Abstract
Key message
Arabidopsis transgenic plants expressing the sunflower transcription factor HaWRKY76 exhibit increased yield and tolerance to drought and flood stresses. The genetic construct containing HaWRKY76 is proposed as a potential biotechnological tool to improve crops.
Abstract
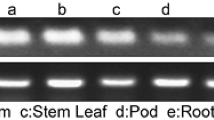
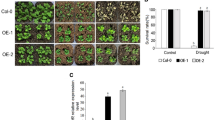
Water deficit and water excess are abiotic stress factors that seriously affect crops worldwide. To increase the tolerance to such stresses without causing yield penalty constitutes a major goal for biotechnologists. In this survey, we report that HaWRKY76, a divergent sunflower WRKY transcription factor, is able to confer both dehydration and submergence tolerance to Arabidopsis transgenic plants without yield penalty. The expression pattern of HaWRKY76 was analyzed in plants grown in standard conditions and under different watering regimes indicating a regulation by water availability. The corresponding cDNA was isolated and cloned under the control of a constitutive promoter and Arabidopsis plants were transformed with this construct. These transgenic plants presented higher biomass, seed production and sucrose content than controls in standard growth conditions. Moreover, they exhibited tolerance to mild drought or flood (complete submergence/waterlogging) stresses as well as the same or increased yield, depending on the stress severity and plant developmental stage, compared with controls. Drought tolerance occurred via an ABA-independent mechanism and induction of stomatal closure. Submergence tolerance can be explained by the carbohydrate (sucrose and starch) preservation achieved through the repression of fermentation pathways. Higher cell membrane stability and chlorenchyma maintenance could be the nexus between tolerance responses in front of both stresses. Altogether, the obtained results indicated that HaWRKY76 can be a potential biotechnological tool to improve crops yield as well as drought and flood tolerances.









Similar content being viewed by others
References
Albacete AA, Martinez-Andujar C, Perez-Alfocea F (2014) Hormonal and metabolic regulation of source-sink relations under salinity and drought: from plant survival to crop yield stability. Biotechnol Adv 32:12–30
Arce AL, Raineri J, Capella M, Cabello JV, Chan RL (2011) Uncharacterized conserved motifs outside the HD-Zip domain in HD-Zip subfamily I transcription factors; a potential source of functional diversity. BMC Plant Biol 11:42
Armstrong W (1979) Aeration in higher plants. In: Woolhouse HWW (ed) Advances in botanical research, vol 7. Academic Press, USA, pp 225–332
Bailey-Serres J, Voesenek L (2008) Flooding stress: acclimations and genetic diversity. Annu Rev Plant Biol 59:313–339
Baud S, Vaultier MN, Rochat C (2004) Structure and expression profile of the sucrose synthase multigene family in Arabidopsis. J Exp Bot 55:397–409
Baxter-Burrell A, Yang Z, Springer PS, Bailey-Serres J (2002) RopGAP4-dependent Rop GTPase rheostat control of Arabidopsis oxygen deprivation tolerance. Science 296:2026–2028
Blokhina O, Fagerstedt KV (2010) Reactive oxygen species and nitric oxide in plant mitochondria: origin and redundant regulatory systems. Physiol Plant 138:447–462
Cabello JV, Chan RL (2012) The homologous homeodomain-leucine zipper transcription factors HaHB1 and AtHB13 confer tolerance to drought and salinity stresses via the induction of proteins that stabilize membranes. Plant Biotechnol J 10:815–825
Chan RL, Gonzalez DH, Dezar CA, Gago GM (2010) Transcription Factor Gene Induced By Water Deficit Conditions And Abscisic Acid From Helianthus Annuus, Promoter And Transgenic Plants. Patent WO2004/099365
