Abstract
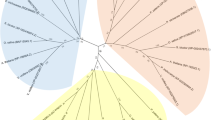
l-asparaginases (EC 3.5.1.1) are hypothesized to play an important role in nitrogen supply to sink tissues, especially in legume-developing seeds. Two plant l-asparaginase subtypes were previously identified according to their K+-dependence for catalytic activity. An l-asparaginase homologous to Lupinus K+-independent enzymes with activity towards β-aspartyl dipeptides, At5g08100, has been previously characterized as a member of the N-terminal nucleophile amidohydrolase superfamily in Arabidopsis. In this study, a K+-dependent l-asparaginase from Arabidopsis, At3g16150, is characterized. The recombinants At3g16150 and At5g08100 share a similar subunit structure and conserved autoproteolytic pentapeptide cleavage site, commencing with the catalytic Thr nucleophile, as determined by ESI-MS. The catalytic activity of At3g16150 was enhanced approximately tenfold in the presence of K+. At3g16150 was strictly specific for l-Asn, and had no activity towards β-aspartyl dipeptides. At3g16150 also had an approximately 80-fold higher catalytic efficiency with l-Asn relative to At5g08100. Among the β-aspartyl dipeptides tested, At5g08100 had a preference for β-aspartyl-His, with catalytic efficiency comparable to that with l-Asn. The phylogenetic analysis revealed that At3g16150 and At5g08100 belong to two distinct subfamilies. The transcript levels of At3g16150 and At5g08100 were highest in sink tissues, especially in flowers and siliques, early in development, as determined by quantitative RT-PCR. The overlapping spatial patterns of expression argue for a partially redundant function of the enzymes. However, the high catalytic efficiency suggests that the K+-dependent enzyme may metabolize l-Asn more efficiently under conditions of high metabolic demand for N.




Similar content being viewed by others
Abbreviations
- Ntn:
-
N-terminal nucleophile
- GUS:
-
β-Glucoronidase
- MES:
-
2-(N-morpholino)ethanesulfonic acid
- AGI:
-
Arabidopsis genome initiative
- ABRC:
-
Arabidopsis Biological Resource Center
- IPTG:
-
Isopropyl β-D-1-thiogalactopyranoside
- DAA:
-
Day after anthesis
- FW:
-
Fresh weight
- DEPC:
-
Diethyl pyrocarbonate
- TAIR:
-
The Arabidopsis Information Resource
- NCBI:
-
National Center for Biotechnology Information
- TIGR:
-
The Institute for Genome Research
- CBR:
-
Canadian Bioinformatics Resource
- LSD:
-
Least significant difference
- EST:
-
Expressed sequence tag
- HsGA:
-
Human glycosylasparaginase
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Aswad DW, Paranandi MV, Schurter BT (2000) Isoaspartate in peptides and proteins: formation, significance, and analysis. J Pharm Biomed Anal 21:1129–1136
Atkins CA, Pate JS, Sharkey PJ (1975) Asparagine metabolism key to the nitrogen nutrition of developing legume seeds. Plant Physiol 56:807–812
Borek D, Jaskolski M (2001) Sequence analysis of enzymes with asparaginase activity. Acta Biochim Pol 48:893–902
Borek D, Podkowinski J, Kisiel A, Jaskolski M (1999) Isolation and characterization of cDNA encoding l-asparaginase from Lupinus luteus (Accession No. AF112444) (PGR 99-050). Plant Physiol 119:1568
Borek D, Michalska K, Brzezinski K, Kisiel A, Podkowinski J, Bonthron DT, Krowarsch D, Otlewski J, Jaskolski M (2004) Expression, purification and catalytic activity of Lupinus luteus asparagine β-amidohydrolase and its Escherichia coli homolog. Eur J Biochem 271:3215–3226
