Abstract
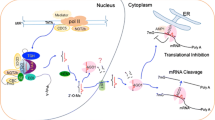
MicroRNAs (miRNAs), a type of endogenous non-coding small RNA, with a length of 20 to 24 nt, represses their target gene expression post-transcriptionally. There are a growing number of studies have discovered that miRNAs are found in animals and plants. Plant miRNAs are involved in growth and development, and they play an important role in physiological and biochemical regulation. This review mainly focuses on components of different miRNA processing pathways and how miRNA effectors inhibit the expression of target genes in plants. We discuss not only the various pathways affecting miRNA biogenesis, but also the different forms of miRNA effector pathways. Previous studies found that there have been some primary advances in the formation of miRNAs, we hope to provide a comprehensive introduction to the biogenesis of plant miRNAs.

Similar content being viewed by others
References
Anand A, Krichevsky A, Schornack S, Lahaye T, Tzfira T, Tang Y, Citovsky V, Mysore KS (2007) Arabidopsis VIRE2 INTERACTING PROTEIN2 is required for Agrobacterium T-DNA integration in plants. Plant Cell 19(5):1695–1708
Anjali N, Nadiya F, Thomas J, Sabu KK (2019) Identification and characterization of drought responsive microRNAs and their target genes in cardamom (Elettaria cardamomum Maton). Plant Growth Regul 87:201–216
Baumberger N, Baulcombe DC (2005) Arabidopsis ARGONAUTE1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102(119):28–33
Berardini TZ, Bollman K, Sun H, Poethig RS (2001) Regulation of vegetative phase change in Arabidopsis thaliana by cyclophilin 40. Science 291:2405–2407
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409:363–366
Bielewicz D, Kalak M, Kalyna M (2013) Introns of plant pri-miRNAs enhance miRNA biogenesis. EMBO Rep 14:622–628
Bologna NG, Mateos JL, Bresso EG, Palatnik JF (2009) A loop-to-base processing mechanism underlies the biogenesis of plant microRNAs miR319 and miR159. EMBO J 28:3646–3656
Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY et al (2008) Widespread translational inhibition by plant miRNAs and siRNAs. Science 320:1185–1190
Brodersen P, Sakvarelidze-Achard L, Schaller H, Khafif M, Schott G, Bendahmane A, Voinnet O (2012) Isoprenoid biosynthesis is required for miRNA function and af–fects membrane association of ARGONAUTE1 in Arabidopsis. Proc Natl Acad Sci USA 109:1778–1783
Cai Q, Liang C, Wang S, Hou Y, Gao L, Liu L, He W, Ma W, Mo B, Chen X (2018) The disease resistance protein SNC1 represses the biogenesis of microRNAs and phased siRNAs. Nat Commun 9:5080
Calderon-Villalobos LI, Kuhnle C, Dohmann EM, Li H, Bevan M, Schwechheimer C (2005) The evolutionarily conserved TOUGH protein is required for proper development of Arabidopsis thaliana. Plant Cell 17:2473–2485
Chaabane SB, Liu R, Chinnusamy V, Kwon Y, Park J, Kim SY, Zhu JK, Yang SW, Lee B (2013) STA1, an Arabidopsis pre-miRNA processing factor 6 homolog, is a new player involved in miRNA biogenesis. Nucleic Acid Res 41:1984–1997
Chadick JZ, Asturias FJ, Biochem T (2005) Structure of eukaryotic mediator complexes. Science 30:264–271
Cho SK, Ben Chaabane S, Shah P, Poulsen CP, Yang SW (2014) COP1 E3 ligase protects HYL1 to retain microRNA biogenesis. Nat Commun 5:5867
Clapier CR, Iwasa J, Cairns BR, Peterson CL (2017) Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol 18:407–422
