Abstract
CYP109E1 is a cytochrome P450 monooxygenase from Bacillus megaterium with a hydroxylation activity for testosterone and vitamin D3. This study reports the screening of a focused library of statins, terpene-derived and steroidal compounds to explore the substrate spectrum of this enzyme. Catalytic activity of CYP109E1 towards the statin drug-precursor compactin and the prodrugs lovastatin and simvastatin as well as biotechnologically relevant terpene compounds including ionones, nootkatone, isolongifolen-9-one, damascones, and β-damascenone was found in vitro. The novel substrates induced a type I spin-shift upon binding to P450 and thus permitted to determine dissociation constants. For the identification of conversion products by NMR spectroscopy, a B. megaterium whole-cell system was applied. NMR analysis revealed for the first time the ability of CYP109E1 to catalyze an industrially highly important reaction, the production of pravastatin from compactin, as well as regioselective oxidations generating drug metabolites (6′β-hydroxy-lovastatin, 3′α-hydroxy-simvastatin, and 4″-hydroxy-simvastatin) and valuable terpene derivatives (3-hydroxy-α-ionone, 4-hydroxy-β-ionone, 11,12-epoxy-nootkatone, 4(R)-hydroxy-isolongifolen-9-one, 3-hydroxy-α-damascone, 4-hydroxy-β-damascone, and 3,4-epoxy-β-damascone). Besides that, a novel compound, 2-hydroxy-β-damascenone, produced by CYP109E1 was identified. Docking calculations using the crystal structure of CYP109E1 rationalized the experimentally observed regioselective hydroxylation and identified important amino acid residues for statin and terpene binding.
Similar content being viewed by others
Introduction
Bacillus megaterium is a gram-positive soil bacterium with industrial importance. The ability of growth on different low-priced media, the absence of alkaline proteases and endotoxins, the stable maintenance of plasmid vectors, and high protein expression capacity make this microorganism an excellent biotechnological production host (Rygus and Hillen 1992; Vary et al. 2007). It is used for the production of industrially important enzymes as well as polyhydroxybutyrate and vitamin B12 (Biedendieck et al. 2010; Korneli et al. 2013; Kulpreecha et al. 2009; Malten et al. 2005; Stammen et al. 2010). Besides that, the full genome sequencing of two B. megaterium strains (Eppinger et al. 2011) gave rise to the identification of several biotechnologically interesting proteins from this bacterium such as cytochromes P450 (P450s) (Abdulmughni et al. 2017; Brill et al. 2014; Jóźwik et al. 2016; Kiss et al. 2015; Milhim et al. 2016).
P450s are heme-containing external monooxygenases. Besides their essential role in steroid hormone biosynthesis and drug metabolism, several more applications of this enzyme class such as bioremediation and implementation of a great variety of chemical reactions are also described (Sono et al. 1996). The broad spectrum of substrates and the ability to perform diverse synthetically challenging reactions under mild conditions provide a high biotechnological potential of these enzymes (Bernhardt 2006; Bernhardt and Urlacher 2014). P450 family members are widely distributed among different species of life. In contrast to mammalian and plant P450s, microbial ones have the advantages that they are soluble proteins and exhibit higher stability simplifying their handling (Fulco 1991; Urlacher et al. 2004). Three P450s from B. megaterium, the self-sufficient fatty acid hydroxylase CYP102A1 (BM3) and two steroid hydroxylases of the CYP106 family, CYP106A1 and CYP106A2, have been extensively studied and described as attractive biocatalysts for potential biotechnological applications (Berg and Rafter 1981; Bleif et al. 2011; Brill et al. 2014; Janocha et al. 2016; Kiss et al. 2015; Putkaradze et al. 2017; Narhi and Fulco 1987; Schmitz et al. 2014; Urlacher et al. 2006; Virus et al. 2006).
CYP109E1 is a new member of Bacillus P450s, recently identified by our laboratory, but is not fully characterized so far. The crystal structure of CYP109E1 in substrate-free and substrate-bound state has provided first insights into the structural background of substrate binding and activity of this enzyme (Jóźwik et al. 2016). Due to the close phylogenetic distance of CYP109E1 to the steroid hydroxylase CYP106A1, its potential as steroidogenic P450 has been investigated. It has been found that CYP109E1 possesses very low or no catalytic activity towards steroidal substrates except for testosterone and vitamin D3 (Abdulmughni et al. 2017; Jóźwik et al. 2016). To extend the substrate range of CYP109E1, a focused library was screened. The library consisted of steroids, statins, and terpenoids reported to be substrates of some Bacillus P450s of the 109 family (Furuya et al. 2009; Girhard et al. 2010) (Scheme 1).
Statins are a class of powerful, widely used drugs against cardiovascular diseases. They effectively reduce plasma LDL-cholesterol levels and coronary heart disease risk by inhibiting the key enzyme 3-hydroxy-3-methylglutaryl-coenzyme-A (HMG-CoA) reductase in the mevalonate pathway (Brown 2007; Endo and Hasumi 1993; Lamon-Fava 2013). The first discovered statin, the natural product compactin (mevastatin) undergoes a stereoselective hydroxylation at the position C6′ resulting in the formation of pravastatin (6′β-hydroxycompactin). Pravastatin as well as the naturally occurring statin lovastatin and its semi-synthetic derivative simvastatin are efficient drugs. The bioconversion of compactin to pravastatin is one of the successful biotechnological applications of P450-based systems with only few examples of P450s described in the literature capable of performing this reaction (Bernhardt and Urlacher 2014; Matsuoka et al. 1989; McLean et al. 2015; Milhim et al. 2016; Sakaki 2012).
Terpenes and terpenoids are the most diverse class of chemicals occurring mainly in plants. These compounds and their derivatives are important for biotechnology since they are applied in different fields, such as chemical, pharmaceutical, or flavor and fragrance industry (Janocha et al. 2015). The regio- and stereoselective oxyfunctionalization of terpenes and terpenoids is of high interest but remains to be a challenging task for synthetic chemistry. Microbial P450 enzymes represent an effective alternative to chemical synthesis since they possess the ability to oxidize diverse biotechnologically important terpene and terpenoid compounds in a highly stereo- and regioselective manner. However, only a limited number of microbial P450s have been identified as terpene hydroxylases and epoxidases until now (Janocha et al. 2015). The well-studied and effective bacterial terpene hydroxylating-P450s include CYP101A1, CYP108A1, and CYP111A1 from Pseudomonas species having camphor, terpineol, and linalool as their natural substrates, respectively (Katagiri et al. 1968; Peterson et al. 1992; Ullah et al. 1990), as well as enzymes from Novosphingobium aromaticivorans such as CYP101B1 (Bell and Wong 2007; Hall and Bell 2014). Two P450s from B. megaterium (CYP106 family) have been shown to hydroxylate di- and triterpenes (Bleif et al. 2011; Brill et al. 2014; Schmitz et al. 2014).
Here, we report highly regioselective oxidations of three statins (compactin 1, lovastatin 2, and simvastatin 3) as well as seven terpenes, including α-ionone 4, β-ionone 5, nootkatone 6, isolongifolen-9-one 7, α-damascone 8, β-damascone 9, and β-damascenone 10 by CYP109E1. Reconstituted P450 and B. megaterium whole-cell systems were successfully applied for the biotransformation of these substrates. Characterization of the reaction products indicated hydroxylase and epoxidase activity of CYP109E1, producing an important drug, pravastatin, as well as known and novel statin drug metabolites and terpene derivatives.
Materials and methods
Reagents and chemicals
Compactin, lovastatin, simvastatin, medroxyprogesterone acetate, norethisterone, and norethisterone acetate were purchased from TCI chemicals (Eschborn, Germany). (+)-Nootkatone, (−)-isolongifolen-9-one, and α- and β-ionone were purchased from Sigma-Aldrich (St. Louis, MI, USA). α-Damascone, β-damascone, and β-damascenone were provided from Bell Flavors & Fragrances (Leipzig, Germany). Bacterial culture media were purchased from Becton, Dickinson and Company (Franklin Lakes, NJ, USA). All other chemicals and solvents were obtained from standard sources and were of the highest purity available.
