Abstract
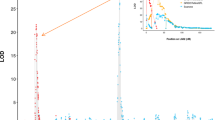
The molecular basis of resistance to diseases in plants can be better understood if the genes coding for resistance can be cloned. The single major dominant gene (R) that confers resistance to the white pine blister rust fungus (Cronartium ribicola Fisch.) in sugar pine (Pinus lambertiana Dougl.) has been previously mapped. The objectives of the present study were to saturate the region flanking R with tightly linked markers and to construct genetic maps for each of four individual seed trees. Bulked segregant analysis (BSA) and haploid segregation analysis were employed to identify random amplified polymorphic DNA (RAPD) markers linked to R. Automated PCR analysis was used to assay 1115 primers with susceptible and resistant DNA pools from each of four seed trees (8920 PCR reactions). Thirteen RAPD loci were identified that were linked to R. The linkage analyses programs JoinMap 1.4 and Mapmaker 2.0 were used to order RAPD loci relative to R and to construct maps for each of the individual seed trees. Two seed trees, 5701 and 6000, had a large number of tightly linked markers flanking R. These trees will be used in subsequent high-resolution mapping experiments to identify very tightly linked markers to facilitate the eventual cloning of R.
Similar content being viewed by others
Author information
Authors and Affiliations
Consortia
Additional information
Received: 1 May 1998 / Accepted: 13 July 1998
Rights and permissions
About this article
Cite this article
Harkins, D., G. N . Johnson., Skaggs, P. et al. Saturation mapping of a major gene for resistance to white pine blister rust in sugar pine. Theor Appl Genet 97, 1355–1360 (1998). https://doi.org/10.1007/s001220051029
Issue Date:
DOI: https://doi.org/10.1007/s001220051029