Abstract
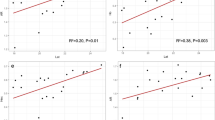
We employed a novel set of six highly polymophic chloroplastic simple sequence repeat (cpSSR) loci to investigate the phylogeography of lodgepole pine (Pinus contorta Dougl. Ex. Loud.), and to examine aspects of the evolutionary process operating on these repetitive DNA sequences. Chloroplast haplotypes of 500 trees, sampled throughout the range of lodgepole pine, were determined. We found a marked association of genetic distance with physical distance within the scale of 0 to 1,000 km, but no association beyond that range. Likewise, geographic clustering was observed only among recent clades in a dendrogram. These phylogeographic patterns are consistant with a rapid rangewide expansion (”big-bang”) followed by recent, local population differentiation (”galaxy formation”). In support of this expansion, coalescent simulations of the genealogical process gave a long-term effective population size in the low thousands, and a time to common ancestry of about 1,500 generations (12,000 years), consistent with a post-Pleistocene population expansion as documented by previous pollen-sediment analyses. Two lines of evidence (mapping mutational events onto a phylogeny, and evaluation of observed versus expected gene diversity) suggest that five of the cpSSR loci evolve primarily by a stepwise model of evolution of single repeat changes (but with a small proportion of changes involving two or more repeats), and the coalescent simulations point to a mutation rate of about 10–3.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 7 December 2000 / Accepted: 17 May 2001
Rights and permissions
About this article
Cite this article
Dawn Marshall, H., Newton, C. & Ritland, K. Chloroplast phylogeography and evolution of highly polymorphic microsatellites in lodgepole pine (Pinus contorta). Theor Appl Genet 104, 367–378 (2002). https://doi.org/10.1007/s001220100687
Issue Date:
DOI: https://doi.org/10.1007/s001220100687