Strains and plasmids
The E. coli C43 (DE3) cells (Lucigen, Middleton, WI, USA) transformed with the plasmid pET17b.CYP109E1 (Abdulmughni et al. 2017) were used for the expression of CYP109E1. B. megaterium MS941 cells (Wittchen and Meinhardt 1995) transformed either with the plasmid pSMF2.1.CYP109E1 (Abdulmughni et al. 2017) or the control plasmid pSMF2.1 (Bleif et al. 2011) were used for whole-cell conversions.
Enzyme purification and spectral measurements
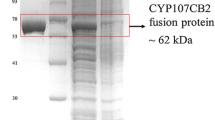
E. coli cells were cultured overnight in Luria-Bertani (LB) medium containing 100 μg/mL ampicillin at 37 °C and 150 rpm in a rotary shaker. Five milliliters of overnight culture was used to inoculate 500 mL Terrific Broth (TB) medium containing 100 μg/mL ampicillin in a 2-L baffled shake flask, and the culture was incubated at 37 °C and 100 rpm. At OD600 of 0.5, the expression of CYP109E1 was induced with 1 mM of isopropyl-thio-β-d-galactopyranoside (IPTG). Simultaneously, 0.5 mM of δ-aminolevulinic acid (δ-ALA) was added to support the heme synthesis. After a 24-h expression, the cells were harvested by centrifugation at 4500×g and stored at − 20 °C until purification of the enzyme. The purification of CYP109E1 was carried out using immobilized metal ion affinity chromatography (TALON™ Resin, Clontech) as described elsewhere (Abdulmughni et al. 2017). Fractions containing CYP109E1 were collected, concentrated, and further purified using size exclusion chromatography (Superdex 75 column, GE Healthcare Life Sciences) in 50 mM potassium phosphate buffer (pH 7.4) with a flow rate of 0.1 mL/min. The purified protein fractions were collected, concentrated by ultrafiltration, and stored at − 20 °C. The concentration of purified CYP109E1 was estimated by carbon monoxide difference spectroscopy using an extinction coefficient (450–490 nm) of 91 mM−1 cm−1 according to the method of Omura and Sato (1964).
Bovine adrenodoxin reductase (AdR) and a truncated form of bovine adrenodoxin (Adx4–108) were expressed and purified as described elsewhere (Sagara et al. 1993; Uhlmann et al. 1992). Protein concentrations were measured spectroscopically using corresponding molar extinction coefficients as described elsewhere (Lisurek et al. 2004).
Spin-state shift and estimation of dissociation constant (K d )
Spin-state shifts were investigated using a double beam spectrophotometer (UV-2101PC, Shimadzu, Japan) and two tandem quartz cuvettes. One chamber of each cuvette contained 10 μM CYP109E1 in 50 mM potassium phosphate buffer (pH 7.4), while the other chamber was filled with buffer alone. The substrates were dissolved in DMSO (5 mM stock solutions) and were added into the chamber with CYP109E1 solution of the sample cuvette while an equal amount of each substrate was also added into the buffer containing a chamber of the reference cuvette. Difference spectra were recorded between 350 and 500 nm. Dissociation constant (K d ) was estimated by titrating each substrate (0–200 μM) until saturation. The data were analyzed by plotting the peak-to-trough differences against the substrate concentrations and fitting them (except for lovastatin and simvastatin) with the hyperbolic function ΔA = ΔA max × [S] / (K d + [S]). Lovastatin and simvastatin exhibited tight binding (K d < 5[E]), and for these compounds, the data were fitted to the tight binding quadratic equation: ΔA = (ΔA max/2[E]) × {(K d + [E] + [S]) − {(K d + [E] + [S])2–4[E][S]}1/2} (Williams and Morrison 1979). ΔA represents the peak-to-trough absorbance difference, ΔA max is the maximum absorbance difference, [E] is the enzyme concentration, and [S] is the substrate concentration. The data processing was done with OriginPro 9.0G software (OriginLab, MA, USA). All K d values represent the mean of three independent measurements with the coefficient of determination (R 2) of 0.99. Spin-state shifts were calculated (to approximately ± 5%) using the ΔA max value of each substrate as a percentage of the maximum expected shift for 10 μM enzyme estimated as described by Luthra et al. (2011).
Measurement of CYP109E1 activity in vitro
The catalytic activity of CYP109E1 towards the selected compounds was investigated using a reconstituted in vitro system consisting of 0.5 or 1 μM CYP109E1, bovine AdR, and Adx4–108 with a molar ratio of 1:2:20. The reactions were carried out in 250 μL of 50 mM potassium phosphate buffer (pH 7.4) with 10% glycerol. For the sufficient electron supply, a NADPH regeneration system containing glucose-6-phosphate-dehydrogenase (1 U), glucose-6-phosphate (5 mM), and MgCl2 (1 mM) was used. The in vitro reactions with 100 μM statin 1–3 (20 mM stock solution in DMSO), 200 μM steroidal 11–13 (20 mM stock solution in DMSO), and terpene 4–10 (50 mM stock solution in ethanol) compounds were started by addition of 1 mM NADPH in 1.5-mL Eppendorf tubes under mixing at 30 °C and 700 rpm and stopped after 15 min by addition of 250 μL of ethyl acetate or chloroform. The reaction mixtures were extracted twice with 500 μL of the organic solvent. The organic phases were combined, evaporated to dryness, and stored at − 20 °C until analysis via either high-performance liquid chromatography (HPLC) or gas chromatography-mass spectrometry (GC-MS).
Whole-cell conversion with CYP109E1 in B. megaterium MS941
The whole-cell conversions were performed in plasmidless B. megaterium MS941 strain transformed with pSMF2.1.CYP109E1 shuttle vector by the method of Barg et al. (2005). As a control of the CYP109E1-based whole-cell biotransformation, MS941 cells transformed with the empty vector pSMF2.1 were used. The main cultures were prepared by inoculating 50 mL of complex medium (24 g/L yeast extract, 12 g/L soytone, 0.5% glycerol (v/v), 2.31 g/L KH2PO4 and 12.5 g/L K2HPO4) with overnight culture (1% of the main culture volume) in 300 mL baffled shake flask and incubated at 37 °C and 160 rpm in a rotary shaker until an OD578 of 0.5 was reached. At this time point, the expression of CYP109E1 was induced by adding 5 g/L xylose. After 24 h expression at 30 °C, the cultures were harvested by centrifugation (4500×g, 4 °C), washed, and resuspended in 50 mM potassium phosphate buffer (pH 7.4) with 2% glycerol. Whole-cell conversions of 100 μM of statins 1–3 and 200 μM of terpenes 4–10 were carried out in 25 mL volume in a rotary shaker at 30 °C and 150 rpm. The conversions of nootkatone 6, isolongifolen-9-one 7, α-ionone 4, β-ionone 5, α-damascone 8, β-damascone 9, and β-damascenone 10 were performed in sealed baffled shake flasks. Substrate stock solutions were prepared by dissolving them in DMSO or ethanol. The biotransformation of each substrate was monitored within 2 h by taking 500-μL culture samples. The samples were extracted with double volume of ethyl acetate, dried, and stored at − 20 °C until HPLC analysis. To obtain sufficient amounts (2–10 mg) of conversion products for the structure elucidation via NMR, the whole-cell reactions were scaled up to 0.5–1 L culture volume. The reactions were stopped after 2 or 4 h and extracted with double volume of ethyl acetate. The organic phases were dried over anhydrous MgSO4, concentrated to dryness using a rotivapor (Büchi R-114), and stored at − 20 °C until purification via HPLC.