Casado A, Caballero JL, Franco AR, Cardenas J, Grant MR, Munoz-Blanco J (1995) Molecular cloning of the gene encoding the L-asparaginase of Arabidopsis thaliana. Plant Physiol 108:1321–1322
Chagas EP, Sodek L (2001) Purification and properties of asparaginase from the testa of immature seeds of pea (Pisum sativum L.). Braz Arch Biol Technol 44:239–245
Chang KS, Farnden KJF (1981) Purification and properties of asparaginase (EC 3.5.1.1) from Lupinus arboreus and Lupinus angustifolius. Arch Biochem Biophys 208:49–58
Chirgwin JM, Przybyla AE, MacDonald RJ, Rutter WJ (1979) Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18:5294–5299
Clarke S (2003) Aging as war between chemical and biochemical processes: protein methylation and the recognition of age-damaged proteins for repair. Ageing Res Rev 2:263–285
Dickson JMJJ, Vincze E, Grant MR, Smith LA, Rodber KA, Farnden KJF, Reynolds PHS (1992) Molecular cloning of the gene encoding developing seed L-asparaginase from Lupinus angustifolius. Plant Mol Biol 20:333–336
Felsenstein J (1989) PHYLIP: phylogeny inference package (version 3.2). Cladistics 5:164–166
Fisher KJ, Tollersrud O, Aronson NN Jr (1990) Cloning and sequence analysis of a cDNA for human glycosylasparaginase. A single gene encodes the subunits of this lysosomal amidase. FEBS Lett 269:440–444
Grant M, Bevan MW (1994) Asparaginase gene expression is regulated in a complex spatial and temporal pattern in nitrogen-sink tissues. Plant J 5:695–704
Hejazi M, Piotukh K, Mattow J, Deutzmann R, Volkmer-Engert R, Lockau W (2002) Isoaspartyl dipeptidase activity of plant-type asparaginases. Biochem J 364:129–136
Hernández-Sebastià C, Marsolais F, Saravitz C, Israel D, Dewey RE, Huber SC (2005) Free amino acid profiles suggest a possible role for asparagine in the control of storage-product accumulation in developing seeds of low- and high-protein soybean lines. J Exp Bot 56:1951–1963
Hsieh JJ, Cheng EH, Korsmeyer SJ (2003) Taspase1: a threonine aspartase required for cleavage of MLL and proper HOX gene expression. Cell 115:293–303
Ireland RJ, Joy KW (1981) Two routes for asparagine metabolism in Pisum sativum L. Planta 151:289–292
Ireland RJ, Joy KW (1983) Subcellular localization of asparaginase and asparagine aminotransferase in Pisum sativum leaves. Plant Physiol 72:1127–1129
Ireland RJ, Lea PJ (1999) The enzymes of glutamine, glutamate, asparagine, and aspartate metabolism. In: Singh BK (ed) Plant amino acids: biochemistry and biotechnology. Dekker, New York, pp 49–109
Ishiyama K, Inoue E, Watanabe-Takahashi A, Obara M, Yamaya T, Takahashi H (2004) Kinetic properties and ammonium-dependent regulation of cytosolic isoenzymes of glutamine synthetase in Arabidopsis. J Biol Chem 279:16598–16605
Kern R, Malki A, Abdallah J, Liebart JC, Dubucs C, Yu MH, Richarme G (2005) Protein isoaspartate methyltransferase is a multicopy suppressor of protein aggregation in Escherichia coli. J Bacteriol 187:1377–1383
Laemlli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lam HM, Wong P, Chan HK, Yam KM, Chen L, Chow CM, Coruzzi GM (2003) Overexpression of the Asn1 gene enhances nitrogen status in seeds of Arabidopsis. Plant Physiol 132:926–935
Lanvers C, Pinheiro JPV, Hempel G, Wuerthwein G, Boos J (2002) Analytical validation of a microplate reader-based method for the therapeutic drug monitoring of L-asparaginase in human serum. Anal Biochem 309:117–126
Laughlin LT, Reed GH (1997) The monovalent cation requirement of rabbit muscle pyruvate kinase is eliminated by substitution of lysine for glutamate 177. Arch Biochem Biophys 348:262–267
