Abstract
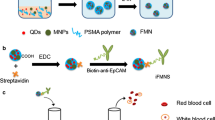
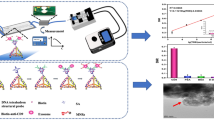
Circulating tumor DNA (ctDNA) is a tumor-derived fragmented DNA in the bloodstream that is not associated with cells. It has been greatly focused in the recent decade because of its potential clinical utility for liquid biopsies. Development of ctDNA analytical techniques with high sensitivity and cost-efficiency will undoubtedly promote the clinical spread of ctDNA testing. In this paper, we propose a novel flow cytometry–based ctDNA sensing strategy which combines enzyme-free amplification and magnetic separation. The target DNA is capable of triggering a hybridization chain reaction, producing a fluorescent long linear assembly of DNA, which can be further captured by magnetic beads to present fluorescent signals using flow cytometry. In comparison with some conventional methods, our strategy has the advantages of easy operation and cost-efficiency, and thereby shows a promising application in clinical diagnosis.

Graphical abstract









Similar content being viewed by others
References
Xiao Han, Junyun Wang, Sun. Y (2017) Circulating tumor DNA as biomarkers for cancer detection. Genom Proteom Bioinf 15 (2):59–72. doi:https://doi.org/10.1016/j.gpb.2016.12.004
Ogawara D, Soda H, Suyama T, Yoshida M, Harada T, Fukuda Y, et al. Pitfalls in diagnosis with the use of circulating tumor-derived epidermal growth factor receptor mutations in lung cancer harboring pretreatment T790M. Thorac Cancer. 2017;9(1):171–4. https://doi.org/10.1111/1759-7714.12538.
Tiberio P, Callari M, Angeloni V, Daidone MG, Appierto V. Challenges in using circulating miRNAs as cancer biomarkers. Biomed Res Int. 2015;2015(731479). https://doi.org/10.1155/2015/731479.
Kishikawa T, Otsuka M, Ohno M, Yoshikawa T, Takata A, Koike K. Circulating RNAs as new biomarkers for detecting pancreatic cancer. World J Gastroenterol. 2015;21(28):8527–40. https://doi.org/10.3748/wjg.v21.i28.8527.
Balmaseda A, Stettler K, Medialdea-Carrera R, Collado D, Jin X, Zambrana JV, et al. Antibody-based assay discriminates Zika virus infection from other flaviviruses. P Natl Acad Sci USA. 2017;114(31):8384–9. https://doi.org/10.1073/pnas.1704984114.
Wee EJH, Wang YL, Tsao SCH, Trau M. Simple, sensitive and accurate multiplex detection of clinically important melanoma DNA mutations in circulating tumour DNA with SERS nanotags. Theranostics. 2016;6(10):1506–13. https://doi.org/10.7150/thno.15871.
Tan YF, Tang TT, Xu HS, Zhu CQ, Cunningham BT. High sensitivity automated multiplexed immunoassays using photonic crystal enhanced fluorescence microfluidic system. Biosens Bioelectron. 2015;73:32–40. https://doi.org/10.1016/j.bios.2015.05.041.
Das J, Ivanov I, Sargent EH, Kelley SO. DNA clutch probes for circulating tumor DNA analysis. J Am Chem Soc. 2016;138(34):11009–16. https://doi.org/10.1021/jacs.6b05679.
Bellassai N, Spoto G. Biosensors for liquid biopsy: circulating nucleic acids to diagnose and treat cancer. Anal Bioanal Chem. 2016;408(26):7255–64. https://doi.org/10.1007/s00216-016-9806-3.
Qi L, Xiao M, Wang X, Wang C, Wang L, Song S, et al. DNA-encoded Raman-active anisotropic nanoparticles for microRNA detection. Anal Chem. 2017;89(18):9850–6. https://doi.org/10.1021/acs.analchem.7b01861.
Farimani AB, Dibaeinia P, Aluru NR. DNA Origami-graphene hybrid nanopore for DNA detection. ACS Appl Mater Interfaces. 2017;9(1):92–100. https://doi.org/10.1021/acsami.6b11001.
Wang D, Tang W, Wu X, Wang X, Chen G, Chen Q, et al. Highly selective detection of single-nucleotide polymorphisms using a quartz crystal microbalance biosensor based on the toehold-mediated strand displacement reaction. Anal Chem. 2012;84(16):7008–14. https://doi.org/10.1021/ac301064g.
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356(6336):438–42. https://doi.org/10.1126/science.aam9321.
Qu X, Zhu D, Yao G, Su S, Chao J, Liu H, et al. An exonuclease III-powered, on-particle stochastic DNA walker. Angew Chem Int Ed. 2017;56(7):1855–8. https://doi.org/10.1002/anie.201611777.
Lo Giudice MC, Herda LM, Polo E, Dawson KA. In situ characterization of nanoparticle biomolecular interactions in complex biological media by flow cytometry. Nat Commun. 2016;7:13475. https://doi.org/10.1038/ncomms13475.
Alvarez-Cubero MJ, Santiago O, Martinez-Labarga C, Martinez-Garcia B, Marrero-Diaz R, Rubio-Roldan A, et al. Methodology for Y chromosome capture: a complete genome sequence of Y chromosome using flow cytometry, laser microdissection and magnetic streptavidin-beads. Sci Rep. 2018;8(1):9436. https://doi.org/10.1038/s41598-018-27819-x.
van Andel E, de Bus I, Tijhaar EJ, Smulders MMJ, Savelkoul HFJ, Zuilhof H. Highly specific binding on antifouling zwitterionic polymer-coated microbeads as measured by flow cytometry. ACS Appl Mater Interfaces. 2017;9(44):38211–21. https://doi.org/10.1021/acsami.7b09725.
