Abstract
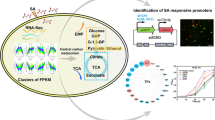
The thermotolerant Kluyveromyces marxianus is a potential candidate for high-temperature ethanol fermentation. Although K. marxianus exhibited high ethanol productivity at 45 °C during the early fermentation stage, we observed a fermentation arrest due to the accumulated inhibitors. The stress responses of K. marxianus during high-temperature fermentation were revealed based on integration of RNA sequencing (RNA-Seq) and metabolite data. High temperature stimulated mitochondrial respiration but repressed the tricarboxylic acid (TCA) cycle, leading to increased generation of reactive oxygen species (ROS) and a lowered ratio of reduced nicotinamide adenine dinucleotide (NADH)/oxidized nicotinamide adenine dinucleotide (NAD+). Glycerol production was enhanced during the early fermentation stage, which might contribute to NADH reoxidation and ROS generation. Excess ROS could be neutralized by reduced nicotinamide adenine dinucleotide phosphate (NADPH) that might be reserved in the following ways: (1) decreased biosynthesis of branched-chain amino acids (BCAAs) reduced NADPH consumption; (2) enhanced acetic acid production increased NADPH regeneration. The degree of fatty acid unsaturation was also reduced to adapt to high temperature. In addition, stress responses were also observed after the fermentation arrest at 45 °C. Genes related to peroxidase activity, iron-sulfur cluster assembly, and flavin mononucleotide (FMN) binding were downregulated, while genes associated with DNA repair and lipid composition of the plasma were upregulated. The yeast also produced more ergosterol to deal with ethanol stress. This study gains comprehensive insights into the K. marxianus transcriptome under various stresses during high-temperature ethanol fermentation, providing rich information for further metabolic engineering towards improved stress tolerance and ethanol production.






Similar content being viewed by others
References
Abdel-Banat BMA, Hoshida H, Ano A, Nonklang S, Akada R (2010) High-temperature fermentation: how can processes for ethanol production at high temperatures become superior to the traditional process using mesophilic yeast? Appl Microbiol Biotechnol 85:861–867. https://doi.org/10.1007/s00253-009-2248-5
Avery AM, Avery SV (2001) Saccharomyces cerevisiae expresses three phospholipid hydroperoxide glutathione peroxidases. J Biol Chem 276:33730–33735. https://doi.org/10.1074/jbc.M105672200
Bai FW, Anderson WA, Moo-Young M (2008) Ethanol fermentation technologies from sugar and starch feedstocks. Biotechnol Adv 26:89–105. https://doi.org/10.1016/j.biotechadv.2007.09.002
Ballweg S, Ernst R (2017) Control of membrane fluidity: the OLE pathway in focus. Biol Chem 398:215–228. https://doi.org/10.1515/hsz-2016-0277
Bernstein H, Bernstein C (2013) Evolutionary origin and adaptive function of meiosis. In: Bernstein C, Bernstein H (eds) Meiosis. IntechOpen, London, pp 41–75
Blank LM, Lehmbeck F, Sauer U (2005) Metabolic-flux and network analysis in fourteen hemiascomycetous yeasts. FEMS Yeast Res 5:545–558. https://doi.org/10.1016/j.femsyr.2004.09.008
Boveris A, Cadenas E, Stoppani AO (1976) Role of ubiquinone in the mitochondrial generation of hydrogen peroxide. Biochem J 156:435–444. https://doi.org/10.1042/bj1560435