Conversion analysis and product isolation via HPLC
The conversion analysis was performed via reversed phase HPLC technique using a Jasco system (a Pu-980 HPLC pump, an AS-950 sampler, an UV-975 UV/Vis detector, a LG-980–02 gradient unit; Jasco, Gross-Umstadt, Germany) and an ec MN Nucleodur C18 (5 μm, 4.0 × 125 mm) column (Macherey-Nagel, Bethlehem, PA, USA). For the purification of conversion products, a preparative ec MN Nucleodur C18 VP (5 μm, 8.0 × 250 mm) column (Macherey-Nagel, Bethlehem, PA, USA) was used. The mobile phase consisted of 10% acetonitrile in water (solvent A) and pure acetonitrile (solvent B). A gradient from 20 to 80% of solvent B was used for the separation of α-ionone 4, β-ionone 5, α-damascone 8, β-damascone 9, β-damascenone 10, compactin 1, and lovastatin 2 conversions, from 20 to 100% for the analysis of nootkatone 6 and isolongifolen-9-one 7 conversions and from 0 to 100% for simvastatin 3, medroxyprogesterone acetate 11, norethisterone 12, and norethisterone acetate 13 conversions. A flow rate of either 1 mL/min (conversion measurements) or 3.5 mL/min (product isolation) and a temperature of 40 °C were set during the analysis of all compounds. The UV detection was accomplished at 236 (statins 1–3), 240 (steroids 11–13, nootkatone 6, and isolongifolen-9-one 7), 228 (α-ionone 4, α-damascone 8, and β-damascone 9), 232 (β-damascenone 10), or 296 nm (β-ionone 5).
Conversion analysis via GC-MS and LC-MS
GC-MS measurements were performed with a system consisting of an AI/AS 3000 autosampler, a DSQ II quadrupole, a Focus GC column oven (Thermo Scientific, Waltham, USA), and a DB-5 column (Agilent) with a length of 25 m, 0.32 mm ID, and 0.52 μm film thickness. The conversions were analyzed as described elsewhere (Litzenburger and Bernhardt 2016).
Analytical HPLC-MS was performed on a Thermo Dionex UltiMate 3000 HPLC using NUCLEOSHELL RP 18plus (100 × 4.6 mm, 2.7 μm) column (Macherey-Nagel, Bethlehem, PA, USA) coupled to a Bruker AmaZon SL ESI-MS system (Bruker, Billerica, MA, USA). The mobile phase consisted of water with 0.1% formic acid (solvent A) and pure acetonitrile with 0.1% formic acid (solvent B). A gradient from 5 to 95% of solvent B was used for the separation. Mass spectra were acquired in positive mode ranging from 100 to 600 m/z using UltraScan mode.
Structure elucidation by NMR spectroscopy
NMR spectra were recorded on a Bruker (Rheinstetten, Germany) DRX 500 NMR spectrometer. A combination of 1H, 13C, 1H,1H-COSY, HSQC, and HMBC experiments was used for structure elucidation. All chemical shifts are relative to CHCl3 (δ = 7.24) or CDCl3 (δ = 77.00) using the standard δ notion in parts per million (ppm).
Molecular docking
For the docking experiments, the recently solved crystal structure of CYP109E1 was used (Jóźwik et al. 2016). Two structurally different substrates, the statin compactin 1 and the norisoprenoid compound α-ionone 4, were docked into the active site of CYP109E1 in its closed conformation (PDB: 5L94) using AutoDock 4.0 (Morris et al. 2009). The Windows version 1.5.6 of AutoDock Tools was used to compute Kollman charges for the enzyme and Gasteiger-Marsili charges for the ligands (Sanner 1999). A partial charge of + 0.400e was assigned manually to the heme iron, which corresponds to Fe(II) that was compensated by adjusting the partial charges of the ligating nitrogen atoms to − 0.348e. While the protein was kept as rigid, the flexible bonds of the ligands were assigned automatically and verified by manual inspection. A cubic grid box with a size of 52 Å × 50 Å × 48 Å was fixed above the heme moiety to cover the whole active site of the enzyme. The spacing between grid points was 0.375 Å. Two hundred docking runs (for each substrate) were carried out applying the Lamarckian genetic algorithm using default parameter settings. The binding conformations were analyzed according to estimated binding energies and distances of the target carbon atom to the heme iron.
Results
Substrate binding to CYP109E1
To screen CYP109E1 for potential new substrates, a library of 13 compounds was used. The library contained the steroids norethisterone 12 and medroxyprogesterone acetate 11, compactin 1, and the terpenes α-ionone 4, β-ionone 5, and nootkatone 6, which previously have been identified as substrates of the CYP109 family (Furuya et al. 2009; Girhard et al. 2010), as well as related compounds with biotechnological or pharmaceutical impact (Scheme 1). First, the potential substrates were investigated for their ability to bind to CYP109E1. This was done by monitoring the high-spin shift induction using difference spectroscopy. When a P450 substrate displaces the axial water ligand, it changes the spin-state of the iron atom from low-spin to high-spin (Schenkman and Jansson 1998). Spectroscopically, this transition is reflected by a shift in the absorption maximum from around 420 to 390 nm (type I difference spectrum). Previously, it has been reported that the spin-shift is not necessary for the catalytic activity of a P450 (Girhard et al. 2010; Simgen et al. 2000). On the other hand, a very large type I spin-shift observed for many bacterial P450s (Bell and Wong 2007) indicating the ability of its substrates to displace almost all water molecules in the active site results in an effective electron transfer and high catalytic activities by substrate-gating mechanism (Sligar and Gunsalus 1976; Fisher and Sligar 1985; Honeychurch et al. 1999, Poulos and Raag 1992). Therefore, this spectroscopic method remains to be well suitable for screening of potential P450 substrates as well as inhibitors (Khatri et al. 2016; Schmitz et al. 2012). All here tested compounds, except for steroids 11–13, were able to induce a type I spin-shift of CYP109E1, which allowed us to investigate the substrate binding spectroscopically (Fig. 1 and S1) and to determine dissociation constants (K d ). Statin compounds 1–3 showed tighter binding to P450 than terpene substrates 4–10 (Table 1). Our results indicate that simvastatin 3 binds tighter to CYP109E1 than the other statins. The K d of simvastatin 3 [K d = 4.7 μM] was shown to be approximately 18 times lower than that of compactin 1 [K d = 84 μM] and two times lower than that of lovastatin 2 [Kd = 9.6 μM] (Fig. 1 and S1). The strongest binding to CYP109E1 among terpenes 4–10 was observed for β-ionone 5 [K d = 91 μM] and the weakest for isolongifolen-9-one 7 [K d = 216 μM] (Fig. S1). Interestingly, α-ionone 4 (51%) and simvastatin 3 (43%) were able to induce the highest spin-state shifts among the investigated substrates, whereas lovastatin 2 shifted the state to a lesser extent (11%).
The absorbance changes plotted against the corresponding concentrations of compactin 1 (a) and α-ionone 4 (b) titrated to CYP109E1 as described in the “Material and methods” section. The mean values were fitted by hyperbolic regression, and the K d value was calculated. The inset shows type I spectral shifts induced by the binding of the increasing amount of the substrate to CYP109E1. Arrows show the direction of spectral changes at increasing substrate concentrations
Biotransformation of medroxyprogesterone acetate, norethisterone, and norethisterone acetate
Although the tested steroids 11–13 did not show any spectral shift when incubated with CYP109E1, their potential biotransformation was investigated in vitro. As observed previously with deoxycorticosterone and testosterone as substrates of CYP106A2 and CYP109B1, respectively, a type I spectral shift is not a necessary prerequisite for bioconversion (Girhard et al. 2010; Simgen et al. 2000). The steroidal substrates medroxyprogesterone acetate 11, norethisterone 12, and norethisterone acetate 13 were thus investigated using a reconstituted P450 system with AdR and Adx4–108 as redox partners. CYP109E1 did not show any activity towards these compounds (data not shown), and they were, therefore, not further tested with the CYP109E1 based whole-cell system in vivo.
Biotransformation of statin substrates
The three statins, compactin 1, lovastatin 2, and simvastatin 3, identified as CYP109E1 ligands by difference spectroscopy, were further tested as potential substrates for CYP109E1 in vitro. The activity of CYP109E1 was reconstituted with bovine AdR and truncated Adx4–108 as described in the “Material and methods” section. CYP109E1 was found to convert all three statin compounds 1–3 resulting in one major reaction product 14, 15 for compactin 1 and lovastatin 2, respectively (Fig. 2a, c) and two products 16, 17 for simvastatin 3 conversion (Fig. 2e). The selectivity of compactin and lovastatin conversions (77 and 94%) was found to be remarkably higher compared to that of simvastatin (Table S1). Interestingly, the conversion ratio of CYP109E1 towards compactin 1 was much lower compared with the other statins resulting in only 11% conversion after 15 min compared with lovastatin 2 (63%) and simvastatin 3 (72%) (Fig. 2a, c, e and S2).