Cui YW, Fang XF, Qi YJ (2016) TRANSPORTIN1 promotes the association of MicroRNA with AGONAUTE1 in Arabidopsis. Plant Cell 28:2576–2585
Cuperus JT, Montgomery TA, Fahlgren N, Burke RT, Townsend T, Sullivan CM, Carrington JC (2010) Identification of MIR390a precursor processing-defective mutantsin Arabidopsis by direct genome sequencing. Proc Natl Acad Sci USA 107:466–471
Deleris A, Gallego-Bartolome J, Bao J, Kasschau KD, Carrington JC, Voinnet O (2006) Hierarchical action and inhibition of plant dicer-like proteins in antiviral defense. Science 313:68–71
Derrien B, Baumberger N, Schepetilnikov M, Viotti C, De Cillia J, Ziegler-Graff V, Isono E, Schumacher K, Genschik P (2012) Degradation of the antiviral component ATGONAUTE1 by the autophagy pathway. Proc Natl Acad Sci USA 109:15942–15946
Dong Z, Han MH, Fedoroff N (2008) The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc Natl Acad Sci USA 105:9970–9975
Eamens AL, Smith NA, Curtin SJ, Wang MB, Waterhouse PM (2009) The Arabidopsis thaliana double-stranded RNA binding protein DRB1 directs guide strand selection from microRNA duplexes. RNA 15:2219–2235
Earley K, Smith M, Weber R, Gregory BD, Poethig RS (2010) An endogenous F box protein regulates ARGONAUTE1 in Arabidopsis thaliana. Silence 1:15
Fang X, Cui Y, Li Y, Qi Y (2015a) Transcription and processing of primary microRNAs are coupled by elongator complex in Arabidopsis. Nat Plants 1:15075
Fang X, Shi Y, Lu X, Chen Z, Qi Y (2015b) CMA33/XCT regulates small RNA production through modulating the transcription of dicer-like genes in Arabidopsis. Mol Plant 8:1227–1236
Fang X, Zhao G, Zhang S, Li Y, Gu H, Li Y, Zhao Q, Qi Y (2019) Chloroplast-to-nucleus signaling regulates microRNA biogenesis in Arabidopsis. Dev Cell 48:371–382
Farrona S, Hurtado L, Bowman JL, Reyes JC (2004) The Arabidopsis thaliana SNF2 homolog AtBRM controls shoot development and flowering. Development 131:4965–4975
Feng S, Xu Y, Guo C, Zheng J, Zhou B, Zhang Y, Ding Y, Zhang L, Zhu Z, Wang H, Wu G (2016) Modulation of miR156 to identify traits associated with vegetative phase change in tobacco (Nicotiana tabacum). J Exp Bot 67(5):1493
Feng S, Zhang J, Mu Z, Wang Y, Wen C, Wu T, Yu C, Li Z, Wang H (2020) Recent progress on the molecular breeding of Cucumis sativus L. in China. Theor Appl Genet 133:1777–1790
Francisco-Mangilet AG, Karlsson P, Kim MH, Eo HJ, Oh SA, Kim JH, Kulcheski FR, Park SK, Manavella PA (2015) THO2, a core member of the THO/TREX complex, is required for microRNA production in Arabidopsis. Plant J 82:1018–1029
Fujii H, Zhu JK (2009) Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Natl Acad Sci USA 106:8380–8385
Furumizu C, Tsukaya H, Komeda Y (2010) Characterization of EMU, the Arabidopsis homolog of the yeast THO complex member HPR1. RNA 16:1809–1817
Grant PA, Duggan L, Cote J, Roberts SM, Brownell JE, Candau R, Ohba R, Owen-Hughes T, Allis CD, Winston F, Berger SL, Workman JL (1997) Yeast Gcn5 funtion in two multisubunit complexes to acetylate nucleosomal histones: characterization of an Ada complex and the SAGA (Spt/Ada) complex. Genes Dev 11:1640–1650
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (2005) Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123:631–640
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Hajheidari M, Farrona S, Huettel B, Koncz Z, Koncz C (2012) CDKF;1 and CDKD protein kinases regulate phosphorylation of serine residues in the C-terminal domain of Arabidopsis RNA polymerase II. Plant Cell 24:1626–1642