Casal M, Cardoso H, Leao C (1996) Mechanisms regulating the transport of acetic acid in Saccharomyces cerevisiae. Microbiology 142:1385–1390. https://doi.org/10.1099/13500872-142-6-1385
Caspeta L, Caro-Bermudez MA, Ponce-Noyola T, Martinez A (2014) Enzymatic hydrolysis at high-solids loadings for the conversion of agave bagasse to fuel ethanol. Appl Energy 113:277–286. https://doi.org/10.1016/j.apenergy.2013.07.036
Davidson JF, Schiestl RH (2001) Mitochondrial respiratory electron carriers are involved in oxidative stress during heat stress in Saccharomyces cerevisiae. Mol Cell Biol 21:8483–8489. https://doi.org/10.1128/MCB.21.24.8483-8489.2001
Diniz RHS, Villada JC, Alvim MCT, Vidigal PMP, Vieira NM, Lamas-Maceiras M, Cerdan ME, Gonzalez-Siso MI, Lahtvee PJ, da Silveira WB (2017) Transcriptome analysis of the thermotolerant yeast Kluyveromyces marxianus CCT 7735 under ethanol stress. Appl Microbiol Biotechnol 101:6969–6980. https://doi.org/10.1007/s00253-017-8432-0
Dong Y, Hu J, Fan L, Chen Q (2017) RNA-Seq-based transcriptomic and metabolomic analysis reveal stress responses and programmed cell death induced by acetic acid in Saccharomyces cerevisiae. Sci Rep 7:42659. https://doi.org/10.1038/srep42659
Fernie AR, Carrari F, Sweetlove LJ (2004) Respiratory metabolism: glycolysis, the TCA cycle and mitochondrial electron transport. Curr Opin Plant Biol 7:254–261. https://doi.org/10.1016/j.pbi.2004.03.007
Fu X, Li P, Zhang L, Li S (2018) RNA-Seq-based transcriptomic analysis of Saccharomyces cerevisiae during solid-state fermentation of crushed sweet sorghum stalks. Process Biochem 68:53–63. https://doi.org/10.1016/j.procbio.2018.02.024
Gao J, Yuan W, Li Y, Xiang R, Hou S, Zhong S, Bai F (2015) Transcriptional analysis of Kluyveromyces marxianus for ethanol production from inulin using consolidated bioprocessing technology. Biotechnol Biofuels 8:115. https://doi.org/10.1186/s13068-015-0295-y
Giannattasio S, Guaragnella N, Zdralevic M, Marra E (2013) Molecular mechanisms of Saccharomyces cerevisiae stress adaptation and programmed cell death in response to acetic acid. Front Microbiol 4:33. https://doi.org/10.3389/fmicb.2013.00033
Goldemberg J (2007) Ethanol for a sustainable energy future. Science 315:808–810. https://doi.org/10.1126/science.1137013
Hillar A, Nicholls P, Switala J, Loewen PC (1994) NADPH binding and control of catalase compound II formation: comparison of bovine, yeast, and Escherichia coli enzymes. Biochem J 300:531–539. https://doi.org/10.1042/bj3000531
Hori A, Yoshida M, Shibata T, Ling F (2009) Reactive oxygen species regulate DNA copy number in isolated yeast mitochondria by triggering recombination-mediated replication. Nucleic Acids Res 37:749–761. https://doi.org/10.1093/nar/gkn993
Ismail KSK, Sakamoto T, Hatanaka H, Hasunuma T, Kondo A (2013) Gene expression cross-profiling in genetically modified industrial Saccharomyces cerevisiae strains during high-temperature ethanol production from xylose. J Biotechnol 163:50–60. https://doi.org/10.1016/j.jbiotec.2012.10.017
Lahtvee PJ, Kumar R, Hallstrom BM, Nielsen J (2016) Adaptation to different types of stress converge on mitochondrial metabolism. Mol Biol Cell 27:2505–2514. https://doi.org/10.1091/mbc.E16-03-0187
Lane MM, Burke N, Karreman R, Wolfe KH, O'Byrne CP, Morrissey JP (2011) Physiological and metabolic diversity in the yeast Kluyveromyces marxianus. Anton Leeuw Int J G 100:507–519. https://doi.org/10.1007/s10482-011-9606-x
