction with N and C metabolism. J Exp Bot 55(404):1831–1842
Koziol AG, Borza T, Ishida KI, Keeling P, Lee RW, Durnford DG (2007) Tracing the evolution of the light-harvesting antennae in chlorophyll a/b-containing organisms. Plant Physiol 143(4):1802–1816
Kropat J, Tottey S, Birkenbihl RP, Depege N, Huijser P, Merchant S (2005) A regulator of nutritional copper signaling in Chlamydomonas is an SBP domain protein that recognizes the GTAC core of copper response element. Proc Natl Acad Sci U S A 102(51):18730–18735
Kruse O, Rupprecht J, Bader KP, Thomas-Hall S, Schenk PM, Finazzi G, Hankamer B (2005) Improved photobiological H-2 production in engineered green algal cells. J Biol Chem 280(40):34170–34177
Kuhlbrandt W, Wang DN, Fujiyoshi Y (1994) Atomic model of plant light-harvesting complex by electron crystallography. Nature 367(6464):614–621
Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouze P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Lien T, Schreiner O (1975) Purification of a derepressible arylsulfatase from Chlamydomonas reinhardti I—properties of the enzyme in intact cells and in purified state. Biochim Biophys Acta 384(1):168–179
Liu ZF, Yan HC, Wang KB, Kuang TY, Zhang JP, Gui LL, An XM, Chang WR (2004) Crystal structure of spinach major light-harvesting complex at 2.72 angstrom resolution. Nature 428(6980):287–292
Lopez Y, Patil A, Nakai K (2013) Identification of novel motif patterns to decipher the promoter architecture of co-expressed genes in Arabidopsis thaliana. BMC Syst Biol 7:S10
Mackerness SAH, Liu LS, Thomas B, Thompson WF, Jordan BR, White MJ (1998) Individual members of the light-harvesting complex II chlorophyll a/b binding protein gene family in pea (Pisum sativum) show differential responses to ultraviolet-B radiation. Physiol Plant 103(3):377–384
Maruyama-Nakashita A, Nakamura Y, Watanabe-Takahashi A, Inoue E, Yamaya T, Takahashi H (2005) Identification of a novel cis-acting element conferring sulfur deficiency response in Arabidopsis roots. Plant J 42(3):305–314
Maruyama-Nakashita A, Nakamura Y, Tohge T, Saito K, Takahashi H (2006) Arabidopsis SLIM1 is a central transcriptional regulator of plant sulfur response and metabolism. Plant Cell 18(11):3235–3251
Masuda T, Tanaka A, Melis A (2003) Chlorophyll antenna size adjustments by irradiance in Dunaliella salina involve coordinate regulation of chlorophyll a oxygenase (CAO) and Lhcb gene expression. Plant Mol Biol 51(5):757–771
Maxwell DP, Laudenbach DE, Huner NPA (1995) Redox regulation of light-harvesting complex II and cab mRNA abundance in Dunaliella salina. Plant Physiol 109(3):787–795
McKim SM, Durnford DG (2006) Translational regulation of light-harvesting complex expression during photo acclimation to high-light in Chlamydomonas reinhardtii. Plant Physiol Biochem 44(11–12):857–865
Melis A, Chen HC (2005) Chloroplast sulfate transport in green algae—genes, proteins and effects. Photosynths Res 86(3):299–307
Melis A, Zhang LP, Forestier M, Ghirardi ML, Seibert M (2000) Sustained photobiological hydrogen gas production upon reversible inactivation of oxygen evolution in the green alga Chlamydomonas reinhardtii. Plant Physiol 122(1):127–135
Mettler T, Muehlhaus T, Hemme D, Schoettler M-A, Rupprecht J, Idoine A, Veyel D, Pal SK, Yaneva-Roder L, Winck FV, Sommer F, Vosloh D, Seiwert B, Erban A, Burgos A, Arvidsson S, Schoenfelder S, Arnold A, Guenther M, Krause U, Lohse M, Kopka J, Nikoloski Z, Mueller-Roeber B, Willmitzer L, Bock R, Schroda M, Stitt M (2014) Systems analysis of the response of photosynthesis, metabolism, and growth to an increase in irradiance in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26(6):2310–2350