Helliwell CA, Chin-Atkins AN, Wilson IW, Chapple R, Dennis ES, Chaudhury A (2001) The Arabidopsis AMP1 gene encodes a putative glutamate carboxypeptidase. Plant Cell 13:2115–2125
Huang X, Feng J, Wang R, Zhang H, Huang J (2018) Comparative analysis of microRNAs and their targets in the roots of two cultivars with contrasting salt tolerance in rice (Oryza sativa L.). Plant Growth Regul 87:139–148
Hurtado L, Farrona S, Reyes JC (2006) The putative SWI/SNF complex subunit BRAHMA activates flower homeotic genes in Arabidopsis thaliana. Plant Mol Biol 62:291–304
Hutvagner G, Simard MJ (2008) Argonaute proteins: key players in RNA silencing. Nat Struct Mol Biol 9:22–32
Iki T, Yoshikawa M, Nishikiori M, Jaudal MC, Matsumoto-Yokoyama E, Mitsuhara I, Meshi T, Ishikawa M (2010) In vitro assembly of plant RNA-induced silencing complexes facilitated by molecular chaperone HSP90. Mol Cell 39:282–291
Jia L, Zhang B, You C, Zhang Y, Zeng L, Li S, Johnson KCM, Yu B, Li X, Chen X (2017) The Arabidopsis MOS4-associated complex promotes microRNA biogenesis and precursor messenger RNA splicing. Plant Cell 29:2626–2643
Jung JH, Seo PJ, Ahn JH, Park CM (2012) Arabidopsis RNA-binding protein FCA regulates microRNA172 processing in thermosensory flowering. J Biol Chem 287:16007–16016
Karlsson P, Christie MD, Seymour DK, Wang H, Wang X, Hagmann J, Kulcheski F, Manacella PA (2015) KH domain protein RCF3 is a tissue-biased regulator of the plant miRNA biogenesis cofactor HYL1. Proc Natl Acad Sci USA 112:14096–14101
Kierzkowski D, Kmieciak M, Piontek P, Wojtaszek P, Szweykowska-Kulinska Z, Jarmolowski A (2009) The Arabidopsis CBP20 targets the cap-binding complex to the nucleus, and is stabilized by CBP80. Plant J 59:814–825
Kim BH, Von Arnim AG (2009) FIERY1 regulates light-mediated repression of cell elongation and flowering time via its 3’ (2’), 5’-bisphosphate nucleotidase activity. Plant J 58:208–219
Kim W, Benhamed M, Servet C, Latrasse D, Zhang W, Delarue M, Zhou DX (2009) Histone acetyltransferase GCN5 interferes with the miRNA pathway in Arabidopsis. Proc Natl Acad Sci USA 19(7):899–909
Kim S, Yang JY, Xu J, Jang IC, Prigge MJ, Chua NH (2008) Two cap-binding proteins CBP20 and CBP80 are involved in processing primary microRNAs. Plant Cell Physiol 49:1634–1644
Kim YJ, Zheng B, Yu Y, Won SY, Mo B, Chen X (2011) The role of Mediator in small and long noncoding RNA production in Arabidopsis thaliana. EMBO J 30:814–822
Köhler A, Hurt E (2007) Exporting RNA from the nucleus to the cytoplasm. Nat Rev Mol Cell Biol 8:761–773
Koiwa H, Barb AW, Xiong L, Li F, McCully MG, Lee BH, Sokolchik I, Zhu J, Gong Z, Reddy M, Sharkhuu A, Manabe Y, Yokoi S, Zhu JK, Bressan RA, Hasegawa PM (2002) C-terminal domain phosphatase-like family members (AtCPLs) differentially regulate Arabidopsis thaliana abiotic stress signaling, growth, and development. Proc Natl Acad Sci USA 99:10893–10898
Kurihara Y, Takashi Y, Watanabe Y (2006) The interaction between DCL1 and HY L1 is important for efficient and precise processing of pri-miRNA in plant microRNA biogenesis. RNA 12(2):206–212
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through dicer-like 1 protein functions. Proc Natl Acad Sci USA 101:12753–12758
Lafos M, Kroll P, Hohenstatt ML, Thorpe FL, Clarenz O, Schubert D (2011) Dynamic regulation of H3K27 Trimethylation during Arabidopsis differentiation. PLoS Genet 7:1002040
