Abstract
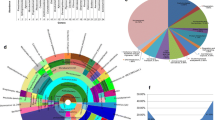
The interior of plants constitutes a unique environment for microorganisms with various organisms inhabiting as endophytes. Unlike subterranean plant parts, aboveground parts are relatively less explored for endophytic microbial diversity. We employed a combination of cultivation and molecular approaches to study the endophytic bacterial diversity in banana shoot-tips. Cultivable bacteria from 20 sucker shoot-tips of cv. Grand Naine included 37 strains under 16 genera and three phyla (Proteobacteria, Actinobacteria, Firmicutes). 16S rRNA gene-ribotyping approach on 799f and 1492r PCR-amplicons to avoid plant organelle sequences was ineffective showing limited bacterial diversity. 16S rRNA metagene profiling targeting the V3-V4 hypervariable region after filtering out the chloroplast (74.2 %), mitochondrial (22.9 %), and unknown sequences (1.1 %) revealed enormous bacterial diversity. Proteobacteria formed the predominant phylum (64 %) succeeded by Firmicutes (12.1 %), Actinobacteria (9.5 %), Bacteroidetes (6.4 %), Planctomycetes, Cyanobacteria, and minor shares (<1 %) of 14 phyla including several candidate phyla besides the domain Euryarchaeota (0.2 %). Microbiome analysis of single shoot-tips through 16S rRNA V3 region profiling showed similar taxonomic richness and diversity and was less affected by plant sequence interferences. DNA extraction kit ominously influenced the phylogenetic diversity. The study has revealed vast diversity of normally uncultivable endophytic bacteria prevailing in banana shoot-tips (20 phyla, 46 classes) with about 2.6 % of the deciphered 269 genera and 1.5 % of the 656 observed species from the same source of shoot-tips attained through cultivation. The predominant genera included several agriculturally important bacteria. The study reveals an immense ecosystem of endophytic bacteria in banana shoot tissues endorsing the earlier documentation of intracellular “Cytobacts” and “Peribacts” with possible roles in plant holobiome and hologenome.




Similar content being viewed by others
Abbreviations
- AP kit:
-
AxyPrep bacterial DNA isolation kit
- FDW:
-
Filter sterilized distilled water post-autoclaving
- PF kit:
-
MO BIO PowerFood microbial DNA isolation kit
- NA:
-
Nutrient agar
- TSA:
-
Trypticasein soy agar
- UC kit:
-
MO BIO UltraClean microbial DNA isolation kit
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Berg G, Grube M, Schloter M, Smalla K (2014) Unraveling the plant microbiome: looking back and future perspectives. Front Microbiol 5:148
Browne HP, Forster SC, Anonye BO et al (2016) Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation. Nature. doi:10.1038/nature17645
Bulgarelli D, Rott M, Schlaeppi K et al (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95
Campisano A, Antonielli L, Pancher M, Yousaf S, Pindo M, Pertot I (2014) Bacterial endophytic communities in the grapevine depend on pest management. PLoS One 9(11), e112763
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Meth 7:335–336
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8:790–803
Chelius MK, Triplett EW (2001) The diversity of archaea and bacteria in association with the roots of Zea mays L. Microb Ecol 41:252–263
Conn VM, Franco CMM (2004) Analysis of the endophytic actinobacterial population in the roots of wheat (Triticum aestivum L.) by terminal restriction fragment length polymorphism and sequencing of 16S rRNA clones. Appl Environ Microbiol 70:1787–1794
de Almeida CV, Andreote FD, Yara R, Tanaka FAO, Azevedo JL, de Almeida M (2009) Bacteriosomes in axenic plants: endophytes as stable endosymbionts. World J Microbiol Biotech 25:1757–1764
DeSantis TZ, Hugenholtz P, Larsen N et al (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Dowd SE, Callaway TR, Wolcott RD et al (2008) Evaluation of the bacterial diversity in the feces of cattle using 16S rDNA bacterial tag-encoded FLX amplicon pyrosequencing (bTEFAP). BMC Microbiol 8:125
Edwards J, Johnson C, Santos-Medellín C et al (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci U S A 112:E911–E920
Hallmann J (2001) Plant interactions with endophytic bacteria. In: Jeger MJ, Spence NJ (eds) Biotic interactions in plant-pathogen associations. CABI Publishing, Oxon, Wallingford, pp 87–119
Hallmann J, Quadt-Hallmann A, Mahaffee WF, Kloepper JW (1997) Bacterial endophytes in agricultural crops. Can J Microbiol 43:895–914
Hardoim PR, van Overbeek LS, Berg G (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79:293–320
Kennedy NA, Walker AW, Berry SH et al (2014) The impact of different DNA extraction kits and laboratories upon the assessment of human gut microbiota composition by 16S rRNA gene sequencing. PLoS One 9, e88982
Köberl M, Dita M, Martinuz A, Staver C, Berg G (2015) Agroforestry leads to shifts within the gammaproteobacterial microbiome of banana plants cultivated in Central America. Front Microbiol 6:91