Chan RL, Cabello JV, Giacomelli JI (2013) HaHB11 provides improved plant yield and tolerance to abiotic stress. Patent WO2013116750 (A1)
Chen Q, Liu Z, Wang B, Wang X, Lai J, Tian F (2015) Transcriptome sequencing reveals the roles of transcription factors in modulating genotype by nitrogen interaction in maize. Plant Cell Rep. doi:10.1007/s00299-015-1822-9
Chory J, Reinecke D, Sim S, Washburn T, Brenner M (1994) A role for cytokinins in de-etiolation in Arabidopsis (det mutants have an altered response to cytokinins). Plant Physiol 104:339–347
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
de Bruxelles GL, Peacock WJ, Dennis ES, Dolferus R (1996) Abscisic acid induces the alcohol dehydrogenase gene in Arabidopsis. Plant Physiol 111:381–391
Ding ZJ, Yan JY, Xu XY, Yu DQ, Li GX, Zhang SQ, Zheng SJ (2014) Transcription factor WRKY46 regulates osmotic stress responses and stomatal movement independently in Arabidopsis. Plant J 79:13–27
Ellis MH, Dennis ES, Peacock WJ (1999) Arabidopsis roots and shoots have different mechanisms for hypoxic stress tolerance. Plant Physiol 119:57–64
FAO (2012) Sunflower: a ray of hope for flood-affected farmers in Sindh. Accessed December 2014 http://www.fao.org/emergencies/fao-in-action/stories/stories-detail/en/c/152907/
Fukao T, Xiong L (2013) Genetic mechanisms conferring adaptation to submergence and drought in rice: simple or complex? Curr Opin Plant Biol 16:196–204
Fukao T, Yeung E, Bailey-Serres J (2011) The submergence tolerance regulator SUB1A mediates crosstalk between submergence and drought tolerance in rice. Plant Cell 23:412–427
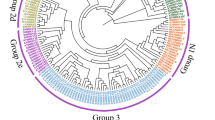
Giacomelli JI, Ribichich KF, Dezar CA, Chan RL (2010) Expression analyses indicate the involvement of sunflower WRKY transcription factors in stress responses, and phylogenetic reconstructions reveal the existence of a novel clade in the Asteraceae. Plant Sci 178:398–410
Giacomelli JI, Weigel D, Chan RL, Manavella PA (2012) Role of recently evolved miRNA regulation of sunflower HaWRKY6 in response to temperature damage. New Phytol 195:766–773
Gibbs DJ, Lee SC, Isa NM, Gramuglia S, Fukao T, Bassel GW, Correia CS, Corbineau F, Theodoulou FL, Bailey-Serres J, Holdsworth MJ (2011) Homeostatic response to hypoxia is regulated by the N-end rule pathway in plants. Nature 479:415–418
Gong X, Zhang J, Hu J, Wang W, Wu H, Zhang Q, Liu JH (2015) FcWRKY70, a WRKY protein of Fortunella crassifolia, functions in drought tolerance and modulates putrescine synthesis by regulating arginine decarboxylase gene. Plant Cell Environ. doi:10.1111/pce.12539
Hattori Y, Nagai K, Ashikari M (2011) Rice growth adapting to deepwater. Curr Opin Plant Biol 14:100–105
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052
Ismond KP, Dolferus R, De Pauw M, Dennis ES, Good AG (2003) Enhanced low oxygen survival in Arabidopsis through increased metabolic flux in the fermentative pathway. Plant Physiol 132:1292–1302
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Johnson JR, Cobb BG, Drew MC (1994) Hypoxic Induction of Anoxia Tolerance in Roots of Adh1 Null Zea mays L. Plant Physiol 105:61–67
Jung C, Seo JS, Han SW, Koo YJ, Kim CH, Song SI, Nahm BH, Choi YD, Cheong JJ (2008) Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis. Plant Physiol 146:623–635
Katinas L, Gutiérrez D, Grossi MA, Crisci JV (2007) Panorama de la familia Asteraceae (Compositae) en la Republica Argentina. Bol Soc Argent Bot 42:113–129
Kerstiens G (1996) Cuticular water permeability and its physiological significance. J Exp Bot 47:1813–1832
Kim EY, Seo YS, Park KY, Kim SJ, Kim WT (2014) Overexpression of CaDSR6 increases tolerance to drought and salt stresses in transgenic Arabidopsis plants. Gene 552:146–154