Lakhwani D, Pandey A, Sharma D, Asif MH, Trivedi PK (2020) Novel microRNAs regulating ripening-associated processes in banana fruit. Plant Growth Regul 90:223–235
Lanet E, Delannoy E, Sormani R, Floris M, Brodersen P, Crete P, Olivier V, Tobaglia C (2009) Biochemical evidence for translational repression by Arabidopsis microRNAs. Plant Cell 21:1762–1768
Lee BH, Kapoor A, Zhu J, Zhu JK (2006) STABILIZED1, a stress upregulated nuclear protein, is required for pre-mRNA splicing, mRNA turnover, and stress tolerance in Arabidopsis. Plant Cell 18:1736–1749
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060
Leng X, Wang P, Zhao P, Wang M, Cui L, Shangguan L, Wang C (2017) Conservation of microRNA-mediated regulatory networks in response to copper stress in grapevine. Plant Growth Regul 82:293–304
Li J, Yang Z, Yu B, Liu J, Chen X (2005) Methylation protects miRNAs and siRNAs from a 30-end uridylation activity in Arabidopsis. Curr Biol 15:1501–1507
Li S, Liu L, Zhuang X, Yu Y, Liu X, Cui X, Ji L, Pan Z, Cao X, Mo B, Zhang F, Raikhel N, Jiang L, Chen X (2013) MicroRNAs inhibit the translation of target mRNAs on the endoplasmic reticulum in Arabidopsis. Cell 153:562–574
Li S, Liu K, Zhang S, Wang X, Rogers K, Ren G, Zhang C, Yu B (2017) STV1, a ribosomal protein, binds primary microRNA transcripts to promote their interacion with the processing complex in Arabidopsis. Proc Natl Acad Sci USA 114(6):1424–1429
Li S, Xu R, Li A, Liu K, Gu L, Li M, Zhang H, Zhang Y, Zhang S, Wang Q, Gao G, Li N, Zhang C, Li Y, Yu B (2018) SMA1, a homolog of the splicing factor Prp28, has a multifaceted role in miRNA biogenesis in Arabidopsis. Nucleic Acids Res 47:9148–9159
Liu C, Axtell MJ, Fedoroff NV (2012) The helicase and RNaseIIIa domains of Arabidopsis DCL1 modulate catalytic parameters during microRNA biogenesis. Plant Physiol 159:748–758
Lobbes D, Rallapalli G, Schmidt DD, Martin C, Clarke J (2006) SERRATE: a new player on the plant microRNA scene. EMBO Rep 7:1052–1058
Lotfi A, Pervaiz T, Jiu S, Faghihi F, Jahanbakhshian Z, Khorizoghi EG, Fang J, Mahdi Seyedi S (2017) Role of microRNAs and their target genes in salinity response in plants. Plant Growth Regul 82:377–390
Lu C, Fedoroff N (2000) A mutation in the Arabidopsis HYL1 gene encoding a dsRNA binding protein affects responses to abscisic acid, auxin, and cytokinin. Plant Cell 12:2351–2366
Lund E, Güttinger S, Calado A, Dahlberg JE, Kutay U (2004) Nuclear export of microRNA precursors. Science 303:95–98
Ma C, Burd S, Lers A (2015) miR408 is involved in abiotic stress responses in Arabidopsis. Plant J 84:169–187
Machida S, Chen HY, Yuan YA (2011) Molecular insights into miRNA processing by Arabidopsis thaliana SERRATE. Nucleic Acids Res 39:7828–7836
Macrae IJ, Zhou K, Li F, Repic A, Brooks AN, Cande WZ, Adams PD, Doudna JA (2006) Structural Basis for Double-Stranded RNA Processing by Dicer. Science 311:195–198
Manavella P, Hagmann J, Ott F, Laubinger S, Franz M, Macek B, Weigel D (2012) Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell 151:859–870
Mi S, Cai T, Hu Y, Chen Y, Hodges E, Ni F, Wu L, Li S, Zhou H, Long C, Chen S, Hannon GJ, Qi Y (2008) Sorting of small RNAs into Arabidopsis argonaute complex is directed by the 5’ terminal nucleotide. Cell 133:116–127
Montgomery TA, Howell MD, Cuperus JT, Li D, Hansen JE, Alexander AL, Chapman EJ, Fahlgren N, Allen E, Carrington JC (2008) Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133:128–141