HPLC chromatograms of in vitro (a, c, e) and in vivo (b, d, f) conversion of statins 1–3. 14, 15, 16, and 17 represent products of compactin 1, lovastatin 2, and simvastatin 3 conversion by CYP109E1, respectively, which have been isolated and characterized. The minor products are labeled with “x” (masses are provided in Table S1) and impurities with asterisk
To prepare and isolate higher amounts of compactin 1, lovastatin 2, and simvastatin 3 conversion products for the analysis via NMR spectroscopy, B. megaterium MS941 whole cells transformed with pSMF2.1.CYP109E1 were used. After in vivo conversion of statins, the main products of the compactin 1 and lovastatin 2 conversion (14 and 15, Fig. 2b, d) as well as two products of simvastatin 3 conversion (16 and 17, Fig. 2f) have been purified in sufficient amounts. NMR analysis (data S7) revealed that CYP109E1 is able to hydroxylate compactin 1 and lovastatin 2 at position C6′β resulting in the formation of pravastatin 14 and 6′β-hydroxy-lovastatin 15, respectively (Scheme 2). The simvastatin 3 metabolites were identified as 3′α-hydroxy-simvastatin 16 and 4″-hydroxy-simvastatin 17. The B. megaterium MS941 strain containing the pSMF2.1 vector, which was used as a negative control for the CYP109E1-dependent conversions, showed no activity towards the investigated substrates (data not shown).
Biotransformation of terpene substrates
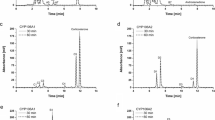
All tested terpenes 4–10 showed type I binding spectra, and the reactions of CYP109E1 with these compounds were further characterized. Similar to statins 1–3, the activity of CYP109E1 towards the potential terpene substrates α-ionone 4, β-ionone 5, nootkatone 6, isolongifolen-9-one 7, α-damascone 8, β-damascone 9, and β-damascenone 10 was initially tested using the reconstituted system. CYP109E1 showed high conversion within 15 min under in vitro conditions (95% of α-ionone 4, 96% of β-ionone 5, 70% of nootkatone 6, 91% of isolongifolen-9-one 7, 72% of α-damascone 8, 70% of β-damascone 9 and β-damascenone 10) (Fig. S2). The reaction selectivity except for β-damascenone conversion was high (77–93%) yielding one main product (Table S1). All these substrates were also successfully converted by B. megaterium cells overexpressing CYP109E1, and the product patterns and selectivities were similar compared to those in vitro. The main products of α-ionone 4, β-ionone 5, α-damascone 8, and β-damascone 9 were compared with authentic standards by GC-MS (Litzenburger and Bernhardt 2016) and identified as 3-hydroxy-α-ionone 18, 4-hydroxy-β-ionone 19, 3-hydroxy-α-damascone 23, and 4-hydroxy-β-damascone 24, respectively (Fig. 3a, b, e, f and S3–S6). Moreover, 3-hydroxy-α-ionone 18 and 3-hydroxy-α-damascone 23 were found to be 3,6-trans-products. The conversion products of nootkatone 6, isolongifolen-9-one 7, and β-damascenone 10 were isolated via HPLC and elucidated by NMR spectroscopy. The products of nootkatone 6 and isolongifolen-9-one 7 conversion were identified as 11(R),12-epoxy-nootkatone 20, 11(S),12-epoxy-nootkatone 21, and 4(R)-hydroxy-isolongifolen-9-one 22, respectively (Fig. 3c, d). Products of in vivo conversion of β-damascenone 10 were found to be 3,4-dihydroxy-β-damascone 27, 2-hydroxy-β-damascenone 25, and 3,4-epoxy-β-damascone 26 (Fig. 4a). The control strain B. megaterium MS941 containing only the pSMF2.1 vector showed low conversion of the investigated terpene substrates 4–10 after 2 h (data not shown), most probably due to the activity of CYP109E1 encoded in the bacterial genome.
HPLC chromatograms of in vivo conversions of α-ionone 4 (a), β-ionone 5 (b), nootkatone 6 (c), isolongifolen-9-one 7 (d), α-damascone 8 (e), and β-damascone 9 (f) with the identified products shown close to the peaks. Masses of the minor products detected in vitro as well as in vivo are provided in Table S1
HPLC chromatogram of β-damascenone 10 conversion by the B. megaterium MS941 cells overexpressing CYP109E1 (a) and time-dependent in vivo conversion within 4 h (b). The minor products are labeled with “x” (masses are provided in Table S1) and impurities with asterisk
Interestingly, the formation of 3,4-dihydroxy-β-damascone 27 was observed in vivo (Fig. 4a), whereas no corresponding peak (Fig. 4a, t R = 2.8 min) was detectable in vitro using the reconstituted CYP109E1-based system (data not shown). For further characterization, the in vivo conversion was monitored over time. The HPLC results showed significant changes in product distribution within 4 h. After 1 h of conversion, 3,4-epoxy-β-damascone 26 and 3,4-dihydroxy-β-damascone 27 were found to be the major and minor products with 34 and 8% of total shares, respectively. The amount of 3,4-dihydroxy-β-damascone 27 increased up to 45% after 4 h of conversion, and the increase was correlated with a decrease of 3,4-epoxy-β-damascone 26 (Fig. 4b). Thus, the time-dependent data suggested that the double hydroxylated product is formed from 3,4-epoxy-β-damascone 26 by a CYP109E1-independent reaction.
Molecular docking
The crystal structure of CYP109E1 in its closed form was used to investigate the enzyme-substrate interaction and observed regioselectivity of hydroxylation for two of the novel substrates, compactin 1 and α-ionone 4, by docking of these structurally different substances into the active site of CYP109E1. Both substrates appeared in docking positions allowing hydroxylation at experimentally identified positions, C6′ (distance ~ 4.4 Å) for compactin 1 and C3 (distance ~ 4 Å) for α-ionone 4, respectively. Eight amino acid residues including Ile85, Leu238, Ile241, Ala242, Thr246, Ala291, Leu292, and His293 were found to form the binding pocket of CYP109E1 with compactin 1 whereas only six (Arg69, Ile85, Ala242, Val289, Leu292, His293) were predicted to interact with the smaller substrate α-ionone 4 (Fig. 5). The results predicted that both substrates were bound by van der Waals forces and hydrophobic interactions. Active site residues with predicted hydrophobic interactions with both substrates are Ile85 (BC-loop, SRS1), Ala242 (I-helix, SRS4), and Leu292 (K-β5-loop, SRS5). Considering the electrostatic interactions with the substrates, only His293 was predicted to form a hydrogen bond with compactin 1 and α-ionone 4 (Fig. 5).
Docking orientations of compactin 1 (a) and α-ionone 4 (b) shown in yellow in the active site of CYP109E1 capable of hydroxylation at C6′ and C3, respectively. The distance of the corresponding carbon atom from the heme iron is given in angstrom. Amino acids forming the active site in the presence of each substrate are shown and named
Discussion
Various P450s from B. megaterium attract interest due to their ability to perform the biotransformation of a broad spectrum of highly interesting compounds such as steroids (Berg and Rafter 1981; Kiss et al. 2015; Putkaradze et al. 2017; Rauschenbach et al. 1993), di- and triterpenes (Bleif et al. 2011; Brill et al. 2014; Schmitz et al. 2014), as well as sesquiterpenes (Sowden et al. 2005), fatty acids, amides, and alcohols (Miura and Fulco 1975). CYP109E1 is a newly identified member of this group, and therefore, elucidation of its substrate range is of great interest. During the initial study on CYP109E1, the crystal structure was solved revealing a highly dynamic active site (Jóźwik et al. 2016). Phylogenetically, CYP109E1 was found to be related to the steroid hydroxylase CYP106A1 and, therefore, a focused library of important steroids was tested as its potential substrates. The study revealed that among the 13 tested steroidal compounds, only testosterone was converted by CYP109E1 resulting in the corresponding 16β-hydroxy product (Jóźwik et al. 2016). Another important compound which was recently identified as substrate of CYP109E1 is the secosteroid vitamin D3 (Abdulmughni et al. 2017). The enzyme showed 25- and 24-hydroxylase activities towards vitamin D3 generating the valuable compound 25-hydroxy vitamin D3 and two new metabolites hydroxylated at position C24(S) and C25 (Abdulmughni et al. 2017).