Knief C (2014) Analysis of plant microbe interactions in the era of next generation sequencing technologies. Front Plant Sci 5:216
Lebeis SL, Paredes SH, Lundberg DS et al (2015) Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349:860–864
Lundberg DS, Lebeis SL, Paredes SH et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90
Manter DK, Delgado JA, Holm DG, Stong RA (2010) Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microbial Ecol 60:157–166
McDonald D, Price MN, Goodrich J et al (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISMEJ 6:610–618
Nogales A, Nobre T, Valadas V et al. (2015) Can functional hologenomics aid tackling current challenges in plant breeding? Briefings Funct Genomics elv030 (doi: 10.1093/bfgp/elv030)
Peiffer JA, Spor A, Koren O et al (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci U S A 110:6548–6553
Pinto C, Pinho D, Sousa S, Pinheiro M, Egas C, Gomes AC (2014) Unravelling the diversity of grapevine microbiome. PLoS One 9, e85622
Pirttilä AM, Laukkanen H, Pospiech H, Myllyla R, Hohtola A (2000) Detection of intracellular bacteria in the buds of Scotch pine (Pinus sylvestris L.) by in situ hybridization. Appl Environ Microbiol 66:3073–3077
Podolich O, Ardanov P, Zaets I, Pirttilä AM, Kozyrovska N (2015) Reviving of the endophytic bacterial community as a putative mechanism of plant resistance. Plant Soil 388:367–377
Qin J, Li R, Raes J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65
Reiter B, Sessitsch A (2006) Bacterial endophytes of the wildflower Crocus albiflorus analyzed by characterization of isolates and by a cultivation-independent approach. Can J Microbiol 52:140–149
Rosenblueth M, Martínez-Romero E (2006) Bacterial endophytes and their interactions with hosts. Mol Plant Microbe In 8:827–837
Rossmann B, Müller H, Smalla K et al (2012) Banana-associated microbial communities in Uganda are highly diverse but dominated by Enterobacteriaceae. Appl Environ Microbiol 78:4933–4941
Schlaeppi K, Dombrowski N, Oter RG, Ver Loren van Themaat E, Schulze-Lefert P (2014) Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc Natl Acad Sci U S A 111:585–592
Sekhar AC, Thomas P (2015) Isolation and identification of shoot-tip associated endophytic bacteria from banana cv. Grand Naine and testing for antagonistic activity against Fusarium oxysporum f. sp. cubense. Am J Plant Sci 6:943–954
Sessitsch A, Hardoim P, Döring J et al (2012) Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol Plant Microbe In 25:28–36
Singh HP, Uma S, Selvarajan R, Karihaloo JL (2011) Micropropagation for Production of Quality Banana Planting Material in AsiaPacific. Asia-Pacific Consortium on Agricultural Biotechnology, New Delhi, India
Sievers F, Wilm A, Dineen D et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Soares MA, Li HY, Kowalski KP, Bergen M, Torres MS, White JF (2016) Functional role of bacteria from invasive Phragmites australis in promotion of host growth. Microb Ecol 72:407–417
Sun L, Qiu F, Zhang X, Dai X, Dong X, Song W (2008) Endophytic bacterial diversity in rice (Oryza sativa L.) roots estimated by 16S rDNA sequence analysis. Microb Ecol 55:415–424
Thomas P, Reddy MK (2013) Microscopic elucidation of abundant endophytic bacteria colonizing the cell wall–plasma membrane peri-space in the shoot-tip tissue of banana. AoB PLANTS 5:plt011
Thomas P, Sekhar AC (2014) Live cell imaging reveals extensive intracellular cytoplasmic colonization of banana by normally non-cultivable endophytic bacteria. AoB PLANTS 6:plu002
Thomas P, Sekhar AC (2016) Effects due to rhizospheric soil application of an antagonistic bacterial endophyte on native bacterial community and its survival in soil: a case study with Pseudomonas aeruginosa from banana. Front Microbiol 7:493
Thomas P, Soly TA (2009) Endophytic bacteria associated with growing shoot tips of banana (Musa sp.) cv. Grand Naine and the affinity of endophytes to the host. Microb Ecol 58:952–964
Thomas P, Kumari S, Swarna GK, Gowda TKS (2007) Papaya shoot tip associated endophytic bacteria isolated from in vitro cultures and host-endophyte interaction in vitro and in vivo. Can J Microbiol 53:380–390
Thomas P, Kumari S, Swarna GK, Prakash DP, Dinesh MR (2007) Ubiquitous presence of fastidious endophytic bacteria in field shoots and index-negative apparently clean shoot-tip cultures of papaya. Plant Cell Rep 26:1491–1499
Thomas P, Mujawar MM, Sekhar AC, Upreti R (2013) Physical impaction injury effects on bacterial cells during spread-plating influenced by cell characteristics of the organisms. J Appl Microbiol 116:911–922
Thomas P, Sekhar AC, Upreti R, Mujawar MM, Pasha SS (2015) Optimization of single plate-serial dilution spotting (SP-SDS) with sample anchoring as an assured method for bacterial and yeast cfu enumeration and single colony isolation from diverse samples. Biotech Rep 8:45–55