Ori N, Cohen AR, Etzioni A, Arnon B, Yanai O, Sharona S, Menda N, Amsellem Z, Efroni I, Pekker I, Alvarez JP, Blum E, Zamir D, Eshed Y (2007) Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet 39:787–791
Park MY, Wu G, Gonzalez-Sulser A, Vaucheret H, Poethig RS (2005) Nuclear processing and export of microRNAs in Arabidopsis. Proc Natl Acad Sci USA 102:3691–3696
Raghuram B, Sheikh AH, Rustagi Y, Sinha AK (2015) MicroRNA biogenesis factor DRB1 is a phosphorylation target of mitogen activated protein kinase MPK3 in both rice and Arabidopsis. FEBS J 282:521–536
Ré D, Cambiagno DA, Arce AL, Tomassi AH, Giustonzzi M, Casati P, Ariel FD, Manavella PA (2020) CURLY LEAF regulates microRNA activity by controlling ARGONAUTE 1 degradation in plants. Mol Plant 13:72–87
Reed R, Cheng H (2005) TREX, SR proteins and export f mRNA. Curr Opin Cell Biol 17:269–273
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR, Ruvkun G (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403:901–906
Ren G, Chen X, Yu B (2012a) Uridylation of miRNAs by hen1 suppressor in Arabidopsis. Curr Biol 22:695–700
Ren G, Xie M, Dou Y, Zhang S, Zhang C, Bin Y (2012b) Regulation of miRNA abundance by RNA binding protein TOUG-H in Arabidopsis. Proc Natl Acad Sci USA 109:12817–12821
Schauer SE, Jacobsen SE, Meinke DW, Ray A (2002) DICER-LIKE1: blind men and elephants in Arabidopsis development. Trends Plant Sci 7:487–491
Schwab R, Speth C, Laubinger S (2013) Enhanced microRNA accumulation through stemloop-adjacent introns. EMBO Rep 14:615–621
Seo HS, Yang JY, Ishikawa M, Bolle C, Ballesteros ML, Chua NH (2003) LAF1 ubiquitination by COP1 controls photomorphogenesis and is stimulated by SPA1. Nature 423:995–999
Silva GFF, Silva EM, Azevedo MS, Guivin MAC, Ramiro DA, Figueiredo CR, Carrer H, Peres LEP, Nogueira FTS (2014) microRNA156-targeted SPL/SBP box transcription factors regulate tomato ovary and fruit development. Plant J 78:604–618
Smith MR, Willmann MR, Wu G, Berardini TZ, Möller B, Weijers D, Poethig RS (2009) Cyclophilin 40 is required for microRNA activity in Arabidopsis. Proc Natl Acad Sci USA 106:5424–5429
Song J, Wang X, Song B, Gao L, Mo X, Yue L, Yang H, Lu J, Ren G, Mo B, Chen X (2019) Prevalent cytidylation and uridylation of precursor miRNAs in Arabidopsis. Nat Plants 5:1260–1272
Song L, Axtell MJ, Fedoroff NV (2010) RNA secondary structural determinants of miRNA precursor processing in Arabidopsis. Curr Biol 20:37–41
Song L, Han MH, Lesicka J, Fedoroff N (2007) Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc Natl Acad Sci USA 104:5437–5442
Strauss EJ, Guthrie C (1994) PRP28, ‘DEAD-box’ protein, is required for the first step of mRNA splicing in vitro. Nucleic Acids Res 22:3187–3193
Su C, Li Z, Cheng J, Li L, Zhong S, Liu L, Zheng Y, Zheng B (2017) The protein Phosphatase 4 and SMEK1 complex dephosphorylates HYL1 to promote miRNA biogenesis by antagonizing the MAPK cascade in Arabidopsis. Dev Cell 41:527–539
Sunkar R, Kapoor A, Zhu JK (2006) Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18:2051–2065
Sun Z, Guo T, Liu Y, Liu Q, Fang Y (2015) The roles of Arabidopsis CDF2 in transcriptional and posttranscriptional regulation of primary microRNAs. PLoS Gene 11:1005598
Tagami Y, Motose H, Watanabe Y (2009) A dominant mutation in DCL1 suppresses the hyl1 mutant phenotype by promoting the processing of miRNA. RNA 15:450–458