In order to further characterize CYP109E1, we aimed to extend the substrate spectrum by investigating its activity towards a focused library of biotechnologically valuable compounds. The enzyme showed no activity for the three tested steroidal compounds 11–13, whereas several statins 1–3 and terpenes 4–10 were successfully converted (Scheme 1). Compactin 1, lovastatin 2, and simvastatin 3, identified as novel substrates of CYP109E1 during our study, belong to the group of statins, effective pharmaceutical agents widely used against lipid disorders. In several studies, a broad range of pleiotropic effects of statins was also described proposing their application for the treatment of inflammatory and neurological diseases as well as tumors (Gazzerro et al. 2012). All three compounds were able to shift the heme iron of CYP109E1 into the high-spin state. Dissociation constants determined by substrate titrations showed strongest binding for simvastatin 3 to CYP109E1, with a K d of 4.7 ± 1.8 μM compared to 9.6 ± 3.5 μM and 84 ± 13 μM for lovastatin 2 and compactin 1, respectively. Investigated statin compounds 1–3 were successfully converted by CYP109E1 into one main product, and the activity of the P450 towards lovastatin 2 and simvastatin 3 was higher than that towards compactin 1 using the heterologous redox partners AdR and Adx4–108 (Fig. 2a, c, e). The CYP109E1-based reactions were further investigated in vivo in a mutant of B. megaterium DSM319, MS941, extensively investigated by our group to generate pharmaceutically important metabolites (Abdulmughni et al. 2017; Kiss et al. 2015; Putkaradze et al. 2017). The observed product patterns of statins in vitro and in vivo (Fig. 2) were very similar and allowed us to scale up the whole-cell reactions to isolate sufficient amounts of the main products for structure identification via NMR spectroscopy. The data revealed that CYP109E1 hydroxylates compactin 1 with a high stereoselectivity at the C6′ position forming the biologically active form of the widely used pharmaceutical pravastatin 14 and this way characterizing CYP109E1 as a novel pravastatin synthase. To the best of our knowledge, so far only two other non-mutated P450 monooxygenases are known being able to convert compactin 1 to active pravastatin variant. The CYP109E1-based whole-cell system produced up to 14 mg/L pravastatin 14 after 4 h of biotransformation which is comparable to the previously described E. coli systems with P450sca-2 (12.9 mg/L within 21 h) (Ba et al. 2013) and CYP107DY1 (13.2 mg/L within 20 h) (Milhim et al. 2016) but provides much higher space-time yield. The activity of this industrially relevant P450 system for pravastatin 14 production might be further optimized by identification of a more effective redox chain, by more suitable production hosts, and by a rational or semi-rational design of the enzyme (Ba et al. 2013; McLean et al. 2015; Milhim et al. 2016). Moreover, to overcome known limitations of cytochrome P450 whole-cell systems on industrial scale, such as substrate solubility, toxicity, and uptake from the cells, expression of CYP109E1 by compactin-producing Penicillium chrysogenum similar to the expression of P450Prava (McLean et al. 2015) might be promising. Besides compactin 1, lovastatin 2 was also converted into the C6′β-hydroxylated product by CYP109E1 while simvastatin 3 was found to be hydroxylated at the C3′α and C4″ positions (Scheme 2). 6′β-Hydroxy-lovastatin 15 and 3′α-hydroxy-simvastatin 16 are human liver metabolites of the statin drugs formed by P450 enzymes, whereas 4″-hydroxy-simvastatin 17 is a synthetic drug derivative. None of these compounds are commercially available although they are highly required in larger quantities for activity and safety studies, particularly due to the reported improved pharmacokinetic properties of some statin metabolites (Kandel et al. 2014) and the known drug-drug interactions of statin pharmaceuticals (Kellick et al. 2014). There are few reports describing the generation of these metabolites, such as chemical synthesis of 3′α-hydroxy-simvastatin 16, electrochemical oxidation, and biotransformation of lovastatin 2 and simvastatin 3 with CYP3A4 or CYP102A1 mutants (Khera and Hu 2013; Kim et al. 2011; Stokker 1994). However, CYP109E1 seems to be a perfect candidate for the production of these metabolites through mild and cost-effective biocatalysis with much higher selectivities and higher product yields compared to previously established methods. Moreover, CYP109E1 represents the first enzyme that is able to introduce a hydroxyl group at the C4″ position into the simvastatin molecule.
In addition to statins 1–3, seven terpene compounds 4–10 were identified as novel substrates of CYP109E1 and their biotransformation was further investigated. Terpene and terpenoid hydroxylation and epoxidation by microbial P450 enzymes have been reported in the literature (Çelik et al. 2005; Hall and Bell 2014; Litzenburger and Bernhardt 2016; Schifrin et al. 2015). However, the high demand of valuable terpene derivatives and their mostly unselective and low-yield production via chemical synthesis makes exploration of new effective biocatalysts for terpene oxyfunctionalization highly important. Here, α-ionone 4, β-ionone 5, α-damascone 8, β-damascone 9, β-damascenone 10, nootkatone 6, and isolongifolen-9-one 7 were found to serve as substrates for CYP109E1 identifying this P450 as a novel terpene hydroxylase and epoxidase. All compounds were able to bind to the enzyme and induce a type I difference spectrum of CYP109E1. Concerning dissociation constants, terpenes 4–10 showed weaker binding to the active site of CYP109E1 than statin substrates 1–3. However, they have been converted by CYP109E1 with high efficiencies and regioselectivities in vitro as well as in vivo. The reaction products of α-ionone 4, β-ionone 5, α-damascone 8, and β-damascone 9 were identified as 3-hydroxy-α-ionone 18, 4-hydroxy-β-ionone 19, 3-hydroxy-α-damascone 23, and 4-hydroxy-β-damascone 24, respectively. Thus, CYP109E1 hydroxylates preferably at allylic positions which are energetically favorable and also preferred by other P450s converting these compounds (Çelik et al. 2005; Hall and Bell 2014; Khatri et al. 2010; Litzenburger and Bernhardt 2016). The resulting hydroxylated ionones and damascones are important compounds which might be used as precursors or building blocks in chemical synthesis of fragrance constituents due to their floral and fruity scents as well as in bioassays due to their bioactive properties. For example, the 4-hydroxy derivative of β-ionone 5 has been reported to be a versatile synthon (More and Bhat 2013) whereas 4-hydroxy-β-damascone 24 was found to possess very strong antifeedant properties against lesser mealworm Alphitobius diaperinus (Gliszczyńska et al. 2016). The CYP109E1-based B. megaterium system used in our study is very suitable for the production of these metabolites. To compare it with the previously established P450-based production systems, whole-cell experiments using higher substrate concentrations and longer incubation times are necessary. However, based on the observed fast conversion of up to 200 μM substrate within 2 h under non-optimized conditions, the whole-cell system used in our study seems to be efficient for the production of these valuable metabolites. The NMR data of nootkatone 6 and β-damascenone 10 conversion products revealed for the first time the epoxidation activity of CYP109E1 resulting in the formation of 11(R),12-epoxy-nootkatone 20, 11(S),12-epoxy-nootkatone 21, and 3,4-epoxy-β-damascone 26, respectively. 3,4-Epoxy-β-damascone 26 has been previously reported to be a conversion product of β-damascone 9 by CYP101C1 from N. aromaticivorans DSM12444 (Ma et al. 2011), whereas 11,12-epoxy-nootkatone has been described to be produced by CYP102A1 mutants as well as unknown fungal proteins. It showed antiproliferative activity against leukemia cell line HL-60 (Gliszczyńska et al. 2011; Sowden et al. 2005). In our studies, 3,4-epoxy-β-damascone 26 was further metabolized in vivo to 3,4-dihydroxy-β-damascone 27 by a CYP109E1-independent reaction proposed to be catalyzed by an unknown epoxide hydrolase from the DSM319 as described for other B. megaterium strains (Michaels et al. 1980; Tang et al. 2001; Zhang et al. 2010). Besides 3,4-epoxy-β-damascone 26, a novel compound, 2-hydroxy-β-damascenone 25, was identified as conversion product of CYP109E1. Its properties are unknown and need to be investigated. Isolongifolen-9-one 7 was hydroxylated regio- and stereoselectively at C4 by CYP109E1 yielding 4(R)-hydroxy-isolongofilen-9-one 22. This compound has been described in the literature as conversion product of isolongifolen-9-one 7 by four fungal cultures, and it has been shown to have an inhibitory activity on tyrosinase, the key enzyme for melanin biosynthesis (Choudhary et al. 2003). In another study, the suppressive effect of 4(R)-hydroxy-isolongofilen-9-one 22 against chemical mutagen-induced SOS response in Salmonella typhimurium TA1535/pSK1002 has been reported (Sakata et al. 2010).