Millar AJ, Kay SA (1991) Circadian control of cab gene transcription and mRNA accumulation in Arabidopsis. Plant Cell 3(5):541–550
Miller G, Suzuki N, Rizhsky L, Hegie A, Koussevitzky S, Mittler R (2007) Double mutants deficient in cytosolic and thylakoid ascorbate peroxidase reveal a complex mode of interaction between reactive oxygen species, plant development, and response to abiotic stresses. Plant Physiol 144(4):1777–1785
Minagawa J, Takahashi Y (2004) Structure, function and assembly of Photosystem II and its light-harvesting proteins. Photosynth Res 82(3):241–263
Morishima A (1998) Identification of preferred binding sites of a light-inducible DNA-binding factor (MNF1) within 5’-upstream sequence of C4-type phosphoenolpyruvate carboxylase gene in maize. Plant Mol Biol 38(4):633–646
Moseley JL, Gonzalez-Ballester D, Pootakham W, Bailey S, Grossman AR (2009) Genetic interactions between regulators of chlamydomonas phosphorus and sulfur deprivation responses. Genetics 181(3):889–905
Mozzo M, Mantelli M, Passarini F, Caffarri S, Croce R, Bassi R (2010) Functional analysis of photosystem I light-harvesting complexes (Lhca) gene products of Chlamydomonas reinhardtii. Biochimt Biophys Acta-Bioenergetics 1797(2):212–221
Naumann B, Stauber EJ, Busch A, Sommer F, Hippler M (2005) N-terminal processing of Lhca3 is a key step in remodeling of the photosystem I-light-harvesting complex under iron deficiency in Chlamydomonas reinhardtii. J Biol Chem 280(21):20431–20441
Neilson JAD, Durnford DG (2010) Structural and functional diversification of the light-harvesting complexes in photosynthetic eukaryotes. Photosynth Res 106(1–2):57–71
Nguyen AV, Thomas-Hall SR, Malnoe A, Timmins M, Mussgnug JH, Rupprecht J, Kruse O, Hankamer B, Schenk PM (2008) Transcriptome for photobiological hydrogen production induced by sulfur deprivation in the green alga Chlamydomonas reinhardtii. Eukaryot Cell 7(11):1965–1979
Niyogi KK, Bjorkman O, Grossman AR (1997) Chlamydomonas xanthophyll cycle mutants identified by video imaging of chlorophyll fluorescence quenching. Plant Cell 9(8):1369–1380
Ohresser M, Matagne RF, Loppes R (1997) Expression of the arylsulphatase reporter gene under the control of the nit1 promoter in Chlamydomonas reinhardtii. Curr Genet 31(3):264–271
Pascal AA, Liu ZF, Broess K, van Oort B, van Amerongen H, Wang C, Horton P, Robert B, Chang WR, Ruban A (2005) Molecular basis of photoprotection and control of photosynthetic light-harvesting. Nature 436(7047):134–137
Peer W, Silverthorne J, Peters JL (1996) Developmental and light-regulated expression of individual members of the light-harvesting complex b gene family in Pinus palustris. Plant Physiol 111(2):627–634
Peers G, Truong TB, Ostendorf E, Busch A, Elrad D, Grossman AR, Hippler M, Niyogi KK (2009) An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462(7272):518–521
Piechulla B (1999) Circadian expression of the light-harvesting complex protein genes in plants. Chronobiol Int 16(2):115–128
Piechulla B, Merforth N, Rudolph B (1998) Identification of tomato Lhc promoter regions necessary for circadian expression. Plant Mol Biol 38(4):655–662
Quinn JM, Kropat J, Merchant S (2003) Copper response element and Crr1-dependent Ni2+-responsive promoter for induced, reversible gene expression in Chlamydomonas reinhardtii. Eukaryot Cell 2(5):995–1002