Takeda A, Iwasaki S, Watanabe T, Utsumi M, Watanabe Y (2008) The mechanism selecting the guide strand from small RNA duplexes is different among argonaute proteins. Plant Cell Physiol 49:493–500
Tian L, Chen ZJ (2011) Blocking histone deacetylation in Arabidopsis induces pleiotropic effects on plant gene regulation and development. Proc Natl Acad Sci USA 98:200–205
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687
Wen Y, Yue Z, Shatkin AJ (1998) Mammalian capping enzyme binds RNA and uses protein tyrosine phosphatase mechanism. Proc Natl Acad Sci USA 95:12226–12231
Wightman B, Ha I, Ruvkun G (1993) Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75:855–862
Wang B, Xue Y, Zhang Z, Ding D, Fu Z, Tang J (2016) Transcriptomic analysis of maize kernel row number-associated miRNAs between a single segment substitution line and its receptor parent. Plant Growth Regul 78:145–154
Wang H, Yu C, Fan P, Bao B, Li T, Zhu Z (2014) Identification of two cucumber putative silicon transporter genes in Cucumis sativus. J Plant Growth Regul 34(2):332–338
Wang H, Zhu Z, Feng Z, Zhang SG, Yu C (2012) Antisense-mediated depletion of GMPase gene expression in tobacco decreases plant tolerance to temperature stresses and alters plant development. Mol Biol Rep 39(12):10413–10420
Wang L, Song X, Gu L, Li X, Cao S, Chu C, Cui X, Chen X, Cao X (2013) NOT2 proteins promote polymerase II-dependent transcription and interact with multiple microRNA biogenesis factors in Arabidopsis. Plant Cell 25:715–727
Wang L, Sun S, Jin J, Fu D, Yang X, Weng X, Xu C, Li X, Xiao J, Zhang Q (2015a) Coordinated regulation of vegetative and reproductive branching in rice. Proc Natl Acad Sci USA 112(50):15504–15509
Wang J, Chen S, Jiang N, Li N, Wang X, Li Z, Li X, Liu H, Li L, Yang Y, Ni T, Yu C, Ma J, Zheng B, Ren G (2019a) Spliceosome disassembly factors ILP1 and NTR1 promote miRNA biogenesis in Arabidopsis thaliana. Nucleic Acids Res 47(15):7886–7900
Wang S, Quan L, Li S, You C, Zhang Y, Gao L, Zeng L, Liu L, Qi Y, Mo B, Chen X (2019b) The PROTEIN PHOSPHATASE4 complex promotes transcription and processing of primary microRNAs in Arabidopsis. Plant Cell 31:486–501
Wang W, Ye R, Xin Y, Fang X, Li C, Shi H, Zhou X, Qi Y (2011) An importin β protein negatively regulates microRNA activity in Arabidopsis. Plant Cell 23:3565–3576
Wang X, Wang Y, Dou Y, Chen L, Wang J, Jiang N, Guo C, Yao Q, Wang C, Liu L, Bin Y, Zheng B, Chekanova JA, Ma J, Ren G (2017) Degradation of unmethylated miRNA/miRNA*s by a DEDDy-type 3’ to 5’ exoribonuclease Atrimmer 2 in Arabidopsis. Proc Natl Acad Sci USA 115:6659–6667
Wang X, Zhang S, Dou Y, Zhang C, Chen X, Yu B, Ren G (2015b) Synergistic and independent actions of multiple terminal nucleotidyl transferases in the 3’tailing of small RNAs in Arabidopsis. PLoS Genet 11:1005091
Wang Z, Ma Z, Castillo-González C, Sun D, Li Y, Yu B, Zhao B, Li P, Zhang X (2018) SWI2/SNF2 ATPase CHR2 remodels pri-miRNAs via Serrate to impede miRNA production. Nature 557:516–521
Wu F, Yu L, Cao W, Mao Y, Liu Z, He Y (2007) The N-terminal double-stranded RNA binding domains of Arabidopsis HYPONASTIC LEAVES1 are sufficient for pre-microRNA processing. Plant Cell 19:914–925
Wu G, Park MY, Conway SR, Wang JW, Weigel D, Poethig RS (2009) The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138:750–759
Wu H, Li B, Iwakawa H, Pan Y, Tang X, Ling-hu Q, Liu Y, Sheng S, Feng L, Zhang H, Zhang X, Tang Z, Xia X, Zhai J, Guo H (2019) Plant 22-nt siRNAs mediate translational repression and stress adaptation. Nature 7806(581):89–93