In addition to the in vitro and in vivo characterization of CYP109E1, in silico experiments were performed using the crystal structure of this enzyme in order to predict residues responsible for substrate binding and to understand the structural basis of the observed hydroxylation regioselectivity. The molecular docking of the two selected structurally different novel substrates, compactin 1 and α-ionone 4, into CYP109E1 indicated the presence of common as well as specific residues interacting with each substrate. The obtained docking orientations of these substrates in the active site supported experimental results, 6′β-hydroxylation of compactin 1 and 3-hydroxylation in the cyclohexene ring of α-ionone 4 (Fig. 5).
Taken together, we were able to characterize CYP109E1 from B. megaterium for the first time as a novel and highly regioselective statin and terpene hydroxylase as well as terpene epoxidase. Thus, CYP109E1 might be used to generate pharmaceutically and biotechnologically interesting compounds, such as the drug pravastatin 14 and the human statin drug metabolites 6′β-hydroxy-lovastatin 15 and 3′α-hydroxy-simvastatin 16 as well as several valuable terpene derivatives. Besides that, 4″-hydroxy-simvastatin 17 and a novel compound, 2-hydroxy-β-damascenone 25, were obtained using CYP109E1 as biocatalyst, and are now available for further investigations. Finally, our B. megaterium whole-cell system was successfully utilized for the production of the statin and terpene metabolites in milligram scale. The established CYP109E1 system is a good candidate for improvement towards industrial scale.
References
Abdulmughni A, Jóźwik IK, Putkaradze N, Brill E, Zapp J, Thunnissen AMWH, Hannemann F, Bernhardt R (2017) Characterization of cytochrome P450 CYP109E1 from Bacillus megaterium as a novel vitamin D3 hydroxylase. J Biotechnol 243:38–47. https://doi.org/10.1016/j.jbiotec.2016.12.023
Ba L, Li P, Zhang H, Duan Y, Lin Z (2013) Semi-rational engineering of cytochrome P450sca-2 in a hybrid system for enhanced catalytic activity: insights into the important role of electron transfer. Biotechnol Bioeng 110:2815–2825. https://doi.org/10.1002/bit.24960
Barg H, Malten M, Jahn M, Jahn D (2005) Protein and vitamin production in Bacillus megaterium. In: Barredo JL (ed) Microbial processes and products. Humana Press, New York, pp 205–223
Bell SG, Wong LL (2007) P450 enzymes from the bacterium Novosphingobium aromaticivorans. Biochem Biophys Res Commun 360(3):666–672. https://doi.org/10.1016/j.bbrc.2007.06.119
Berg A, Rafter JJ (1981) Studies on the substrate specificity and inducibility of cytochrome P-450meg. Biochem J 196:781–786
Bernhardt R (2006) Cytochromes P450 as versatile biocatalysts. J Biotechnol 124:128–145. https://doi.org/10.1016/j.jbiotec.2006.01.026
Bernhardt R, Urlacher VB (2014) Cytochromes P450 as promising catalysts for biotechnological application: chances and limitations. Appl Microbiol Biotechnol 98:6185–6203. https://doi.org/10.1007/s00253-014-5767-7
Biedendieck R, Malten M, Barg H, Bunk B, Martens JH, Deery E, Leech H, Warren MJ, Jahn D (2010) Metabolic engineering of cobalamin (vitamin B12) production in Bacillus megaterium. Microb Biotechnol 3:24–37. https://doi.org/10.1111/j.1751-7915.2009.00125.x
Bleif S, Hannemann F, Zapp J, Hartmann D, Jauch J, Bernhardt R (2011) A new Bacillus megaterium whole-cell catalyst for the hydroxylation of the pentacyclic triterpene 11-keto-β-boswellic acid (KBA) based on a recombinant cytochrome P450 system. Appl Microbiol Biotechnol 93:1135–1146. https://doi.org/10.1007/s00253-011-3467-0
Brill E, Hannemann F, Zapp J, Brüning G, Jauch J, Bernhardt R (2014) A new cytochrome P450 system from Bacillus megaterium DSM319 for the hydroxylation of 11-keto-β-boswellic acid (KBA). Appl Microbiol Biotechnol 98:1701–1717. https://doi.org/10.1007/s00253-013-5029-0
Brown AJ (2007) Cholesterol, statins and cancer. Clin Exp Pharmacol Physiol 34:135–141. https://doi.org/10.1111/j.1440-1681.2007.04565.x
Çelik A, Flitsch SL, Turner NJ (2005) Efficient terpene hydroxylation catalysts based upon P450 enzymes derived from Actinomycetes. Org Biomol Chem 3:2930–2934. https://doi.org/10.1039/b506159h
Choudhary MI, Musharraf SG, Khan MTH, Abdelrahman D, Parvez M, Shaheen F, Rahman A-u (2003) Microbial transformation of isolongifolen-4-one. Helv Chim Acta 86:3450–3460. https://doi.org/10.1002/hlca.200390289
Endo A, Hasumi K (1993) HMG-CoA reductase inhibitors. Nat Prod Rep 10:541–550. https://doi.org/10.1039/NP9931000541
Eppinger M, Bunk B, Johns MA, Edirisinghe JN, Kutumbaka KK, Koenig SSK, Creasy HH, Rosovitz MJ, Riley DR, Daugherty S, Martin M, Elbourne LD, Paulsen I, Biedendieck R, Braun C, Grayburn S, Dhingra S, Lukyanchuk V, Ball B, Ul-Qamar R, Seibel J, Bremer E, Jahn D, Ravel J, Vary PS (2011) Genome sequences of the biotechnologically important Bacillus megaterium strains QM B1551 and DSM319. J Bacteriol 193:4199–4213. https://doi.org/10.1128/JB.00449-11
Fisher MT, Sligar SG (1985) Control of heme protein redox potential and reduction rate: linear free energy relation between potential and ferric spin state equilibrium. J Am Chem Soc 107(17):5018–5019
Fulco AJ (1991) P450BM-3 and other inducible bacterial P450 cytochromes: biochemistry and regulation. Annu Rev Pharmacol Toxicol 31:177–203. https://doi.org/10.1146/annurev.pa.31.040191.001141
Furuya T, Shibata D, Kino K (2009) Phylogenetic analysis of Bacillus P450 monooxygenases and evaluation of their activity towards steroids. Steroids 74:906–912. https://doi.org/10.1016/j.steroids.2009.06.005
Gazzerro P, Proto MC, Gangemi G, Malfitano AM, Ciaglia E, Pisanti S, Santoro A, Laezza C, Bifulco M (2012) Pharmacological actions of statins: a critical appraisal in the management of cancer. Pharmacol Rev 64:102–146. https://doi.org/10.1124/pr.111.004994
Girhard M, Klaus T, Khatri Y, Bernhardt R, Urlacher VB (2010) Characterization of the versatile monooxygenase CYP109B1 from Bacillus subtilis. Appl Microbiol Biotechnol 87:595–607. https://doi.org/10.1007/s00253-010-2472-z
Gliszczyńska A, Łysek A, Janeczko T, Świtalska M, Wietrzyk J, Wawrzeńczyk C (2011) Microbial transformation of (+)-nootkatone and the antiproliferative activity of its metabolites. Bioorg Med Chem 19:2464–2469. https://doi.org/10.1016/j.bmc.2011.01.062