Yu B, Li F, Zhao T, Li F, Zhou B, Xu H. Hybridization chain reaction-based flow cytometric bead sensor for the detection of emetic Bacillus cereus in milk. Sensors Actuators B Chem. 2018;256:624–31. https://doi.org/10.1016/j.snb.2017.09.199.
Muratore KA, Grundhofer HM, Arriaga EA. Capillary electrophoresis with laser-induced fluorescent detection of immunolabeled individual autophagy organelles isolated from liver tissue. Anal Chem. 2016;88(23):11691–8. https://doi.org/10.1021/acs.analchem.6b03270.
Poulcharidis D, Belfor K, Kros A, van Kasteren SI. A flow cytometry assay to quantify intercellular exchange of membrane components. Chem Sci. 2017;8(8):5585–90. https://doi.org/10.1039/c7sc00260b.
Daniele JR, Heydari K, Arriaga EA, Dillin A. Identification and characterization of mitochondrial subtypes in Caenorhabditis elegans via analysis of individual mitochondria by flow cytometry. Anal Chem. 2016;88(12):6309–16. https://doi.org/10.1021/acs.analchem.6b00542.
Xu J, Wang Y, Yang L, Gao Y, Li B, Jin Y. A cytometric assay for ultrasensitive and robust detection of human telomerase RNA based on toehold strand displacement. Biosens Bioelectron. 2017;87:1071–6. https://doi.org/10.1016/j.bios.2016.08.038.
Zhu LP, Chen DS, Lu XH, Qi Y, He P, Liu CH, et al. An ultrasensitive flow cytometric immunoassay based on bead surface-initiated template-free DNA extension. Chem Sci. 2018;9(32):6605–13. https://doi.org/10.1039/c8sc02752h.
Dirks RM, Pierce NA. Triggered amplification by hybridization chain reaction. P Natl Acad Sci USA. 2004;101(43):15275–8. https://doi.org/10.1073/pnas.0407024101.
Bi S, Yue S, Zhang S. Hybridization chain reaction: a versatile molecular tool for biosensing, bioimaging, and biomedicine. Chem Soc Rev. 2017;46(14):4281–98. https://doi.org/10.1039/c7cs00055c.
Wu S, Peng P, Wang H-H, Li T. Stimuli-Triggered strand displacement-based multifunctional gene detection platform controlled by a multi-input DNA logic gate. Chin J Anal Chem. 2018;46(5):E1832–7. https://doi.org/10.1016/s1872-2040(18)61084-9.
Seelig G, Soloveichik D, Zhang DY, Winfree E. Enzyme-free nucleic acid logic circuits. Science. 2006;314(5805):1585–8. https://doi.org/10.1126/science.1132493.
Yurke B, Turberfield AJ, Mills AP, Simmel FC, Neumann JL. A DNA-fuelled molecular machine made of DNA. Nature. 2000;406(6796):605–8. https://doi.org/10.1038/35020524.
Khodakov DA, Khodakova AS, Huang DM, Linacre A, Ellis AV. Protected DNA strand displacement for enhanced single nucleotide discrimination in double-stranded DNA. Sci Rep. 2015;5. https://doi.org/10.1038/srep08721.
Lai W, Ren L, Tang Q, Qu X, Li J, Wang L, et al. Programming chemical reaction networks using intramolecular conformational motions of DNA. ACS Nano. 2018;12(7):7093–9. https://doi.org/10.1021/acsnano.8b02864.
Qu X, Wang S, Ge Z, Wang J, Yao G, Li J, et al. Programming cell adhesion for on-chip sequential Boolean logic functions. J Am Chem Soc. 2017;139(30):10176–9. https://doi.org/10.1021/jacs.7b04040.
Zhang DY, Winfree E. Control of DNA strand displacement kinetics using toehold exchange. J Am Chem Soc. 2009;131(47):17303–14. https://doi.org/10.1021/ja906987s.
Genot AJ, Zhang DY, Bath J, Turberfield AJ. Remote toehold: a mechanism for flexible control of DNA hybridization kinetics. J Am Chem Soc. 2011;133(7):2177–82. https://doi.org/10.1021/ja1073239.
Meng X, Yang G, Li F, Liang T, Lai W, Xu H. Sensitive detection of Staphylococcus aureus with vancomycin-conjugated magnetic beads as enrichment carriers combined with flow cytometry. ACS Appl Mater Interfaces. 2017;9(25):21464–72. https://doi.org/10.1021/acsami.7b05479.
Chen C, Liu W, Tian S, Hong T. Novel surface-enhanced Raman spectroscopy techniques for DNA, protein and drug detection. Sensors-Basel. 2019;19(7). https://doi.org/10.3390/s19071712.
Baek S, Ahn JK, Won BY, Park KS, Park HG. A one-step and label-free, electrochemical DNA detection using metal ion-mediated molecular beacon probe. Electrochem Commun. 2019;100:64–9. https://doi.org/10.1016/j.elecom.2019.01.023.
Funding
This work is financially supported by the National Key R&D Program of China (2018YFC0115705), Youth Innovation Promotion Association CAS (2015261), the National Natural Science Foundation of China (Grant No. 31500805, 81873539, 21575088), “333 High-level Talents Training Project of Jiangsu Province,” and Science and Technology Program of Suzhou (SYG201508).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhai, X., Li, J., Cao, Y. et al. A nanoflow cytometric strategy for sensitive ctDNA detection via magnetic separation and DNA self-assembly. Anal Bioanal Chem 411, 6039–6047 (2019). https://doi.org/10.1007/s00216-019-01985-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01985-x