Ravina CG, Chang CI, Tsakraklides GP, McDermott JP, Vega JM, Leustek T, Gotor C, Davies JP (2002) The sac mutants of Chlamydomonas reinhardtii reveal transcriptional and posttranscriptional control of cysteine biosynthesis. Plant Physiol 130(4):2076–2084
Rochaix JD (2007) Role of thylakoid protein kinases in photosynthetic acclimation. FEBS Lett 581(15):2768–2775
Rouached H, Secco D, Arpat B, Poirier Y (2011) The transcription factor PHR1 plays a key role in the regulation of sulfate shoot-to-root flux upon phosphate starvation in Arabidopsis. BMC Plant Biol 11:19
Ruban AV, Horton P (1995) Regulation of non-photochemical quenching of chlorophyll fluorescence in plants. Aust J Plant Physiol 22(2):221–230
Ruban AV, Berera R, Ilioaia C, van Stokkum IHM, Kennis JTM, Pascal AA, van Amerongen H, Robert B, Horton P, van Grondelle R (2007) Identification of a mechanism of photoprotective energy dissipation in higher plants. Nature 450(7169):575–578
Schaffner AR, Sheen J (1991) Maize rbcS promoter activity depends on sequence elements not found in dicot rbcS promoters. Plant Cell 3(9):997–1012
Schreiner O, Lien T, Knutsen G (1975) The capacity for arylsulfatase synthesis in synchronous and synchronized cultures of Chlamydomonas reinhardtii. Biochim Biophys Acta 384(1):180–193
Shin R (2011) Transcriptional regulatory components responding to macronutrient limitation. J Plant Biol 54(5):286–293
Standfuss J, Terwisscha van Scheltinga AC, Lamborghini M, Kuhlbrandt W (2005) Mechanisms of photoprotection and nonphotochemical quenching in pea light-harvesting complex at 2.5 A resolution. EMBO J 24(5):919–928
Sugimoto K, Sato N, Tsuzuki M (2007) Utilization of a chloroplast membrane sulfolipid as a major internal sulfur source for protein synthesis in the early phase of sulfur starvation in Chlamydomonas reinhardtii. FEBS Lett 581(23):4519–4522
Sugimoto K, Midorikawa T, Tsuzuki M, Sato N (2008) Upregulation of PG synthesis on sulfur-starvation for PSI in Chlamydomonas. Biochem Biophys Res Commun 369(2):660–665
Takahashi H, Braby CE, Grossman AR (2001) Sulfur economy and cell wall biosynthesis during sulfur limitation of Chlamydomonas reinhardtii. Plant Physiol 127(2):665–673
Takahashi H, Iwai M, Takahashi Y, Minagawa J (2006) Identification of the mobile light-harvesting complex II polypeptides for state transitions in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 103(2):477–482
Teng CY, Qin S, Liu JG, Yu DZ, Liang CW, Tseng CK (2002) Transient expression of lacZ in bombarded unicellular green alga Haematococcus pluvialis. J Appl Phycol 14(6):497–500
Teramoto H, Ono T, Minagawa J (2001) Identification of Lhcb gene family encoding the light-harvesting chlorophyll-a/b proteins of photosystem II in Chlamydomonas reinhardtii. Plant Cell Physiol 42(8):849–856
Teramoto H, Nakamori A, Minagawa J, Ono T (2002) Light-intensity-dependent expression of Lhc gene family encoding light-harvesting chlorophyll-a/b proteins of photosystem II in Chlamydomonas reinhardtii. Plant Physiol 130(1):325–333
Terashima M, Specht M, Naumann B, Hippler M (2010) Characterizing the Anaerobic Response of Chlamydomonas reinhardtii by Quantitative Proteomics. Mol Cell Proteom 9(7):1514–1532
Timmins M, Zhou WX, Rupprecht J, Lim L, Thomas-Hall SR, Doebbe A, Kruse O, Hankamer B, Marx UC, Smith SM (2009) Schenk PM (2009) The metabolome of Chlamydomonas reinhardtii following induction of anaerobic H-2 production by sulfur depletion. J Biol Chem 284(51):35996