Wu X, Shi Y, Li J, Xu L, Fang Y, Li X, Qi Y (2013) A role for the RNA-binding protein MOS2 in microRNA maturation in Arabidopsis. Cell Res 23:645–657
Xie Z, Allen E, Fahlgren N, Calamar A, Givan SA, Carrington JC (2005) Expression of Arabidopsis MIRNA genes. Plant Physiol 138:2145–2154
Xiong L, Lee H, Ishitani M, Tanaka Y, Stevenson B, Koiwa H, Bressan RA, Hasegawa PM, Zhu JK (2002) Repression of stress-responsive genes by FIERY2, a novel transcriptional regulator in Arabidopsis. Proc Natl Acad Sci USA 99:10899–20904
Xu F, Xu S, Wiermer M, Zhang Y, Li X (2012) The cyclin L homolog MOS12 and the MOS4-associated complex are required for the proper splicing of plant resistance genes. Plant J 70:916–928
Xu J, Yang JY, Niu QW, Chua NH (2006) Arabidopsis DCP1, DCP2 and VARICOSE form a decapping complex required for postembryonic development. Plant Cell 18(12):3386–3398
Xu M, Leichty AR, Hu T, Poethig RS (2018) H2A.Z promotes the transcription of MIR156A and MIR156C in Arabidopsis by facilitating the deposition of H3K4me3. Development 145(2):152868
Xu Y, Guo C, Zhou BY, Li C, Wang H, Zheng B, Ding Y, Zhu Z, Peragine A, Cui Y, Poethig S, Gang Wu (2016) Regulation of vegetative phase change by SWI2/SNF2 chromatin remodeling ATPase BRAHMA. Plant Physiol 172:2416–2428
Yan J, Wang P, Wang B, Hsu CC, Tang K, Zhang H, Hou YJ, Zhao Y, Wang Q, Zhao C, Zhu X, Tao WA, Li J, Zhu JK (2017) The SnRK2 kinases modulate miRNA accumulation in Arabidopsis. PLoS Genet 13(4):1006753
Yang L, Liu Z, Lu F, Dong A, Huang H (2006) SERRATE is a novel nuclear regulator in primary microRNA processing in Arabidopsis. Plant J 47:841–850
Yang L, Wu G, Poethig RS (2012) Mutations in the GW-repeat protein SUO reveal a developmental function for microRNA-mediated translational repression in Arabidopsis. Proc Natl Acad Sci USA 109:315–320
Yang SW, Chen HY, Yang J, Machida S, Chua NH, Yuan YA (2010) Structure of Arabidopsis HYPONASTIC LEAVES1 and its molecular implications for miRNA processing. Cell 18:594–605
Yang X, Ren W, Zhao Q, Zhang P, Wu F, He Y (2014) Homodimerization of HYL1 ensures the correct selection of cleavage sites in primary miRNA. Nucleic Acids Res 42:12224–12236
Yang Z, Huang Y, Yang J, Yao S, Zhao K, Wang D, Qin Q, Bian Z, Li Y, Lan Y, Zhou T, Wang H, Liu C, Wang W, Qi Y, Xu Z, Li Y (2020) Jasmonate signaling enhances RNA silencing and antiviral defense in rice. Cell Host Microb 28:89–103
Yu B, Bi L, Zheng B, Ji L, Chevalier D, Agarwal M, Ramachandran V, Li W, Lagrange T, Walker JC, Chen X (2008) The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc Natl Acad Sci USA 105:10073–10078
Yu B, Yang Z, Li J, Minakhina S, Yang M, Padgett RW, Steward R, Chen X (2005) Methylation as a crucial step in plant microRNA biogenesis. Science 307:932–935
You C, He W, Hang R, Zhang C, Cao X, Guo H, Chen X, Cui J, Mo B (2019) FIERY1 promotes microRNA accumulation by suppressing rRNA-derived small interfering RNAs in Arabidopsis. Nat Commun 10(1):1–15
Yu N, Niu QW, Ng KH, Chua NH (2015) The role of miR156/SPLs modules in Arabidopsis lateral root development. Plant J 83:673–685
Yu Y, Jia T, Chen X (2017a) The ‘how’ and ‘where’ of plant microRNAs. New Phytol 216(4):1002–1017
Yu Y, Ji L, Le BH, Zhai J, Chen J, Luscher E, Gao L, Liu C, Cao X, Mo B, Ma J, Meyers BC, Chen X (2017b) ARGONAUTE10 promotes the degradation of miR165/6 through the SDN1 and SDN2 exonucleases in Arabidopsis. PLoS Biol 15(2):2001272