Thomas P, Swarna GK, Patil P, Rawal RD (2008) Ubiquitous presence of normally non-culturable endophytic bacteria in field shoot-tips of banana and their gradual activation to quiescent cultivable form in tissue cultures. Plant Cell Tiss Org Cult 93:39–54
Thomas P, Swarna GK, Roy PK, Patil P (2008) Identification of culturable and originally non-culturable endophytic bacteria isolated from shoot tip cultures of banana cv. Grand Naine. Plant Cell Tiss Org Cult 93:55–63
Tkacz A, Poole P (2015) Role of root microbiota in plant productivity. J Exp Bot 66:2167–2175
Tkacz A, Cheema J, Chandra G, Grant A, Poole PS (2015) Stability and succession of the rhizosphere microbiota depends upon plant type and soil composition. ISMEJ 9:2349–2359
Truyens S, Weyens N, Cuypers A, Vangronsveld J (2014) Bacterial seed endophytes: genera, vertical transmission and interaction with plants. Environ Microbiol Rep 7:40–50
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14:209
Upreti R, Thomas P (2015) Root-associated bacterial endophytes from Ralstonia solanacearum resistant and susceptible tomato cultivars and their pathogen antagonistic effects. Front Microbiol 6:255
Vandenkoornhuyse P, Quaiser A, Duhamel M, Le Van A, Dufresne A (2015) The importance of the microbiome of the plant holobiont. New Phytol 206:1196–1206
Walker AW, Martin JC, Scott P, Parkhill J, Flint HJ, Scott KP (2015) 16S rRNA gene-based profiling of the human infant gut microbiota is strongly influenced by sample processing and PCR primer choice. Microbiome 3:26
Acknowledgments
The study was funded partly under the ICAR-AMAAS Net-work project “Exploration of the endophytic microbial diversity in horticultural crops through metagenomics and cultivation” funded through the National Bureau of Agriculturally Important Microorganisms, Mau Nath Bhanjan, Uttar Pradesh, India, and partly by ICAR-Indian Institute of Horticultural Research, Bengaluru, India. The authors thank Dr. T. P. Rajendran, Former Acting Director, National Institute of Biotic Stress Management, Raipur, India, for the critical reading of the manuscript and the suggestions. The supports provided by Bipul Banerjee, Srinivas Vudathala, Chandan Badapanda (Xcelris Labs), Beena, P. S., Rajadurai C. P. and Jubina Benny (SciGenom Labs) during the metagene profiling studies are gratefully acknowledged. Thanks to Sadiq Pasha and N. Shivarudriah for the technical help during the conduct of some of the experiments. The publication bears ICAR-IIHR contribution No. 97/2015.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Table S1
Categorization of 100 clones derived from the two batches of 10 shoot-tips each of banana as per the ribotyping pattern and the suggested identity of the organisms (DOC 73 kb)
Table S2
16S rRNA metagene profiling data for MG07 and MG08 samples derived through Illumina MiSeq and phylogenetic distribution of OTUs as per direct QIIME analysis (QIIME-I) (DOC 61 kb)
Table S3
16S rRNA metagenome data for MG07 and MG08 samples in QIIME analysis-II excluding chloroplast, mitochondrial and unassigned reads showing the distribution of reads and OTUs (DOC 59 kb)
Table S4
Metagenome data for MG33-V3 and MG34-V3 samples derived through Illumina MiSeq and phylogenetic distribution of OTUs as per direct QIIME analysis (QIIME-I) of 16S rRNA V3 sequence data (DOC 55 kb)
Fig. S1
Phylogenetic distribution of cultivable endophytic bacterial strains isolated from banana ‘Grand Naine’ suckers in two batches (a and b) of 10 suckers; ×n indicates the number of strains (DOC 151 kb)
Fig. S2
OTU table heat map showing taxonomy assignment for each OTU. The OTU heat map displays raw OTU counts per sample where the counts are coloured based on the contribution of each OTU to the total OTU count present in that sample (blue: contributes low percentage of OTUs to sample; red: contributes high percentage of OTUs) (DOC 98 kb)
Fig. S3
Rarefaction plot of metagenome sequences of banana shoot-tip DNA derived through PowerFood (MG07) and AxyPrep (MG08) DNA isolation kits after QIIME-II analysis (DOC 73 kb)
Data set S1 (Excel file)
Taxonomic distribution of OTUs in MG07 sample (V3-V4 profiling) at phylum, class, order, family and genus levels as per QIIME-II analysis (XLS 71 kb)
Data set S2 (Excel file)
Taxonomic distribution of OTUs in MG08 sample (V3-V4 profiling) at phylum, class, order, family and genus levels as per QIIME-II analysis (XLS 39 kb)
Data set S3 (Excel file)
Taxonomic distribution as per OTUs and reads in MG33 and MG34 samples (V3 profiling) at genus level as per QIIME analysis-II (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Thomas, P., Sekhar, A.C. Cultivation Versus Molecular Analysis of Banana (Musa sp.) Shoot-Tip Tissue Reveals Enormous Diversity of Normally Uncultivable Endophytic Bacteria. Microb Ecol 73, 885–899 (2017). https://doi.org/10.1007/s00248-016-0877-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0877-7