Tokutsu R, Iwai M, Minagawa J (2009) CP29, a monomeric light-harvesting complex II Protein, is essential for state transitions in Chlamydomonas reinhardtii. J Biol Chem 284(12):7777–7782
Turkina MV, Kargul J, Blanco-Rivero A, Villarejo A, Barber J, Vener AV (2006) Environmentally modulated phosphoproteome of photosynthetic membranes in the green alga Chlamydomonas reinhardtii. Mol Cell Proteom 5(8):1412–1425
Vaahtera L, Brosche M (2011) More than the sum of its parts—how to achieve a specific transcriptional response to abiotic stress. Plant Sci 180(3):421–430
Vandenbon A, Miyamoto Y, Takimoto N, Kusakabe T, Nakai K (2008) Markov chain-based promoter structure modeling for tissue-specific expression pattern prediction. DNA Res 15(1):3–11
Villand P, Eriksson M, Samuelsson G (1997) Carbon dioxide and light regulation of promoters controlling the expression of mitochondrial carbonic anhydrase in Chlamydomonas reinhardtii. Biochem J 327:51–57
Walker JC, Howard EA, Dennis ES, Peacock WJ (1987) DNA sequences required for anaerobic expression of the maize alcohol dehydrogenase 1 gene. Proc Natl Acad Sci U S A 84(19):6624–6628
Wang CH, Wang YY, Su Q, Gao XR (2007) Transient expression of the GUS gene in a unicellular marine green alga, Chlorella sp MACC/C95, via electroporation. Biotechnol Bioproc Eng 12(2):180–183
Waters MT, Langdale JA (2009) The making of a chloroplast. EMBO J 28(19):2861–2873
White MJ, Fristensky BW, Falconet D, Childs LC, Watson JC, Alexander L, Roe BA, Thompson WF (1992) Expression of the chlrophyll-a/b-protein multigene family in pea (Pisum ativum L.), Evidence for distinct developmental resonses. Planta 188(2):190–198
Wilson KE, Huner NPA (2000) The role of growth rate, redox-state of the plastoquinone pool and the trans-thylakoid Delta pH in photoacclimation of Chlorella vulgaris to growth irradiance and temperature. Planta 212(1):93–102
Wollman FA (2001) State transitions reveal the dynamics and flexibility of the photosynthetic apparatus. EMBO J 20(14):3623–3630
Xiao XD, Marzluf GA (1996) Identification of the native NIT2 major nitrogen regulatory protein in nuclear extracts of Neurospora crassa. Genetica 97(2):153–163
Yakushevska AE, Keegstra W, Boekema EJ, Dekker JP, Andersson J, Jansson S, Ruban AV, Horton P (2003) The structure of photosystem II in Arabidopsis: localization of the CP26 and CP29 antenna complexes. Biochem (Mosc) 42(3):608–613
Yanagisawa S (1995) A novel DNA-binding domain that may form a single zinc finger motif. Nucleic Acids Res 23(17):3403–3410
Yang T, Poovaiah BW (2002) A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J Biol Chem 277(47):45049–45058
Yildiz FH, Davies JP, Grossman AR (1994) Characterization of Sulfate Transport in Chlamydomonas reinhardtii during sulfur-limited and sulfur-sufficient growth. Plant Physiol 104(3):981–987
Yu LM, Lamb CJ, Dixon RA (1993) Purification and biochemical characterization of proteins which bind to the H-box cis-element implicated in transcriptional activation of plant defense genes. Plant J 3(6):805–816
Zhang ZD, Shrager J, Jain M, Chang CW, Vallon O, Grossman AR (2004) Insights into the survival of Chlamydomonas reinhardtii during sulfur starvation based on microarray analysis of gene expression. Eukaryot Cell 3(5):1331–1348
Acknowledgments