Zeng Y, Cullen BR (2004) Structural requirements for pre-microRNA binding and nuclear export by exportin 5. Nucleic Acid Res 32:4776–4785
Zhang CJ, Zhou JX, Liu J, Ma ZY, Zhang SW, Dou K, Huang HW, Cai T, Liu R, Zhu JK, He XJ (2013a) The splicing machinery promotes RNA-directed DNA methylation and transcriptional silencing in Arabidopsis. EMBO J 32:1128–1140
Zhang L, Ding H, Jiang H, Wang H, Chen K, Duan J, Feng S, Wu G (2019) Regulation of cadmium tolerance and accumulation by miR156 in Arabidopsis. Chemosphere 242:125168
Zhang L, Volinia S, Bonome T et al (2008) Genomic and epigenetic alterations deregulate microRNA expression in human epithelial ovarian cancer. Proc Natl Acad Sci USA 105:7004–7009
Zhang S, Liu Y, Yu B (2014) PRL1, an RNA-binding protein, positively regulates the accumulation of miRNAs and siRNAs in Arabidopsis. PLoS Genet 10:1004841
Zhang S, Xie M, Ren G, Yu B (2013b) CDC5, a DNA binding protein, positively regulates posttranscriptional processing and/or transcription of primary microRNA transcripts. Proc Natl Acad Sci USA 110:17588–17593
Zhang X, Clarenz O, Cokus S, Bernatavichute YV, Pellegrini M, Goodrich J, Jacobsen SE (2007) Whole-genome analysis of histone H3 lysine 27 Trimethylation in Arabidopsis. PLoS Biol 5:129
Zhang Y, Cheng YT, Bi D, Palma K, Li X (2005) MOS2, a protein containing G-patch and KOW motifs, is essential for innate immunity in Arabidopsis thaliana. Curr Biol 15:1936–1942
Zhang Z, Guo X, Ge C, Ma Z, Jiang M, Li T, Koiwa H, Yang SW, Zhang X (2017a) KETCH1 imports HYL1 to nucleus for miRNA biogenesis in Arabidopsis. Proc Natl Acad Sci USA 114:4011–4016
Zhang Z, Hu F, Sung MW, Shu C, Castillo-Gonzalez C, Koiwa H, Tang G, Dickman M, Li P, Zhang X (2017b) RISC-interacting clearing 3’-5’ exoribonucleases (RICEs) degrade uridylated cleavage fragments to maintain functional RISC in Arabidopsis thaliana. eLife 6:24466
Zhan X, Wang B, Li H, Liu R, Kalia RK, Zhu JK, Chinnusamy V (2012) Arabidopsis proline rich protein important for development and abiotic stress tolerance is involved in microRNA biogenesis. Proc Natl Acad Sci USA 109(44):18198–18203
Zhao Y, Yu Y, Zhai J, Ramachandran V, Dinh TT, Meyers BC, Mo B, Chen X (2012) The Arabidopsis nucleotidyl transferase HESO1 uridylates unmethylated small RNAs to trigger their degradation. Curr Biol 22(8):689–694
Zhu H, Hu F, Wang R, Zhou X, Sze SH, Liou LW, Barefoot A, Dickman M, Zhang X (2011) Arabidopsis argonaute10 specifically sequesters miR166/165 to regulate shoot apical meristem development. Cell 145:242–256
Zhu H, Zhou Y, Castillo-González C, Lu A, Ge C, Zhao YT, Duan L, Li Z, Axtell MJ, Wang XJ, Zhang X (2013) Bidirectional processing of pri-miRNAs with branched terminal loops by Arabidopsis Dicer-like1. Nat Struct Mol Biol 20(9):1106–1115
Acknowledgements
This work was supported by the Public Projects of Zhejiang Province (No. LGN19C150007), National Natural Science Foundation of China (No. 31701915) and (Grant No. 31872105), and start-up fund from Zhejiang Agriculture & Forestry University (2017FR006). We thank Gang Wu for his advices on our review.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, Z., Nie, J. & Wang, H. MicroRNA biogenesis in plant. Plant Growth Regul 93, 1–12 (2021). https://doi.org/10.1007/s10725-020-00654-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00654-9