Koch KE, Ying Z, Wu Y, Avigne WT (2000) Multiple paths of sugar-sensing and a sugar/oxygen overlap for genes of sucrose and ethanol metabolism. J Exp Bot 51:417–427
Komatsu S, Deschamps T, Hiraga S, Kato M, Chiba M, Hashiguchi A, Tougou M, Shimamura S, Yasue H (2011) Characterization of a novel flooding stress-responsive alcohol dehydrogenase expressed in soybean roots. Plant Mol Biol 77:309–322
Lakshmanan M, Zhang Z, Mohanty B, Kwon JY, Choi HY, Nam HJ, Kim DI, Lee DY (2013) Elucidating rice cell metabolism under flooding and drought stresses using flux-based modeling and analysis. Plant Physiol 162:2140–2150
Licausi F, Kosmacz M, Weits DA, Giuntoli B, Giorgi FM, Voesenek LA, Perata P, van Dongen JT (2011) Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479:419–422
Liu H, Yang W, Liu D, Han Y, Zhang A, Li S (2011) Ectopic expression of a grapevine transcription factor VvWRKY11 contributes to osmotic stress tolerance in Arabidopsis. Mol Biol Rep 38:417–427
Liu QL, Zhong M, Li S, Pan YZ, Jiang BB, Jia Y, Zhang HQ (2013) Overexpression of a chrysanthemum transcription factor gene, DgWRKY3, in tobacco enhances tolerance to salt stress. Plant Physiol Biochem 69:27–33
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408
Loreti E, Poggi A, Novi G, Alpi A, Perata P (2005) A genome-wide analysis of the effects of sucrose on gene expression in Arabidopsis seedlings under anoxia. Plant Physiol 137:1130–1138
Luo X, Bai X, Sun X, Zhu D, Liu B, Ji W, Cai H, Cao L, Wu J, Hu M, Liu X, Tang L, Zhu Y (2013) Expression of wild soybean WRKY20 in Arabidopsis enhances drought tolerance and regulates ABA signalling. J Exp Bot 64:2155–2169
Ma D, Pu G, Lei C, Ma L, Wang H, Guo Y, Chen J, Du Z, Wang H, Li G, Ye H, Liu B (2009) Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the amorpha-4,11-diene synthase gene, a key gene of artemisinin biosynthesis. Plant Cell Physiol 50:2146–2161
Manzur ME, Grimoldi AA, Insausti P, Striker GG (2009) Escape from water or remain quiescent? Lotus tenuis changes its strategy depending on depth of submergence. Ann Bot 104:1163–1169
Nishiuchi S, Yamauchi T, Takahashi H, Kotula L, Nakazono M (2012) Mechanisms for coping with submergence and waterlogging in rice. Rice 5:1–14
O’Brien D, Belshe D, Meyer R, Falk J, Olson B (2009) High Plains Sunflower Production Handbook. K-State Research and Extension Publication, MF-2384
Okay S, Derelli E, Unver T (2014) Transcriptome-wide identification of bread wheat WRKY transcription factors in response to drought stress. Mol Genet Genomics 289:765–781
Panero JL, Funk VA (2008) The value of sampling anomalous taxa in phylogenetic studies: major clades of the Asteraceae revealed. Mol Phylogenet Evol 47:757–782
Quartacci MF, Navari-Izzo F (1992) Water stress and free radical mediated changes in sunflower seedlings. J Plant Physiol 139:621–625
Raineri J, Wang S, Peleg Z, Blumwald E, Chan RL (2015) The rice transcription factor OsWRKY47 is a positive regulator of the response to water deficit stress. Plant Mol Biol 88:401–413. doi:10.1007/s11103-015-0329-7
Rengel D, Arribat S, Maury P, Martin-Magniette ML, Hourlier T, Laporte M, Varès D, Carrère S, Grieu P, Balzergue S, Gouzy J, Vincourt P, Langlade NB (2012) A gene-phenotype network based on genetic variability for drought responses reveals key physiological processes in controlled and natural environments. PLoS One 7:e45249
Rivero RM, Kojima M, Gepstein A, Sakakibara H, Mittler R, Gepstein S, Blumwald E (2007) Delayed leaf senescence induces extreme drought tolerance in a flowering plant. Proc Natl Acad Sci USA 104:19631–19636