Le HD, Thanonkeo P, Le VV (2013) Impact of high temperature on ethanol fermentation by Kluyveromyces marxianus immobilized on banana leaf sheath pieces. Appl Biochem Biotechnol 171:806–816. https://doi.org/10.1007/s12010-013-0411-z
Lertwattanasakul N, Kosaka T, Hosoyama A, Suzuki Y, Rodrussamee N, Matsutani M, Murata M, Fujimoto N, Suprayogi TK, Limtong S, Fujita N, Yamada M (2015) Genetic basis of the highly efficient yeast Kluyveromyces marxianus: complete genome sequence and transcriptome analyses. Biotechnol Biofuels 8:47. https://doi.org/10.1186/s13068-015-0227-x
Lertwattanasakul N, Shigemoto E, Rodrussamee N, Limtong S, Thanonkeo P, Yamada M (2009) The crucial role of alcohol dehydrogenase Adh3 in Kluyveromyces marxianus mitochondrial metabolism. Biosci Biotechnol Biochem 73:2720–2726. https://doi.org/10.1271/bbb.90609
Li BZ, Cheng JS, Ding MZ, Yuan YJ (2010) Transcriptome analysis of differential responses of diploid and haploid yeast to ethanol stress. J Biotechnol 148:194–203. https://doi.org/10.1016/j.jbiotec.2010.06.013
Li J, Li S, Han B, Yu M, Li G, Jiang Y (2013) A novel cost-effective technology to convert sucrose and homocelluloses in sweet sorghum stalks into ethanol. Biotechnol Biofuels 6:174. https://doi.org/10.1186/1754-6834-6-174
Li P, Fu X, Li S, Zhang L (2018a) Engineering TATA-binding protein Spt15 to improve ethanol tolerance and production in Kluyveromyces marxianus. Biotechnol Biofuels 11:207. https://doi.org/10.1186/s13068-018-1206-9
Li P, Fu X, Zhang L, Li S (2018b) CRISPR/Cas-based screening of a gene activation library in Saccharomyces cerevisiae identifies a crucial role of OLE1 in thermotolerance. Microb Biotechnol. https://doi.org/10.1111/1751-7915.13333
Li P, Fu X, Zhang L, Zhang Z, Li J, Li S (2017) The transcription factors Hsf1 and Msn2 of thermotolerant Kluyveromyces marxianus promote cell growth and ethanol fermentation of Saccharomyces cerevisiae at high temperatures. Biotechnol Biofuels 10:289. https://doi.org/10.1186/s13068-017-0984-9
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967. https://doi.org/10.1093/bioinformatics/btp336
Li Y, Chang WQ, Zhang M, Li XB, Jiao Y, Lou HX (2015) Diorcinol D exerts fungicidal action against Candida albicans through cytoplasm membrane destruction and ROS accumulation. PLoS One 10:e0128693. https://doi.org/10.1371/journal.pone.0128693
Limtong S, Sringiew C, Yongmanitchai W (2007) Production of fuel ethanol at high temperature from sugar cane juice by a newly isolated Kluyveromyces marxianus. Bioresour Technol 98:3367–3374. https://doi.org/10.1016/j.biortech.2006.10.044
Lobs AK, Engel R, Schwartz C, Flores A, Wheeldon I (2017) CRISPR-Cas9-enabled genetic disruptions for understanding ethanol and ethyl acetate biosynthesis in Kluyveromyces marxianus. Biotechnol Biofuels 10:164. https://doi.org/10.1186/S13068-017-0854-5
Ludovico P, Sousa MJ, Silva MT, Leao C, Corte-Real M (2001) Saccharomyces cerevisiae commits to a programmed cell death process in response to acetic acid. Microbiology 147:2409–2415. https://doi.org/10.1099/00221287-147-9-2409
Minard KI, McAlister-Henn L (2001) Antioxidant function of cytosolic sources of NADPH in yeast. Free Radic Biol Med 31:832–843. https://doi.org/10.1016/S0891-5849(01)00666-9
Morano KA, Grant CM, Moye-Rowley WS (2012) The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 190:1157–1195. https://doi.org/10.1534/genetics.111.128033