Gliszczyńska A, Gładkowski W, Dancewicz K, Gabryś B, Szczepanik M (2016) Transformation of β-damascone to (+)-(S)-4-hydroxy-β-damascone by fungal strains and its evaluation as a potential insecticide against aphids Myzus persicae and lesser mealworm Alphitobius diaperinus Panzer. Catal Commun 80:39–43. https://doi.org/10.1016/j.catcom.2016.03.018
Hall EA, Bell SG (2014) The efficient and selective biocatalytic oxidation of norisoprenoid and aromatic substrates by CYP101B1 from Novosphingobium aromaticivorans DSM12444. RSC Adv 5:5762–5773. https://doi.org/10.1039/c4ra14010a
Honeychurch MJ, Hill AO, Wong LL (1999) The thermodynamics and kinetics of electron transfer in the cytochrome P450cam enzyme system. FEBS Lett 451(3):351–353
Janocha S, Schmitz D, Bernhardt R (2015) Terpene hydroxylation with microbial cytochrome P450 monooxygenases. Adv Biochem Eng Biotechnol 148:215–250. https://doi.org/10.1007/10_2014_296
Janocha S, Carius Y, Hutter M, Lancaster CRD, Bernhardt R (2016) Crystal structure of CYP106A2 in substrate-free and substrate-bound form. Chembiochem 17(9):852–860. https://doi.org/10.1002/cbic. 201500524
Jóźwik IK, Kiss FM, Gricman Ł, Abdulmughni A, Brill E, Zapp J, Pleiss J, Bernhardt R, Thunnissen AWH (2016) Structural basis of steroid binding and oxidation by the cytochrome P450 CYP109E1 from Bacillus megaterium. FEBS J 283:4128–4148. https://doi.org/10.1111/febs.13911
Kandel SE, Wienkers LC, Lampe JN (2014) Cytochrome P450 enzyme metabolites in lead discovery and development. Annu Rep Med Chem 49:347–359. https://doi.org/10.1016/B978-0-12-800167-7.00022-5
Katagiri M, Ganguli BN, Gunsalus IC (1968) A soluble cytochrome P-450 functional in methylene hydroxylation. J Biol Chem 243(12):3543–3546
Kellick KA, Bottorff M, Toth PP (2014) A clinician’s guide to statin drug-drug interactions. J Clin Lipidol 8:30–46. https://doi.org/10.1016/j.jacl.2014.02.010
Khatri Y, Girhard M, Romankiewicz A, Ringle M, Hannemann F, Urlacher VB, Hutter MC, Bernhardt R (2010) Regioselective hydroxylation of norisoprenoids by CYP109D1 from Sorangium cellulosum So ce56. Appl Microbiol Biotechnol 88:485–495. https://doi.org/10.1007/s00253-010-2756-3
Khatri Y, Ringle M, Lisurek M, von Kries JP, Zapp J, Bernhardt R (2016) Substrate hunting for the myxobacterial CYP260A1 revealed new 1α-hydroxylated products from C-19 steroids. Chembiochem 17(1):90–101. https://doi.org/10.1002/cbic.201500420
Khera S, Hu N (2013) Generation of statin drug metabolites through electrochemical and enzymatic oxidations. Anal Bioanal Chem 405:6009–6018. https://doi.org/10.1007/s00216-013-7021-z
Kim KH, Kang JY, Kim DH, Park SH, Park SH, Kim D, Park KD, Lee YJ, Jung HC, Pan JG, Ahn T, Yun CH (2011) Generation of human chiral metabolites of simvastatin and lovastatin by bacterial CYP102A1 mutants. Drug Metab Dispos 39(1):140–150. https://doi.org/10.1124/dmd.110.036392
Kiss FM, Schmitz D, Zapp J, Dier TKF, Volmer DA, Bernhardt R (2015) Comparison of CYP106A1 and CYP106A2 from Bacillus megaterium—identification of a novel 11-oxidase activity. Appl Microbiol Biotechnol 99:8495–8514. https://doi.org/10.1007/s00253-015-6563-8
Korneli C, Biedendieck R, David F, Jahn D, Wittmann C (2013) High yield production of extracellular recombinant levansucrase by Bacillus megaterium. Appl Microbiol Biotechnol 97:3343–3353. https://doi.org/10.1007/s00253-012-4567-1
Kulpreecha S, Boonruangthavorn A, Meksiriporn B, Thongchul N (2009) Inexpensive fed-batch cultivation for high poly(3-hydroxybutyrate) production by a new isolate of Bacillus megaterium. J Biosci Bioeng 107:240–245. https://doi.org/10.1016/j.jbiosc.2008.10.006
Lamon-Fava S (2013) Statins and lipid metabolism: an update. Curr Opin Lipidol 24:221–226. https://doi.org/10.1097/MOL.0b013e3283613b8b
Lisurek M, Kang MJ, Hartmann RW, Bernhardt R (2004) Identification of monohydroxy progesterones produced by CYP106A2 using comparative HPLC and electrospray ionisation collision-induced dissociation mass spectrometry. Biochem Biophys Res Commun 319:677–682
Litzenburger M, Bernhardt R (2016) Selective oxidation of carotenoid-derived aroma compounds by CYP260B1 and CYP267B1 from Sorangium cellulosum So ce56. Appl Microbiol Biotechnol 100:4447–4457. https://doi.org/10.1007/s00253-015-7269-7
Luthra A, Denisov IG, Sligar SG (2011) Spectroscopic features of cytochrome P450 reaction intermediates. Arch Biochem Biophys 507(1):26–35. https://doi.org/10.1016/j.abb.2010.12.008
Ma M, Bell SG, Yang W, Hao Y, Rees NH, Bartlam M, Zhou W, Wong LL, Rao Z (2011) Structural analysis of CYP101C1 from Novosphingobium aromaticivorans DSM12444. Chembiochem 12:88–99
Malten M, Hollmann R, Deckwer WD, Jahn D (2005) Production and secretion of recombinant Leuconostoc mesenteroides dextransucrase DsrS in Bacillus megaterium. Biotechnol Bioeng 89:206–218. https://doi.org/10.1002/bit.20341
Matsuoka T, Miyakoshi S, Tanzawa K, Nakahara K, Hosobuchi M, Serizawa N (1989) Purification and characterization of cytochrome P-450sca from Streptomyces carbophilus. Eur J Biochem 184:707–713. https://doi.org/10.1111/j.1432-1033.1989.tb15070.x
McLean KJ, Hans M, Meijrink B, van Scheppingen WB, Vollebregt A, Tee KL, van der Laan JM, Leys D, Munro AW, van den Berg MA (2015) Single-step fermentative production of the cholesterol-lowering drug pravastatin via reprogramming of Penicillium chrysogenum. Proc Natl Acad Sci U S A 112:2847–2852. https://doi.org/10.1073/pnas.1419028112
Michaels BC, Ruettinger RT, Fulco AJ (1980) Hydration of 9,10-epoxypalmitic acid by a soluble Enzyme from Bacillus megaterium. Biochem Biophys Res Commun 92(4):1189–1195
Milhim M, Putkaradze N, Abdulmughni A, Kern F, Hartz P, Bernhardt R (2016) Identification of a new plasmid-encoded cytochrome P450 CYP107DY1 from Bacillus megaterium with a catalytic activity towards mevastatin. J Biotechnol 240:68–75. https://doi.org/10.1016/j.jbiotec.2016.11.002
Miura Y, Fulco AJ (1975) ω-1, ω-2 and ω-3 hydroxylation of long-chain fatty acids, amides and alcohols by a soluble enzyme system from Bacillus megaterium. Biochim Biophys Acta 388:305–317
More GP, Bhat SV (2013) Facile lipase catalysed syntheses of (S)-(+)-4-hydroxy-β-ionone and (S)-(+)-4-hydroxy-β-damascone: chiral flavorants and synthons. Tetrahedron Lett 54:4148–4149. https://doi.org/10.1016/j.tetlet.2013.05.089