Lea PJ, Ireland RJ (1999) Nitrogen metabolism in higher plants. In: Singh BK (ed) Plant amino acids: biochemistry and biotechnology. Dekker, New York, pp 1–47
Liepman AH, Olsen LJ (2001) Peroxisomal alanine: glyoxylate aminotransferase (Agt1) is a photorespiratory enzyme with multiple substrates in Arabidopsis thaliana. Plant J 25:487–498
Liepman AH, Olsen LJ (2004) Genomic analysis of aminotransferases in Arabidopsis thaliana. Crit Rev Plant Sci 23:73–89
Lough TJ, Chang KS, Canne A, Monk BC, Reynolds PHS, Farnden KJF (1992a) L-Asparaginase from developing seeds of Lupinus arboreus. Phytochemistry 31:1519–1527
Lough TJ, Reddington BD, Grant MR, Hill DF, Reynolds PHS, Farnden KJF (1992b) The isolation and characterisation of a cDNA clone encoding L-asparaginase from developing seeds of lupin (Lupinus arboreus). Plant Mol Biol 19:391–399
Meinke DW (1994) Seed development in Arabidopsis thaliana. In: Meyerowitz EM, Somerville CR (ed) Arabidopsis. Cold Spring Harbor Laboratory Press, Plainview, pp 253–295
Meinke DW, Sussex IM (1979) Embryo-lethal mutants of Arabidopsis thaliana. A model system for genetic analysis of plant embryo development. Dev Biol 72:50–61
Michalska K, Brzezinski K, Jaskolski M (2005) Crystal structure of isoaspartyl aminopeptidase in complex with L-aspartate. J Biol Chem 280:28484–28491
Möllering H (1985) L-Aspartate and L-asparagine. In: Bergmeyer J, Grassl M (eds) Metabolites 3: lipids, amino acids and related compounds, 3rd edn. VCH Verlagsgesellschaft, Weinheim, pp 350–357
Mudgett MB, Clarke S (1994) Hormonal and environmental responsiveness of a developmentally regulated protein repair L-isoaspartyl methyltransferase in wheat. J Biol Chem 269:25605–25612
Mudgett MB, Clarke S (1996) A distinctly regulated protein repair L-isoaspartylmethyltransferase from Arabidopsis thaliana. Plant Mol Biol 30:723–737
Mudgett MB, Lowenson JD, Clarke S (1997) Protein repair L-isoaspartyl methyltransferase in plants. Phylogenetic distribution and the accumulation of substrate proteins in aged barley seeds. Plant Physiol 115:1481–1489
Murray DR, Kennedy IR (1980) Changes in activities of enzymes of nitrogen metabolism in seed coats and cotyledons during embryo development in pea seeds. Plant Physiol 66:782–786
Murray AJS, Blackwell RD, Joy KW, Lea PJ (1987) Photorespiratory N donors, aminotransferase and photosynthesis in mutant of barley deficient in serine: glyoxylate aminotransferase activity. Planta 172:106–113
Palmiter RD (1974) Magnesium precipitation of ribonucleoprotein complexes. Expedient techniques for the isolation of undegraded polysomes and messenger ribonucleic acid. Biochemistry 13:3606–3615
Pate JS (1976) Nitrogen mobilization and cycling: case studies for carbon and nitrogen in organs of a legume. In: Wardlow IF, Passioura JB (eds) Transport and transfer processes in plants. Academic Press, New York, pp 447–462
Pate JS (1989) Origin, destination and fate of phloem solutes in relation to organ and whole plant functioning. In: Baker DA, Milburn JA (eds) Transport of photoassimilates. Longman, Harlow, pp 138–166
Ramírez-Silva L, Ferreira ST, Nowak T, de Gómez-Puyou MT, Gómez-Puyou A (2001) Dimethylsulfoxide promotes K+-independent activity of pyruvate kinase and the acquisition of the active catalytic conformation. Eur J Biochem 268:3267–3274
Robinson NE, Robinson AB (2001) Molecular clocks. Proc Natl Acad Sci USA 98:944–949