Navarro-Tapia E, Nana RK, Querol A, Perez-Torrado R (2016) Ethanol cellular defense induce unfolded protein response in yeast. Front Microbiol 7:189. https://doi.org/10.3389/Fmicb.2016.00189
Nielsen J, Larsson C, van Maris A, Pronk J (2013) Metabolic engineering of yeast for production of fuels and chemicals. Curr Opin Biotechnol 24:398–404. https://doi.org/10.1016/j.copbio.2013.03.023
Palma M, Guerreiro JF, Sa-Correia I (2018) Adaptive response and tolerance to acetic acid in Saccharomyces cerevisiae and Zygosaccharomyces bailii: a physiological genomics perspective. Front Microbiol 9:274. https://doi.org/10.3389/fmicb.2018.00274
Pan S, Jia B, Liu H, Wang Z, Chai M-Z, Ding M-Z, Zhou X, Li X, Li C, Li B-Z, Yuan Y-J (2018) Endogenous lycopene improves ethanol production under acetic acid stress in Saccharomyces cerevisiae. Biotechnol Biofuels 11:107. https://doi.org/10.1186/s13068-018-1107-y
Pocsi I, Prade RA, Penninckx MJ (2004) Glutathione, altruistic metabolite in fungi. Adv Microb Physiol 49:1–76. https://doi.org/10.1016/S0065-2911(04)49001-8
Postmus J, Tuzun I, Bekker M, Muller WH, de Mattos MJT, Brul S, Smits GJ (2011) Dynamic regulation of mitochondrial respiratory chain efficiency in Saccharomyces cerevisiae. Microbiology 157:3500–3511. https://doi.org/10.1099/mic.0.050039-0
Remize F, Andrieu E, Dequin S (2000) Engineering of the pyruvate dehydrogenase bypass in Saccharomyces cerevisiae: role of the cytosolic Mg2+ and mitochondrial K+ acetaldehyde dehydrogenases Ald6p and Ald4p in acetate formation during alcoholic fermentation. Appl Environ Microbiol 66:3151–3159. https://doi.org/10.1128/AEM.66.8.3151-3159.2000
Salvo A, Brito J, Artaxo P, Geiger FM (2017) Reduced ultrafine particle levels in Sao Paulo’s atmosphere during shifts from gasoline to ethanol use. Nat Commun 8:77. https://doi.org/10.1038/s41467-017-00041-5
Scherz-Shouval R, Elazar Z (2007) ROS, mitochondria and the regulation of autophagy. Trends Cell Biol 17:422–427. https://doi.org/10.1016/j.tcb.2007.07.009
Shui WQ, Xiong Y, Xiao WD, Qi XN, Zhang Y, Lin YP, Guo YF, Zhang ZD, Wang QH, Ma YH (2015) Understanding the mechanism of thermotolerance distinct from heat shock response through proteomic analysis of industrial strains of Saccharomyces cerevisiae. Mol Cell Proteomics 14:1885–1897. https://doi.org/10.1074/mcp.M114.045781
Steensels J, Snoek T, Meersman E, Picca Nicolino M, Voordeckers K, Verstrepen KJ (2014) Improving industrial yeast strains: exploiting natural and artificial diversity. FEMS Microbiol Rev 38:947–995. https://doi.org/10.1111/1574-6976.12073
Suutari M, Liukkonen K, Laakso S (1990) Temperature adaptation in yeasts: the role of fatty acids. J Gen Microbiol 136:1469–1474. https://doi.org/10.1099/00221287-136-8-1469
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Tarrio N, Garcia-Leiro A, Cerdan ME, Gonzalez-Siso MI (2008) The role of glutathione reductase in the interplay between oxidative stress response and turnover of cytosolic NADPH in Kluyveromyces lactis. FEMS Yeast Res 8:597–606. https://doi.org/10.1111/j.1567-1364.2008.00366.x
Vanegas JM, Contreras MF, Faller R, Longo ML (2012) Role of unsaturated lipid and ergosterol in ethanol tolerance of model yeast biomembranes. Biophys J 102:507–516. https://doi.org/10.1016/j.bpj.2011.12.038
Vogel C, Marcotte EM (2012) Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet 13:227–232. https://doi.org/10.1038/nrg3185