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Narhi LO, Fulco AJ (1987) Identification and characterization of two functional domains in cytochrome P-450BM-3, a catalytically self-sufficient monooxygenase induced by barbiturates in Bacillus megaterium. J Biol Chem 262:6683–6690
Omura T, Sato R (1964) The carbon monoxide-binding pigment of liver microsomes. I. Evidence for its hemoprotein nature. J Biol Chem 239:2370–2378
Peterson JA, Lu JY, Geisselsoder J, Graham-Lorence S, Carmona C, Witney F, Lorence MC (1992) Cytochrome P-450terp. Isolation and purification ofthe protein and cloning and sequencing of its operon. J Biol Chem 267(20):14193–14203
Poulos TL, Raag R (1992) Cytochrome P450cam: crystallography, oxygen activation, and electron transfer. FASEB J 6(2):674–679
Putkaradze N, Kiss FM, Schmitz D, Zapp J, Hutter MC, Bernhardt R (2017) Biotransformation of prednisone and dexamethasone by cytochrome P450 based systems—identification of new potential drug candidates. J Biotechnol 242:101–110. https://doi.org/10.1016/j.jbiotec.2016.12.011
Rauschenbach R, Isernhagen M, Noeske-Jungblut C, Boidol W, Siewert G (1993) Cloning sequencing and expression of the gene for cytochrome P450meg, the steroid-15β-monooxygenase from Bacillus megaterium ATCC 13368. Mol Gen Genet 241:170–176
Rygus T, Hillen W (1992) Catabolite repression of the xyl operon in Bacillus megaterium. J Bacteriol 174:3049–3055
Sagara Y, Wada A, Takata Y, Waterman MR, Sekimizu K, Horiuchi T (1993) Direct expression of adrenodoxin reductase in Escherichia coli and the functional characterization. Biol Pharm Bull 16:627–630
Sakaki T (2012) Practical application of cytochrome P450. Biol Pharm Bull 35:844–849
Sakata K, Oda Y, Miyazawa M (2010) Suppression of SOS-inducing activity of chemical mutagens by metabolites from microbial transformation of (−)-isolongifolene. J Agric Food Chem 58:2164–2167. https://doi.org/10.1021/jf903651c
Sanner MF (1999) Python: a programming language for software integration and development. J Mol Graph Modell 17(1):57–61
Schenkman JB, Jansson I (1998) Spectral analyses of cytochromes P450. In: Phillips I, Shephard EA (eds) Cytochrome P450 protocols. Humana Press, New York, pp 25–34
Schifrin A, Litzenburger M, Ringle M, Ly TTB, Bernhardt R (2015) New sesquiterpene oxidations with CYP260A1 and CYP264B1 from Sorangium cellulosum So ce56. Chembiochem 16:2624–2632. https://doi.org/10.1002/cbic.201500417
Schmitz D, Zapp J, Bernhardt R (2012) Hydroxylation of the triterpenoid dipterocarpol with CYP106A2 from Bacillus megaterium. FEBS J 279:1663–1674. https://doi.org/10.1111/j.1742-4658.2012. 08503.x
Schmitz D, Zapp J, Bernhardt R (2014) Steroid conversion with CYP106A2—production of pharmaceutically interesting DHEA metabolites. Microb Cell Factories 13:81. https://doi.org/10.1186/1475-2859-13-81
Simgen B, Contzen J, Schwarzer R, Bernhardt R, Jung C (2000) Substrate binding to 15β-hydroxylase (CYP106A2) probed by FT infrared spectroscopic studies of the iron ligand CO stretch vibration. Biochem Biophys Res Commun 269:737–742. https://doi.org/10.1006/bbrc.2000.2348
Sligar SG, Gunsalus IC (1976) A thermodynamic model of regulation: modulation of redox equilibria in camphor monoxygenase. Proc Natl Acad Sci 73(4):1078–1082
Sono M, Roach MP, Coulter ED, Dawson JH (1996) Heme-containing oxygenases. Chem Rev 96:2841–2888
Sowden RJ, Yasmin S, Rees NH, Bell SG, Wong LL (2005) Biotransformation of the sesquiterpene (+)-valencene by cytochrome P450cam and P450BM-3. Org Biomol Chem 3:57–64. https://doi.org/10.1039/b413068e
Stammen S, Müller BK, Korneli C, Biedendieck R, Gamer M, Franco-Lara E, Jahn D (2010) High-yield intra- and extracellular protein production using Bacillus megaterium. Appl Environ Microbiol 76:4037–4046. https://doi.org/10.1128/AEM.00431-10
Stokker GE (1994) Synthesis of the 3′(S)-hydroxy derivative of simvastatin. Bioorg Med Chem Lett 4:1767–1770. https://doi.org/10.1016/S0960-894X(00)80377-7
Tang YF, Xu JH, Ye Q, Schulze B (2001) Biocatalytic preparation of (S)-phenyl glycidyl ether using newly isolated Bacillus megaterium ECU1001. J Mol Catal B Enzym 13:61–68. https://doi.org/10.1016/S1381-1177(00)00230-7
Uhlmann H, Beckert V, Schwarz D, Bernhardt R (1992) Expression of bovine adrenodoxin in E. coli and site-directed mutagenesis of /2FE-2S/ cluster ligands. Biochem Biophys Res Commun 188:1131–1138
Ullah AJ, Murray RI, Bhattacharyya PK, Wagner GC, Gunsalus IC (1990) Protein components of a cytochrome P-450 linalool 8-methyl hydroxylase. J Biol Chem 265(3):1345–1351
Urlacher VB, Lutz-Wahl S, Schmid RD (2004) Microbial P450 enzymes in biotechnology. Appl Microbiol Biotechnol 64:317–325. https://doi.org/10.1007/s00253-003-1514-1
Urlacher VB, Makhsumkhanov A, Schmid RD (2006) Biotransformation of beta-ionone by engineered cytochrome P450 BM-3. Appl Microbiol Biotechnol 70:53–59. https://doi.org/10.1007/s00253-005-0028-4
Vary PS, Biedendieck R, Fuerch T, Meinhardt F, Rohde M, Deckwer WD, Jahn D (2007) Bacillus megaterium—from simple soil bacterium to industrial protein production host. Appl Microbiol Biotechnol 76:957–967. https://doi.org/10.1007/s00253-007-1089-3
Virus C, Lisurek M, Simgen B, Hannemann F, Bernhardt R (2006) Function and engineering of the 15β-hydroxylase CYP106A2. Biochem Soc Trans 34:1215–1218. https://doi.org/10.1042/BST0341215
Williams JW, Morrison JF (1979) The kinetics of reversible tight-binding inhibition. Methods Enzymol 63:437–467
Wittchen KD, Meinhardt F (1995) Inactivation of the major extracellular protease from Bacillus megaterium DSM319 by gene replacement. Appl Microbiol Biotechnol 42:871–877. https://doi.org/10.1007/ BF00191184
Zhang Z, Sheng Y, Jiang K, Wang Z, Zheng Y, Zhu Q (2010) Bio-resolution of glycidyl (o, m, p)-methylphenyl ethers by Bacillus megaterium. Biotechnol Lett 32:513–516. https://doi.org/10.1007/s10529-009-0181-4
Acknowledgements
The authors thank Birgit Heider-Lips for the purification of AdR and Adx4-108, Dr. Josef Zapp for the NMR measurements, and Ghamdan Beshr from the Helmholtz Institute for Pharmaceutical Research Saarland (HIPS) for LC-MS measurements.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 1608 kb)
About this article
Cite this article
Putkaradze, N., Litzenburger, M., Abdulmughni, A. et al. CYP109E1 is a novel versatile statin and terpene oxidase from Bacillus megaterium . Appl Microbiol Biotechnol 101, 8379–8393 (2017). https://doi.org/10.1007/s00253-017-8552-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8552-6