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258
Rushton DL, Tripathi P, Rabara RC, Lin J, Ringler P, Boken AK, Langum TJ, Smidt L, Boomsma DD, Emme NJ, Chen X, Finer JJ, Shen QJ, Rushton PJ (2012) WRKY transcription factors: key components in abscisic acid signalling. Plant Biotechnol J 10:2–11
Sachs MM, Freeling M, Okimoto R (1980) The anaerobic proteins of maize. Cell 20:761–767
Sasidharan R, Mustroph A, Boonman A, Akman M, Ammerlaan AM, Breit T, Schranz ME, Voesenek LA, van Tienderen PH (2013) Root transcript profiling of two Rorippa species reveals gene clusters associated with extreme submergence tolerance. Plant Physiol 163:1277–1292
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Setter TL, Bhekasut P, Greenway H (2010) Desiccation of leaves after de-submergence is one cause for intolerance to complete submergence of the rice cultivar IR 42. Funct Plant Biol 37:1096–1104
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Skirycz A, Vandenbroucke K, Clauw P, Maleux K, De Meyer B, Dhondt S, Pucci A, Gonzalez N, Hoeberichts F, Tognetti VB, Galbiati M, Tonelli C, Van Breusegem F, Vuylsteke M, Inzé D (2011) Survival and growth of Arabidopsis plants given limited water are not equal. Nat Biotechnol 29:212–214
Sukumaran N, Weiser C (1972) Freezing injury in potato leaves. Plant Physiol 50:564–567
Sun Y, Yu D (2015) Activated expression of AtWRKY53 negatively regulates drought tolerance by mediating stomatal movement. Plant Cell Rep 34:1295–1306 doi:10.1007/s00299-015-1787-8
Timme RE, Simpson BB, Randal Linder C (2007) High-resolution phylogeny for Helianthus (Asteraceae) using the 18S-26S ribosomal DNA external transcribed spacer. Am J Bot 94:1837–1852
Trinder P (1969) Determination of blood glucose using 4-amino phenazone as oxygen acceptor. J Clin Pathol 22:246
Tripathi P, Rabara RC, Rushton PJ (2014) A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 239:255–266
Vanderauwera S, Vandenbroucke K, Inzé A, van de Cotte B, Mühlenbock P, De Rycke R, Naouar N, Van Gaever T, Van Montagu MC, Van Breusegem F (2012) AtWRKY15 perturbation abolishes the mitochondrial stress response that steers osmotic stress tolerance in Arabidopsis. Proc Natl Acad Sci USA 109:20113–20118
Voesenek LA, Bailey-Serres J (2013) Flooding tolerance: O2 sensing and survival strategies. Curr Opin Plant Biol 16:647–653
Xiong L, Ishitani M, Lee H, Zhu JK (1999) HOS5-a negative regulator of osmotic stress-induced gene expression in Arabidopsis thaliana. Plant J 19:569–578
Yan H, Jia H, Chen X, Hao L, An H, Guo X (2014) The cotton WRKY transcription factor GhWRKY17 functions in drought and salt stress in transgenic Nicotiana benthamiana through ABA signaling and the modulation of reactive oxygen species production. Plant Cell Physiol 55:2060–2076
Acknowledgments
This work was supported by Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, PICT-PAE 37100 and PICT 2011 850), Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET, PIP 2011 11420100100278) and Universidad Nacional del Litoral (UNL, CAID 2011 50120110100399 and 50120110100349). KFR and RLC are members of CONICET and UNL; JR is a Fellow of CONICET and member of UNL. We thank Dr. Julieta Cabello and Dr. Gabriel Céccoli (IAL–CONICET UNL) for their valorous technical advice.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Menossi.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Raineri, J., Ribichich, K.F. & Chan, R.L. The sunflower transcription factor HaWRKY76 confers drought and flood tolerance to Arabidopsis thaliana plants without yield penalty. Plant Cell Rep 34, 2065–2080 (2015). https://doi.org/10.1007/s00299-015-1852-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1852-3