The authors would like to thank Ms Erin Ahern for constructing the initial LHCBM9 promoter vector (pLM) and for providing assistance with the preliminary cloning work, transformations and luciferase assays. The authors would also like to thank Dr Melanie Oey, for constructing the positive control vector, pMO59-luc, and for assistance with the cloning work. This work was supported by Grants to BH from the Australian Research Council (ARC) [Grant numbers DP130100346 and DP110101699] and an Australian Postgraduate Award scholarship to AS.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2015_2249_MOESM1_ESM.tif
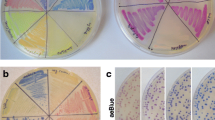
Supplementary material 1 Appearance of cultures after sulphur deprivation. Photo of sulphur-replete (+S) and sulphur-deprived (−S) untransformed Stm6 (-ve) and full-length LHCBM9 promoter (L9) transformants after 48 h of S deprivation. (TIFF 17116 kb)
425_2015_2249_MOESM2_ESM.tif
Supplementary material 2 Time course of luciferase activity. Luciferase activity was measured for the full-length LHCBM9 promoter transformant L9_83b, the deletion construct transformant L9Δ2_4 and the constitutive control (pMO59-luc5) over a period of 6 to 56 h post sulphur deprivation. The experiment was performed in triplicate in a 96 well plate with 200 µL of culture under 40 µE m−2 s−1 constant white light. The negative control was untransformed Stm6. Luciferase activity is reported in relative light units (RLU). An exponential line of best fit was plotted. (TIFF 879 kb)
425_2015_2249_MOESM3_ESM.tif
Supplementary material 3 Extended time course of luciferase activity. Luciferase activity is shown for the full-length LHCBM9 promoter transformant L9_83b, the LHCBM9 promoter deletion transformant L9Δ2_4 and the constitutive control (pMO59-luc5) 48–78 h post sulphur deprivation. The experiment was performed in triplicate in a 96 well plate with 200 µL of culture under 40 µE m−2s−1 constant white light. Luciferase activity is reported in relative light units (RLU). An exponential line of best fit was plotted. Note that the high apparent luciferase activity seen in the negative controls (untransformed Stm6) was an artefact caused by an unknown stress as evidenced by a lighter green colour. (TIFF 897 kb)
425_2015_2249_MOESM4_ESM.tif
Supplementary material 4 LHCBM9 promoter replicates. Luciferase activity of the full-length LHCBM9 promoter transformant L9_83b, the LHCBM9 promoter deletion transformant L9Δ2_4 and the positive control pMO59-luc5 after 48 h of sulphur (S) deprivation. Luciferase activity is reported in relative light units (RLU) and fold induction is displayed above the respective columns. Circles represent the RLU value obtained for single transformants and the black and grey lines indicate the mean values. Nine biological replicates were performed for each construct. The negative control was untransformed Stm6. A Student’s t test showed that L9_83b and L9Δ2_4 had significantly higher luciferase activity under –S conditions (P < 0.01). (TIFF 131 kb)
425_2015_2249_MOESM5_ESM.docx
Supplementary material 5 Clustal W alignment of the major light harvesting complex promoters. The coding sequence and first 1 kb upstream of the translation start site (ATG) was aligned using the Clustal W multiple alignment tool. Putative sulphur responsive elements (SURE) and the ATG are highlighted. Note that although the complete coding sequences were aligned, only the first ~ 50 bp are displayed in this figure. (DOCX 21 kb)
About this article
Cite this article
Sawyer, A.L., Hankamer, B.D. & Ross, I.L. Sulphur responsiveness of the Chlamydomonas reinhardtii LHCBM9 promoter. Planta 241, 1287–1302 (2015). https://doi.org/10.1007/s00425-015-2249-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-015-2249-9