Voordeckers K, Kominek J, Das A, Espinosa-Cantu A, De Maeyer D, Arslan A, Van Pee M, van der Zande E, Meert W, Yang Y, Zhu B, Marchal K, DeLuna A, Van Noort V, Jelier R, Verstrepen KJ (2015) Adaptation to high ethanol reveals complex evolutionary pathways. PLoS Genet 11:e1005635. https://doi.org/10.1371/journal.pgen.1005635
Wang D, Wu D, Yang X, Hong J (2018) Transcriptomic analysis of thermotolerant yeast Kluyveromyces marxianus in multiple inhibitors tolerance. RSC Adv 8:14177–14192. https://doi.org/10.1039/c8ra00335a
Welker S, Rudolph B, Frenzel E, Hagn F, Liebisch G, Schmitz G, Scheuring J, Kerth A, Blume A, Weinkauf S, Haslbeck M, Kessler H, Buchner J (2010) Hsp12 is an intrinsically unstructured stress protein that folds upon membrane association and modulates membrane function. Mol Cell 39:507–520. https://doi.org/10.1016/j.molcel.2010.08.001
Woo JM, Yang KM, Kim SU, Blank LM, Park JB (2014) High temperature stimulates acetic acid accumulation and enhances the growth inhibition and ethanol production by Saccharomyces cerevisiae under fermenting conditions. Appl Microbiol Biotechnol 98:6085–6094. https://doi.org/10.1007/s00253-014-5691-x
Yarimizu T, Nonklang S, Nakamura J, Tokuda S, Nakagawa T, Lorreungsil S, Sutthikhumpha S, Pukahuta C, Kitagawa T, Nakamura M, Cha-aim K, Limtong S, Hoshida H, Akada R (2013) Identification of auxotrophic mutants of the yeast Kluyveromyces marxianus by non-homologous end joining-mediated integrative transformation with genes from Saccharomyces cerevisiae. Yeast 30:485–500. https://doi.org/10.1002/yea.2985
Ying WH (2008) NAD+/NADH and NADP+/NADPH in cellular functions and cell death: regulation and biological consequences. Antioxid Redox Signal 10:179–206. https://doi.org/10.1089/ars.2007.1672
Acknowledgements
We are grateful to Mr. Shengkai Pan, Chinese Academy of Sciences, for his help in analyzing RNA-Seq data. We thank Dr. Savitree Limtong, Kasetsart University, Thailand, for her help in purchasing K. marxianus DMKU3-1042. We also thank Ms. Lan Summer Sun, China National Knowledge Infrastructure, for her generous help in the English editing.
Funding
This study was funded by the National Natural Science Foundation of China (grant numbers 31600067, 21506113, 31470229), the National Key R&D Program of China (grant number 2016YFE0108500), and the China Postdoctoral Science Foundation (grant numbers 2017T100064, 2015M581074).
Author information
Authors and Affiliations
Contributions
XF and PL contributed equally to this work. PL, SL, and LZ designed the experiments. XF performed the fermentation experiments, HPLC analysis, assays of NAD(H) and NADP(H), and RNA extraction. PL and XF performed the assays of ROS and pyruvate. PL analyzed the RNA-Seq data and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Online Resource 1
(PDF 1463 kb)
Online Resource 2
(XLSX 21 kb)
Online Resource 3
(XLSX 485 kb)
Online Resource 4
(XLSX 24 kb)
Online Resource 5
(XLSX 446 kb)
Rights and permissions
About this article
Cite this article
Fu, X., Li, P., Zhang, L. et al. Understanding the stress responses of Kluyveromyces marxianus after an arrest during high-temperature ethanol fermentation based on integration of RNA-Seq and metabolite data. Appl Microbiol Biotechnol 103, 2715–2729 (2019). https://doi.org/10.1007/s00253-019-09637-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09637-x