Rolletschek H, Hosein F, Miranda M, Heim U, Gotz KP, Schlereth A, Borisjuk L, Saalbach I, Wobus U, Weber H (2005) Ectopic expression of an amino acid transporter (Vfaap1) in seeds of Vicia narbonensis and pea increases storage proteins. Plant Physiol 137:1236–1249
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Shimizu T, Matsuoka Y, Shirasawa T (2005) Biological significance of isoaspartate and its repair system. Biol Pharm Bull 28:1590–1596
Sieciechowicz K, Ireland RJ, Joy KW (1985) Diurnal variation of asparaginase in developing pea leaves. Plant Physiol 77:506–508
Sieciechowicz K, Ireland RJ, Joy KW (1988a) Diurnal changes in asparaginase activity in pea leaves. II. Regulation of activity. J Exp Bot 39:707–721
Sieciechowicz K, Joy KW, Ireland RJ (1988b) Diurnal changes in asparaginase activity in pea leaves. I. The requirement for light for increased activity. J Exp Bot 39:695–706
Sieciechowicz K, Joy KW, Ireland RJ (1988c) The metabolism of asparagine in plants. Phytochemistry 27:663–671
Sodek L, Lea PJ (1993) Asparaginase from the testa of developing lupin and pea seeds. Phytochemistry 34:51–56
Sodek L, Lea PJ, Miflin BJ (1980) Distribution and properties of a potassium dependent asparaginase EC 3.5.1.1 isolated from developing seeds of Pisum sativum cultivar Feltham-First and other plants. Plant Physiol 65:22–26
Suelter CH (1970) Enzymes activated by monovalent cations. Science 168:789–795
Thapar N, Kim AK, Clarke S (2001) Distinct patterns of expression but similar biochemical properties of protein L-isoaspartyl methyltransferase in higher plants. Plant Physiol 125:1023–1035
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tonin GS, Sodek L (1990) Asparaginase, allantoinase and glutamine synthetase activities in soybean cotyledons grown in vitro. Phytochemistry 29:2829–2831
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun TP (2004) Della proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019
Walbot V, Clutter M, Sussex IM (1972) Reproductive development and embryogeny in Phaseolus. Phytomorphology 22:59–68
Wehner A, Harms E, Jennings MP, Beacham IR, Derst C, Bast P, Roehm KH (1992) Site-specific mutagenesis of Escherichia coli Asparaginase II. None of the three histidine residues is required for catalysis. Eur J Biochem 208:475–480
Xu Q, Belcastro MP, Villa ST, Dinkins RD, Clarke SG, Downie AB (2004) A second protein L-isoaspartyl methyltransferase gene in Arabidopsis produces two transcripts whose products are sequestered in the nucleus. Plant Physiol 136:2652–2664
Yanagisawa S, Akiyama A, Kisaka H, Uchimiya H, Miwa T (2004) Metabolic engineering with Dof1 transcription factor in plants: improved nitrogen assimilation and growth under low-nitrogen conditions. Proc Natl Acad Sci USA 101:7833–7838
Acknowledgements
We thank the ABRC (Ohio State University, Colombus, OH) for the At5g08100 cDNA clone, and Dr. Soon Park (Agriculture and Agri-Food Canada, Greenhouse and Processing Crops Research Centre, Harrow, Ontario) for seeds of white navy bean (P. vulgaris) cv. AC Compass. We also thank Alex Molnar for his expert graphical assistance in the preparation of figures.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bruneau, L., Chapman, R. & Marsolais, F. Co-occurrence of both l-asparaginase subtypes in Arabidopsis: At3g16150 encodes a K+-dependent l-asparaginase. Planta 224, 668–679 (2006). https://doi.org/10.1007/s00425-006-0245-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-006-0245-9